Task Decomposition and Synchronization for Semantic Biomedical Image Segmentation
Semantic segmentation is essentially important to biomedical image analysis. Many recent works mainly focus on integrating the Fully Convolutional Network (FCN) architecture with sophisticated convolution implementation and deep supervision. In this …
Authors: Xuhua Ren, Lichi Zhang, Sahar Ahmad
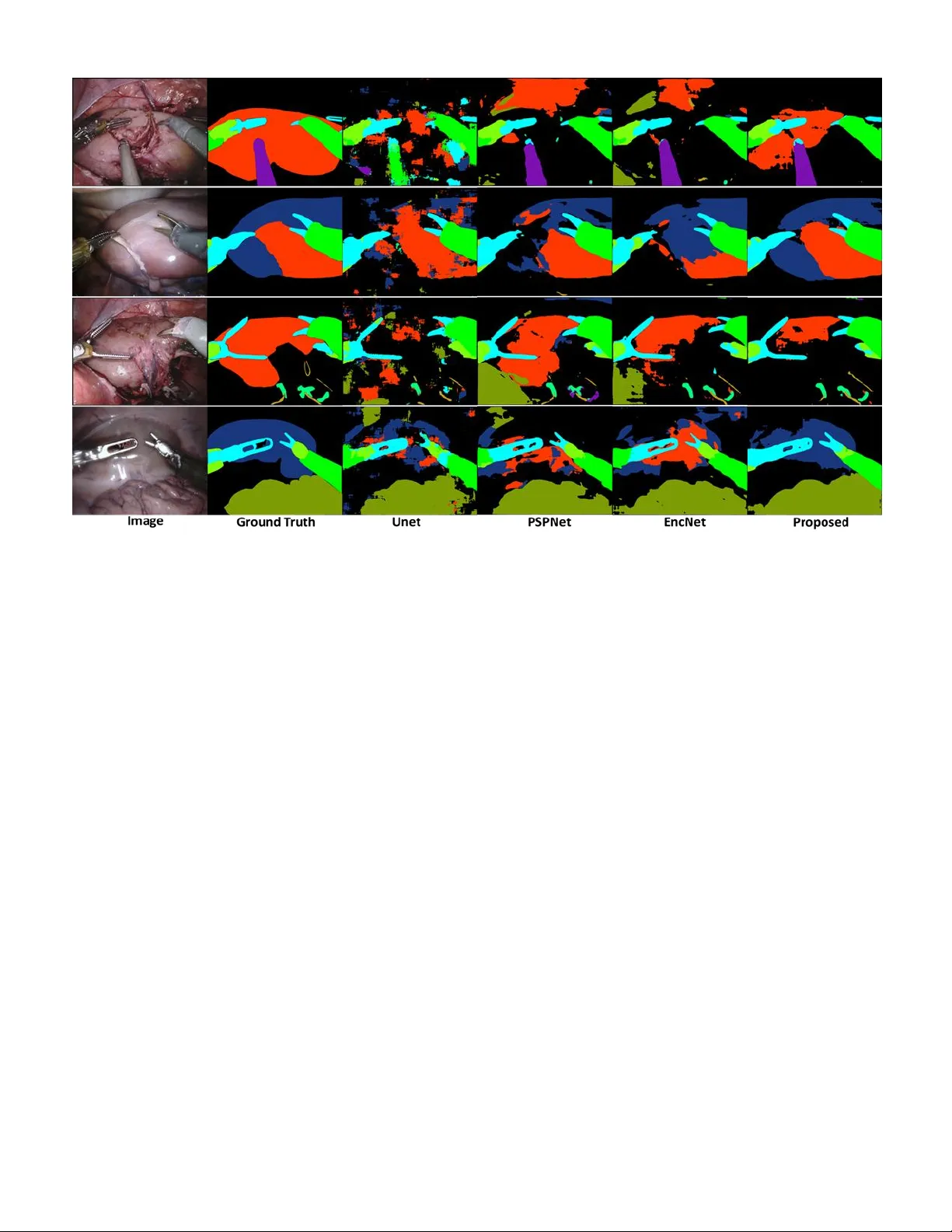
Abstract — Semantic segmentation is essentially important to biomedical image analysis. Many recent works mainly focus on integrating the Fully Convolutional Network (FCN) architecture with sophisticated c onvolution imp lementation and deep supervision. In this paper, w e propose to decompose the sing l e segmentation task into three subsequent sub-tasks, including (1) pixel-wise image segmentation, (2) prediction of t he class l abels of the objects within the image, and (3) classification of the scene the image belonging to. While the se three sub-ta sks are trained to optimize their individual loss functions of different percep tual levels, w e propose to let them interact by the task-task context ensemble. M oreover, w e propose a novel sync -regularization to penalize the deviation betw een the o utputs of t he p ix el-wise segmentation a nd t he class prediction tasks. These effective regularizations help FCN utilize context informa tion comprehensive ly and attain accu rate semantic segmentation, e ven though the nu mber of t he images fo r training m ay be li mited in many bio medica l ap plications. We h ave successfully app lied our framework to three diverse 2D/3 D medical image datasets, including Robotic Scene Seg mentation Challenge 18 (ROBOT18), Brain Tum or Segmentation Challenge 18 (BRATS18), a nd Re tinal Fundus Glaucoma Challenge (REFUG E18). We have achieved top-tier performance in all three challen ges. Index Terms — semantic segmentation, fully convolutional network, task decomposition, sync-regularization, deep learning I. I NTRODUCTI ON emantic seg mentation is a classical p roblem i n the field of computer v ision, where a pre-defined class label needs to be assigned to each pixel. The input image is thus divided into the regions corresponding to different class labels of a ce rtain scene [1]. A n optimal solution to segm entation usually relies on complicated representations including object class, location, scene and context. * C orresponding authors: Qian Wang (wang.qian@sjtu.edu.c n), Dinggang Shen (dgshen @med.unc.edu) Author, Xuhua Ren, Lichi Z hang, Xi ang L ei and Qian W ang are with the Institute for Medical Imaging Technology , School of B iomedical Engineering, Shanghai Jiao T ong Univer sity, 200030, S hanghai, China . Fig. 1 . Multi-level representations observed in BRATS18 and ROBOT18 datasets f or sem antic segmentation . Besides assigning pixel-wise label in the segmentation task, there are 4 classes and 2 scenes for BRATS18 , 12 classes and 3 scenes for ROBOT18 , respectively. The three tasks, as well as the ir multi-level representations, are closely coupled. Currently, m ost sta te- of -the-art semantic segmentation approaches are based on the FCN frame w ork [2 - 7] . FCN has a powerful enco der to extract image features. Then, the decoder gradually fuses the hig h-level features at top layers of the encoder with the lo w-level features at botto m layers, which is essential for the deco der to generate high -quality se mantic segmentation result [8 ]. The recent success of FCN can be attributed to the v ery deep network architecture, w hich poo ls the features into pyramid representations effectivel y. T he d eep supervision also contributes to the search for the net work para meters. F or example, EncNet [3] has a ResNet-101 encoder [9], and adopts Sahar Ahmad, Fan Yang, Do ng Nie and Dinggang Shen are with the Department of Radiology and BRI C, University of North Carolina at Chapel Hill, Chape l Hil l, Nor th Carolina 27599, USA. Dingga ng Shen is also with Department of Br ain and Cogn itive Enginee ring, Ko rea University, Se oul 02841, Republic of Ko rea. . T ask Decomposition and Synchronization for Semantic Biomedical Image Segm entation Xuhua Ren, Lichi Zha ng, Sahar Ahmad, Don g Nie, Fa n Yang, Lei Xiang, Qi an Wang*, Member, IEEE and Dinggang Sh en*, Fellow , IEEE S Fig. 2. The exam ple of ROBOT18 w ith the proposed task decomposition framework for segmentation. The image is processed through the encoder and task-tas k context ensemble to arrive at the latent space. Then, the segmentation, class, a nd scene tas ks are solved through individual decoders. A strong sync-regularization betw een the segm entation and class tasks is f urther used t o augm ent the coherence of m ulti-task learning. dilated convolution [10] in both encoder and dec oder. A context encoding module strengthens deep supervision b y incorporating semantic encoding loss. P yramid scene parsing network (P SPNet ) [2] has the traditio nal dilated F CN architecture for pix el prediction, w hile this network extends t he pixel-level features to the specially designed glob al p yramid pooling features. T he local and global cues to gether make the final prediction more reliable. However, a sophisticated n etwork in deep learning often needs a m assive a mount o f dat a to train. Whereas a co mmon yet critical challenge in bio medical image segmentation arises due to the limited size of the dataset – ROBOT 18 1 , REFUGE18 2 and BRATS18 [1 1], which are all widely used benchmark datasets in biomedical image segmentation and considered in this pap er as well, h ave o nly 2,235, 400 and 285 subjects f or training, respectively. To this end, it is e ssential to probe how to adapt the n etwork for small me dical image datasets. While the datasets used in this p aper are highly diverse, they all share the same goal of semantic segmentation. 1) Taking ROBOT 18 for example (c.f. Fig. 1 ), the d ataset is often used to validate instrument trac king a nd segmentati on, which enab les surgeo ns to conduct ro bot -assisted minimal invasive surgery (R MIS) m o re pr ecisely. The ima ge can be classified into three scenes , i.e., “preparing” surgery, “peeling” the covered kidney, and “incising” the kidne y parenchyma. The e xistence o f a cer tain clas s i n the image can also be determined, followed by ca reful pixel -wis e segmentation . 2) As a second example, BRATS18 is often used to validate automatic brain tumor segmentation, which is challe nging 1 https://endovis.gr and-challenge .org/ due to the diversity o f t umor lo cation, s hape, size and appearance [12]. Given a patient i mage, a clinician may first classify t he scene as high -grade glio ma (HGG) or low- grade glioma (LGG), since the t wo t ypes of tumors usually app ear different in images. T hen, it can be determined w hether the image renders certai n su b-re gion of the t umor (or class , including ET , NCR /NET, ED, etc.), follo wed by care ful pixel-wise segmentatio n . 3) REFUGE18 is us ed to evaluate and compare automated algorithms f or glaucoma detection and optic disc/cup segmentation on r etinal f undus image s. Given a patient i mage, a clinician may first classify the scene as glaucoma patient or normal, as the patients may be different in the ratio b etween optic cu p and disk. Then, i t c an be determined w hether t he image contai ns t he cla sses o f optic cup and disk, while the image can b e processed through careful pixel - wise segmentation finally. In general, it is evident that human expert segmentation is accurately co nducted only i f c onsidering multi -level tas ks a nd representations jointly. Motivated by the above, w e adopt task decom position as a generalized solutio n to biomedical image segmentatio n ( first contribution ). T ask d ecomposition is to perfor m multi-level tasks and representatio ns which dec ompose a single task in to several relative sub-tasks . While mu lti -level representations are essentially important to se mantic segme ntation, we deco mpose the segmentation task to th ree sub-ta sks, i.e., to d etermine (1) pixel-wise segmentation, (2) objec t class, and (3) image sce ne. Although the three sub-tas ks are clo sely coupled , tr aditional 2 https://refuge .grand-challe nge.org/Home / FCNs are only supervised b y the pixel-wise segm entation loss, which is i nsufficient to d ecode the lo w-le vel task w hen ignori ng high-level tas k/representation. By combing multi -level tasks, the n etwork is able to comprehensively encod e the context information for the low -level seg mentation task. The seco nd contribution is the s ynchronization acro ss different sub-tasks. C oncerning the inco herence to lear n multiple sub-tasks, we use task-task context ense mble to d erive the co mmon latent space, from which t he features maps ar e forward to solve all sub-tasks jointly [13, 14] . Moreover, we propose a strong sync -regularization bet ween the segmentation and class tasks, as t he two tasks are ver y closely related with each other. Intuitively, in ROBOT18, if a class label (e.g., “kidney”) is deter mined to ha ve a certain 2D ima ge, the pi xel- wise seg mentation should b e co nsistent with some pixels labeled as “kidney”. If inconsistent outco mes are detected, one may immediately realize the failure of the jo int learn ing of the two sub-tasks. Therefore, the sync-regularization can supervise th e network to better generalize multi -level representation s. The third contribution of our w ork is to i mplement semantic segmentation to d iverse 2D/3D m edical scenario s. Specifically, we adopt a s hallow CNN as the encoder, followed by a spatial p yramid dilation module to ensemble context information for scene/clas s interpretation [15 , 16]. Instead of using a Unet -like net work [17], we adop t the PSPnet architecture [2]. W e achieve to p-tier per formance in B RATS18, REFUGE18 and ROBOT 18 challenges . In general, we summarize our major contributions in this paper as follo ws: 1) W e propose the task dec omposition strategy to ease the challenging segmentatio n task in bio medical images. 2) W e propose sync -regularization to coo rdinate the decomposed tas ks, which gains advantage from m ulti-task learning toward i mage segmentation. 3) W e build a practical framework f or diverse biomedical image semantic segmentation and successfull y apply it to three different challe nge datasets. The rest of this paper are organized as follo ws. We begin b y reviewing literature reports related to semantic segmentation in Section II. The details of our f ramework an d its components are presented in Sectio n III. To verify the effectiveness of our method, extensive experiments are co nducted and compared in Section IV. We conclude this work in Section V with extensive discussions in Section VI. II. RELATED WO RKS The FCN and its variants have d emonstrated significant power on se mantic segmentatio n. T here are also many works to highlight the ro le of conte xt information for the segmentation task. Semantic segmentation : CNN and FCN have become s tate- of -the-art m ethods in se mantic segmentation [18- 22] . In FCN , fully connected layers are i mplemented as convolutions with large rec eptive fields, and s egmentation is achieved using coarse class score maps obtained by feedfor warding th e inp ut image. The FCN netwo rk de monstrates impress ive performance. Bad rinarayanan et al. [23] present ed SegNet which is the fir st encoder-deco der a rchitecture for s emantic pixel-wise segmentation, which consists of an encoder network and a corresp onding decoder network followed by a pixel -wise classification la yer. In semantic segme ntation for medical images , Ro nneberger et al. [6] pr esent ed U-net and pro posed a training strate gy with effective data augmentation for the limited number of annotated samples. Follo wing the tremendous p erformance of th e U-net architect ure , Milletari et al. [24] proposed V-net for fully 3D image segmentation. Th eir CNN i s trained e nd- to -end o n MRI, and learns to segment the whole volume at once. There are also several attempts to address the degraded feature maps due to p ooling, including Atrous Spatial Pyramid P ooling (ASPP), encoder -decoder, and dilated convolution [ 25]. Recently, so me models such as PSPnet [2] and DeepLabV3+ [ 25] per form ASPP at various scales or apply dilated con volution in parallel. These models have bee n shown p romising capabilit y of handling semantic segmentation. I n general, a t ypical segmentation ne two rk nowadays usuall y integrates: (1) the encod er module that gradually increases the r eceptive field to capture high -level context, ( 2) the decoder module that graduall y recovers spat ial information w ith skip con nection, and (3) the dilated convolution t hat is e ffective to context ensemble. However, such a so phisticated network often needs large amount of d ata to optimize, w hich is a critical challenge in bio medical image segmentation task. Deep multi-task learning: Dee p multi-task learning aims at improving ge neralization ca pability by leveragi ng different domain-specific information [2 6] . T here are many works of multi-task lear ning to ward image seg mentation and detec tion. Dai et al. [27] proposed Mul ti -task Network Ca scades for instance-aware se mantic segmentation, which is designed as cascaded structure to share the convolutional features. He et al. [28] proposed Mask R-CNN which jointly opti mizes three tasks, i.e., detection, segmentatio n and classification , and outperformed sin gle-task competitors. This ap proach efficiently detected objects in an image while si multaneously generating a high-quality segmentation mask for each instance. In the field o f medical image multi-task learning , Ravi et al. [29] proposed a multi-task transfer learning DCNN with the aim of translating the ‘kno wledge’ le arned fro m non - medical i mages to medical segmentation tasks b y simultaneo usly learning auxiliary ta sks. Chen et al. [30] proposed a net work wh ich contains multi-level context features fro m t he hier archical architecture and explores auxiliary supervision for accurate gland segmentatio n. Whe n incorp orated w it h multi -task regularization d uring the training, the discriminative capabilit y of latent features can be further i mproved. However, none of the methods m entioned above has co mbined multi-task learning with the idea o f deco mposing the se gmentation tas k into sub - tasks and synchroniz ing them for better segmentation. Fig. 3 . Ty pical segmentation results for ROBOT 18 by usin g d iffe rent m ethods. From left to right are the origin al ima ge, grou nd tr uth, Unet result , PSPNet result, EncNet result, and the proposed method ’s result. Using deep multi-tas k learning with d ecomposition and sync- regularization could increase the generalizat ion cap ability of the FCN model, and thus reduce the difficulty in semantic segmentation [3]. III. PROPOSED MET HOD We propose the task deco mposition framework as in Fig. 2 . Given an input image (2D or 3D), we first use a dense convolutional encoder to extract feature maps. Because of the diversity of the inp ut 2D/3D data, we design specific encod er for each of the three challenges in this paper. Then, w e feed the extracted feat ures to the task -task context ensemble module. The context ensemble module contains multi-scale dilat ed convolution, so the recep tive fields ar e enlarged along the paths to co mbine feat ures of different scales by d ifferent d ilated rates . Moreover, the th ree parallel context ensemble modules are generated a s tas k-task co ntext ensemble module and each of module ar e co nnected by two branches which we ca lled latent space. Fin ally, the net work is deco mposed from the latent space into th ree branches, co rresponding to (1) the segmentation task , (2) the class task, and (3) the scene task. T he decoders are trained for each d ecomposed ta sk, incl uding up sampling for the segmentation tas k, and also g lobal average poo ling and fully - connected la yers for the class/scene tasks. No te that the three decomposed tas ks share the same late nt space for dec oding. A. Task Decompo sition and Sync -Regularization We aim to exploit m ulti-lev el rep resentations for semantic segmentation. While state- of -the-art net works for segmentation often require tons o f training data, it is possible to rely on t he mutual depende ncy o f the d ecomposed tasks to ease the parameter sear ching in FCN, especiall y concer ning t he li mited numbers of medical i mages for tr aining . W e particularly decompose the sema ntic se gmentation ta sk into (1) p ixel-wise image seg mentation, (2) p rediction o f the object classes within the image, and (3 ) classification of the sce ne the image belonging to. The three tasks are designed to optimize their individual loss functions. W e utilize a hybrid loss to supervise the lo w- level segmentation ta sk: Ω (1) is the cross-e ntropy between the se gmentation ground truth and the estimated for the image in the training set . divides the i ntersection area over the union bet ween and . For b oth the high-level class/scene tasks, we adop t the same loss design as: Ω Ω (2) is the binary cross -entropy, and are the ground truth and the estimated class labels, respectively, and and are the labels for the scene task. B. Task-Task Context E nsemble The three tasks are trained to s hare the s ame latent space, while the tas k-task interaction is atta ined b y co ntext ensemble in our framework. The size of the receptive field, which is critical to explore context in formation, can o ften b e too lo w [31] . To this end, we are inspired by Dlinknet [1 6] and cascade the dilat ed convolution for the conte xt ensemble module as in in Fig. 2 . Specifically, b y couplin g each pair of the d ecomposed tas ks, the receptive fields enlarge their s izes, throu gh the stacked dilated convolution with the dila ted r ates 63, 31, 15, 3 , and 1 , respectively , in the outp ut o f the module . The outputs of the three task-task CE m odules are f urther parallel ed and cascaded , which are resulted in the latent space. I n detail, the three parall el context ense mble mod ules are generated and each of module are connected by tw o branches which w e called th is structure is latent space. Note that o ur con text ensemble module can k eep the spatial resol ution/size of the feature maps, which significantly benefits seg mentation accurac y. C. Sync -Regularization As the network is decomposed into multiple branches for decoding, it is necessary to balance different tasks when generalizing the latent space. In particular, the class task can be regarded as a pr ojection of the segmentation task. For example, the class task can determine wh ether “kidney” e xists in a ROBOT18 image. One may also infer fro m the outpu t of the segmentation tas k, and identify whether certain pixels are wrongly lab eled as “kidne y”. By examining the deviatio n between the t wo tasks, we prop ose a novel sync-regularizati on strategy which has demonstrated po werful capability of learning the late nt space jointly for better se gmentation performance . Specifically, we estimate the volume of eac h segmented label from the segmentation task’s output, w hich is then converted to a boolean value (the pixel number of the spec ific class in segmentation map greater than 0 is set to 1). The vector of the boolean values, , is then compared w i th the output of the clas s task, which results in the following loss functio n: (3) Here, is the binary cross-entropy. The sync-regularization provides feedbacks in bac k-pr opagation through t he branch for the class task, and t hus e nforces cons istency chec k to optim ize the rep resentations in the latent space. Note that the sce ne task is ex cluded from sync-regulariza tion, as its association with the segmentation ta sk is relativel y loose . It’s difficult to decompose segmentation map or classification vector into scene vector. E.g., converti ng seg mentation maps of brai n tu mor to HGG/LGG classes i s hard to implement. In general, the total loss in our fra mework is (4) where , , and are weights for tasks , , and , and also the hyper-para meters tuned in t he experi ments. We tu ned th e parameters in sequence, and the set of val ue is 0 .0 , 0.2, 0.4, 0.8, 1 .0 . D. Encoder and Decoder We design different encoders for 2D and 3D scenarios. In ROBOT18 and REFUGE18 that are 2D image datasets, we use VGG16 [32] pre-trained o n ImageNet [33] as the encoders following LinkNet [34] setting. While for BRATS18 of 3D images, we start from Wang et al. [15] by using multiple layers of anisotro pic and dilated convolution filters for encod er and decoder architectures. Many m edical image segmentation m ethods prefer to us e the encoder-decod er structure an d argue that it pro vides detai led boundary infor mation to the o bject under consideration [35] . Here w e have found that dilated convolution as in t he PSP net - like arc hitecture ca n also extract multi-scale dense feature maps effectively an d can achieve high p erformance in image segmentation. It is thus noted that connecting lo w- level and high-level layers w ith skip con nection will possibl y increase t he difficulty o f op timization in biomedical image segmentati on. Particularly, for the decod ers to the class/scene tasks, we fi rst feed the feature maps throug h global average pooling [36] , and then a fully con nected la yer deco des the desired outputs (c.f. Fig. 2 ) . E. Training Proced ure It is known that th e task-task interaction incurs additional difficulty to train in multi -task lear ning. T o this end, we implement the task dec omposition framework in a t hree-s tep training procedure: 1) W e train the segmentation a nd class tasks together. 2) W e utilize the early estimate d parameters as initialization and refine the segmentati on/class tasks w ith sync - regularization enforced . 3) W e add in the sce ne task and refine all lo ss functions jointly. Consequently, we ob tain the fine -tuned FCN model tha t produces superior p erformance in t he provided medical ima ge datasets, while our training procedure tends to b e robust in all experiments. IV. EXPERIMENTAL RESULTS In this section, w e firs t p rovide i mplementation details for o ur task deco mposition framework. T hen, we eval uate i t on three diverse challenge data sets, includin g ROBOT1 8, BRATS18, and RUFUGE18 . A. Implementatio n Details Our implementation is based o n Pytorch [37 ] . Regarding the hyper-para meters, the basic learning rate is set to 0 .0001. For multi-task lear ning, the lear ning rate dec reases gradually, i. e., 1× 10 -4 , 5× 10 -5 , 2× 10 -5 after ev ery 50 iterations . The momentum and weight deca y are set to 0.9 a nd 0.0001, r espectively . In ROBOT18 , we apply vertical/horizo ntal flip, RGB shift, random brightness (limit: 0.9 -1.1) and random contrast (limit: 0.9-1.1) to augment the training data. For the validation dataset, no augmentation is adopted. Note that, for ROBOT 18, there are 12 classes (i.e., sha ft, intesti ne, wrist, thread, pr obe, kidney, covered, clasper , suction, clamps, needle a nd background) and Fig. 4 . Ty p ical segmentation resu lts for BRATS18 by using diff erent methods. Fr om le ft to right are for t he o riginal image, the ground truth, and results by Unet and our proposed method, respectively. 3 scenes (i.e., prep aring, pee ling and incising). We use 256 neurons in the hidden la yers for both class and scene tasks ( c.f. Fig. 2 ) . The seco nd dataset BRAT S18 is co mmonly used to validate brain tumor s egmentation. For 3D medical im ages, the receptive field, model complexity and memory consumption of the net work sho uld be balanc ed. As a trad e-off, we ado pt t he anisotropic setting that sets lar ge receptive field in 2D slice but relatively small in the direction per pendicular to the slice. All images ar e thus cropped to the size of 1 44× 144× 19. For data augmentation, we on ly utilize r andom flip in three directions during trai ning. There are 4 classes ( i.e., ET , NCR/NET, ED and background) and 2 scenes (i.e., low -grade and high-grade gliomas) that are identified for BRATS18. We also use 128 neurons in t he hidden la yers, w hich is the same with R OBOT18. The third dataset REFUGE18 is used to validate automated segmentation of optic disc/cup . We utilize same configuration of learnin g rate, data au gmentation and trai ning strate gies with ROBOT18 . There are 3 classes (i.e ., optic disk, optic cup and background) and 2 scenes (i.e., glaucoma, normal). T here ar e also 256 neuro ns used in the hidden la yers. B. ROBOT: Robo tic Scene Segmentation Challenge There are 85 teams participating into the ROBOT 18 challenge . Table 1 . Comparisons o f single/multi-task learning, as well as the components o f the proposed method, in solvin g RO BOT18 segmentation challenge. ( “ Base+Class+Syn+Scene ” indicates the proposed method.) IoU: % Dice: % Single Task of Segmentation Only ResNet34+Unet 74.28 78.31 ResNet50+Unet 51.33 55.27 VGG16+Unet 75.82 79.17 VGG16+CE+Unet 75.90 79.23 VGG16+CE+ASPP ( Base ) 77.81 81.40 Multiple Tasks of Segmentation , Class , and Scene Base+Class (using Training Step 1) 80.03 83.73 Base+Class+Sync (Steps 1- 2) 81.99 85.56 Base+Class+Sync+Scene (Steps 1- 3) 82.08 85.70 Table 2 . Comparisons of th e prop osed method and other state- of -the- art methods in solving ROBOT18 segmentation challenge. IoU: % Dice: % PSPnet+ResNet34 69.86 73.06 PSPnet+ResNet50 65.59 68.87 Encnet+ResNet34 67.85 71.07 Encnet+ResNet50 67.02 70.09 Proposed ( Base+Class+Sync+Scene ) 82.08 85.70 The challen ge data has 2,2 35 training i mages, where the occurrence of certain cla ss can be low. At the end of the challenge, 1,00 0 unseen i mages are released for test. T he challenge is ranked on the m ean IoU m etric, which i s computed per class and then averaged for the score of the entire test image . We can also compute Dice coe fficient ( Dice), a co mmonly used metric, to quantitati vely assess the segmentation results. To validate th e d esign of our network, we co mpare seve ral different sett ings and repo rt the results in Table 1 . First, we adopts VGG16 as the encoder while ASPP as th e decoder (“ VGG16+CE+ASPP ”), while one may also c hoose o ther network architectures. The IoU/Dice scores of o ur implementation are 77.8 1% and 81.40%, both of which are higher than the alternatives. Particularly, we notice that the context ense mble (designated as “CE” in the table) module is effective even though only th e segmentation task is considered. It is worth n oting that, alt hough the module is d esigned to couple a pair of decomposed tasks, w e can also apply it to the case when only the single segmentation task is co nsidered. Second, with the ne twork ar chitecture (“ Base ”) validated in the single -task learning, we further verify the co ntribution of the p roposed task decompositio n fra mework. Since t here are three steps in o ur trainin g proced ure, we gradually add i n the class/scene tasks and e nforce the sync -regularization step b y step. T he experimental r esults in the botto m part o f Table 1 show that, with task deco mposition, the multi-tas k solution to semantic segmentation outperforms the single -task solution, implying the effectiveness of the decomposed class/scene tasks toward image segme ntation. Moreo ver, the s ync-regularization also y ields significant i mprovement for both IoU and Di ce. Therefore, we concl ude that the p roposed task d ecomposition and s ync-regularization are ef fective to t he seg mentation ta sk (IoU: 82.08%; Dice : 85.70%). Fig. 5 . Visual comparison on R EFUGE18. Our proposed netw ork achieves m ore accurate a nd detailed results. Fro m lef t to right is original image, ground truth, and results by Unet and our proposed method, respectively. Finally, we compare our proposed method w ith o ther state - of - the-art algorithms in Table 2 . Considering the small size of the dataset, we adopt light-weighted encoders for PSP net and Encnet. The resu lts show that the proposed metho d (“ Base+Class+Syn+Scene ”) outperforms all m ethods under comparison in the validatio n set. Moreover, our method has demonstrated top -tier performance in t he on -site testing set i n ROBOT 18 (rank second, IoU=61 %, compared to 62% of the challenge winner ). Note that the on -site test data IoU score is different with what is s hown in Table 2 , since the i mages in the on -site testing s et and the validation set are not co herent with each oth er. We have also pro vided visual inspection of t ypical se gmentation resu lts of ROBOT 18 in Fig. 3 , where our method clearl y performs better than the alternat ives und er consideration. C. BRATS: B rain Tumor Segmentation Challenge BRATS is focusing o n the evaluation o f state - of -the-art methods for the segmentatio n o f brain t umors i n multi -mod al magnetic reso nance (M R) sca ns [ 38]. There are more than 20 0 teams participating in the BRATS18. In our w ork there are 2 85 subjects used for tr aining, ea ch of which co mes with a 240× 240× 155 sized 3D image. We also use the data of 66 subjects for o n-site validation. Slightl y different from ROBOT18 , the B RATS18 challenge uses Dice a nd Hausdorff distance (HD) as the perfor mance metrics. All comparisons are conducted o n the o n-site valid ation data, while the resul ts are submitted directly through the official website 3 . The org anizers 3 https://www .med.u penn.edu/sbia/bra ts2018 Table 3 . Comparisons o f single/multi-task learning, as well as the components of the proposed method, in solving BRATS18 segmentation challenge. ( “ Base+Class+Syn+Scene ” indicates th e proposed method.) Dice: % HD: mm Single Task of Segmentation Only Wnet+Unet 79.67 17.09 Wnet +CE+Unet 79.69 16.43 Wnet +CE+ASPP ( Base ) 80.15 15.75 Multiple Tasks of Segmentation , Class , and Scene Base+Class (using Training Step 1) 80.41 10.30 Base+Class+Sync (Steps 1- 2) 80.87 10.14 Base+Class+Sync+Scene (Steps 1- 3) 80.88 9.74 Table 4 . Comparisons of th e prop osed method and oth er state - of -the- art methods in solving BRATS18 segmentation challenge. Dice: % HD: mm Unet 79.45 12.76 Deepmedic 80.42 10.54 GTNet 82.29 11.30 Proposed ( Base+Class+Sync+Scene ) 80.88 9.74 Table 5 . Comparisons of o ur result and other top-rank ed results submitted to the on -site validation of BR ATS18 segmentation challenge. 4 Team Dice: % HD: mm ET WT TC ET WT TC NVDL 82.5 91.2 87.0 4.0 4.5 6.8 MIC 80.9 91.3 86.3 2.4 4.3 6.5 BIGS2 80.5 91.0 85.1 2.8 4.8 7.5 Proposed 83.2 91.5 88.3 2.9 3.9 7.7 return the ranks of all participants, as our submission ranks f irst in B RATS18. In o rder to validate th e design of our n etwork, we compare several different setti ngs and report respective results in Table 3 , w hich is similar to early experiments in ROBOT18. Our implementation adopts state- of -the-art “ Wnet ” as th e encoder for 3D medical image segme ntation. Me anwhile, we use ASPP as the decoder. T herefore, w e term our implementation as “ Wnet+CE +ASPP ”. The Dice/HD scores of our method are 80.15% and 15 .75 mm on the on-site validati on se t, bo th of which are better than other cases (c.f. Table 3 , the single-task setting). Particula rly, we notice that th e cont ext ensemble module is effective even though o nly t he single segmentation task is cons idered. We also conduct the exper iment o f sema tic se gmentation using the proposed task d ecomposition strategy, a nd co mpare its perfor mance with t he alternati ve confi gurations, including “ Base+Class ”, “ Base+Class+Syn c ”, and “ Base+Class+Sync+S cene ” in Table 3 . T he findings in Ta ble 3 are also si milar with those in Table1 : using the whole three steps as the multi-ta sk sol ution ca n f urther i mprove the segmentation perfor mance; t he propo sed task decomposition and sync -regularization are also considered as b eneficial to the segmentation task for the BRATS18 d ataset (Dice: 8 0.88 %; HD: 9. 74 mm). Besides, w e co mpare o ur proposed method with other state- of -the-art algorithms in Table 4 . For sing le state- of - the-art 4 https://www . cbica.upenn.edu/BraT S18/ Table 6 . Comparisons o f single/multi-task learning, as well as the components of the proposed method, in solving REF UGE18 segmentation challenge. ( “ Base+Class+Syn+Scene ” indicates the proposed method.) IoU(%) Dice(%) Single Task of Segmentation Only ResNet34+Unet 75.04 84.78 ResNet50+Unet 76.85 85.98 VGG16+Unet 81.26 89.13 VGG16+CE+Unet 82.47 90.43 VGG16+CE+ASPP ( Base ) 82.67 90.72 Multiple Tasks of Segmentation , Class , and Scene Base+Class (using Training Step 1) 82.91 91.02 Base+Class+Sync (Steps 1- 2) 83.44 91.21 Base+Class+Sync+Scene (Steps 1- 3) 83.49 91.29 Table 7 . Comparisons of th e pro posed method and oth er state - of -the- art methods in solving REFUGE18 segmentation challenge. IoU: % Dice: % PSPnet+ResNet34 75.52 84.99 PSPnet+ResNet50 72.96 82.62 Encnet+ResNet34 75.57 85.31 Encnet+ResNet50 73.02 83.46 Proposed ( Base+Class+Sync+Scene ) 83.49 91.29 models like 3D Unet [39] and Deepmedic [40], our method could outperform these metho ds on bo th evaluation metrics. Comparing our method with the winner of the BRATS challenge in 2017 (GTNet) [15 ] , w e al so ha ve a better performance o n HD score while worse Dice. Note that GT Net requires to train 9 dif ferent segmentation models f or th e coarse- to -fine ensemble. Moreover, our m ethod has obtained the to p -rank ed score o n the o n-site validatio n set in B RATS18 as in Table 5 . Our final submission achieves 8 3.2, 91.5, 8 8.3 for Dice (%) and 2 .9, 3.9, 7.7 f or HD (mm) o n t hree foreground classes (i.e., ET, WT, and TC). We have also provided visual inspectio n of t he segmentation results of BR ATS18 in Fig. 4 . D. REFUGE: Retin al Fundus Glaucoma Chal lenge The number of tea ms participating in the REFUGE18 is more than 400 . Here we use 400 training images in the REFUGE 18 challenge dataset. A t th e end of the challenge, 400 un seen images are released for test. T he challenge is ranked on t he mean Dice metric, which is co mputed per class and then averaged for the sco re of the entire test i mage. We ca n also compute Intersect ion over Union (IoU) to quantitatively a ssess the segmentation results. We also conduct the e xperiments of performance evaluatio n for the REFUGE1 8 dataset, and com pare the res ults using several differe nt settings in single -task learning first. Table 6 verifies the Io U and th e Dice m easurements for the experimental results. While our implementation adopts VGG16 as the encoder and ASPP as the deco der (“ VGG16+CE+ASPP ”), th e IoU/Dice scores of o ur method are 82.67% and 90.72%, bo th of which ar e higher tha n o ther choices. The experi mental results for the propo sed task decomposition f ramework are also sh own in Table 6 , where we also evaluate the validit y of the three training steps for the REFUGE18 dataset. Sim ilar with what are previously s hown in Table 1 and Table 3 , the segmentation performance using all three steps outper forms the cases when the class/scene ta sks and the sync-re gularization co mponent ar e not fully app lied. The IoU and the Dice scor es for o ur method (“ Base+Class+Sync+Scene ”) reach 83.49% and 91.29%, respectively. Finally, we compare our proposed method with other state - of -the-art algorithms in Tab le 7 . Here we adopt light -weighted encoders for PSPnet and E ncnet due to the limited size of the training datasets. The results show that the proposed method outperforms al l methods under comparison in th e validation s et. Moreover, the on-site score of our method for REFUGE18 can be mostly comparable with t he winner score of the c hallenge (IoU=91%, compared to 92% of the win ner). We have also provided visual inspectio n of the seg mentation res ults o f REFUGE 18 in Fig. 5 . V. D ISCUSSION AND C ONCLUSION As a summary, w e ha ve pro posed an effective task decomposition fra mework for semantic segmentatio n of di verse biomedical i mages i n this work . Specificall y, we dec ompose the ver y challe nging semantic segmentation task to seek for helps fro m auxiliar y class/scene tasks. T he d ecomposed tasks are associated w it h low-level to high-level r epresentations a nd reduce the complexity to solve image segmentatio n. Moreo ver, in addition to co ntext ense mble in the latent space, w e pr opose sync-regularization to p enalize the deviat ion b etween differ ent tasks and to coordinate multi-task learning for the sake of semantic segmentation. We hav e conducted comprehensive experiments on three ver y diverse yet p opular medical im age datasets. Our results are curr ently top -ranking in all three challenges. In th is work, t he entire training phase o f o ur method take s about 4 , 7, 2 hours on a single NVI DIA Titan X GPU for ROBOT18 , B RATS18 an d REFUGE 18 dataset, resp ectively . It is worth noti ng t hat it costs only 0.1s (2s) to gen erate the final segmentation map for one 2D ( 3D ) i mage in the te sting stage . While the accurac y of the CN N for m edical image segmentation is mostl y co mparable to (or better than) state - of -the-art algorithms, the runtime is sho wn to outperform other mu lti-task segmentation app roach (e.g., Mask R-CNN [2 8]) for medical image seg mentation. T his w ill b e particularly b eneficial in large clinical en vironments where hundreds or so metimes thousan ds of people are screened every day. There are so me limitations for our propo sed m ethod . Our proposed encoder and decoder try to find a trad e-off bet ween depth and computational feasibility. W hile increasing depth addresses lar ger semantic regions d uring pr ocessing, contraction ele ments such as max-pooling red uces the number of parameters but loses information ab out sp ecific locations. In contrast, dilated convolutio n op eration may lead to increased parameters but keep more i nformation in d etail . Although the proposed method has shown its pro mising result s in the experiments, it would be interesting ho w to obtain an optimal architecture which could keep necessary information but c ost less in computation. Moreover, in this w ork our s ync- regularization is a sin gle-directional loss which only matc h es the segme ntation task with classi fication one. However, the reversible loss is a m o re reasonab le w ay to measure t he variance between the t wo tasks considering classificat ion task may misguide the optimization of segmentation task. Finally , we only explore the b iomedical image datase ts which only have limited number of images in o ur work. It w o uld be valuable to explore this f urther and see ho w t he method performs on more traditional natural i mages, par ticularly when varyi ng the amount of training data. In future work, we will further i mprove method fro m the following two aspects. 1) W e will sim plify the sync- regularization as the single de stination loss in t he network. We need to explor e a feasible w ay to build a reversible loss between segmentation a nd classi fication tas ks i n our propo sed network. 2) Our prop osed innovation modules, i ncluding task decomposition a nd s ynchronization , can be further explor ed especially in a large a mount of training dataset. VI. R EFERENCE [1] J. Long, E. Shelhamer, and T. Darrell , "Fully convolutional netw orks for semantic segmentation," in Proceedings of the IEEE conference on computer visi on and pattern re cognition , 2015, p p. 3431-3440. [2] H. Zhao, J. Shi, X. Qi, X. Wang, and J. Jia, "Pyramid scene p arsing networ k," in IEEE Conf. on Computer Vision and Pattern Recognition (CV PR) , 2017, pp. 2881-2890. [3] H. Zhang et al. , "Context encoding for semantic segmentation," in The IEEE Conference on Computer Vision and Pattern Recognition (CVPR) , 2018. [4] L. -C. C hen, G . Papandre ou, I. Kokkinos, K. Mu rphy, and A. L . Yuille , "Deeplab: Semantic i mage segme ntation with deep convolutional nets, atrous convolution, and fully conn ected crfs," IEEE transactions on pattern analysis and machine intelligence, vol. 40, no. 4 , pp. 8 34-848, 2018. [5] Z. Zhang, X. Zha ng, C. Peng, D. Cheng, and J. Sun, "ExFuse : Enhancing Feature Fusion for Semanti c Segme ntation," a rXiv preprint arXiv:1804.0382 1, 2018. [6] O. Ronneber ger, P. Fischer, and T . Br ox, "U-net: Convolutio nal netwo rks for biomedical im age segme ntation," in Internationa l Conference on Medical image computing and computer-assisted intervention , 2015, p p. 234-241: Springe r. [7] H. Noh, S. Ho ng, and B. Han, "Learning deconvolution networ k f or semantic segmentation," in Proceedings of the IEEE internati onal conference on c omputer vision , 2015, pp. 1 520 -1528. [8] V. Badrinaray anan, A. Ke ndall, and R. Cipolla, "Segnet: A deep convolutional encoder-decoder architecture for image segmentation," arXiv preprint arXiv:1511.00 561, 2015. [9] K. He , X. Zhang, S. Ren, and J. Sun, "Deep residual learning for i ma ge recognition," in Proceedings of t he IEEE conference on computer vision and pattern rec ognition , 2016, pp. 770-778. [10] F. Yu and V. Koltun, "Multi-scale context aggregation by dilated convolutions," arXi v preprint arXi v:1511.07122, 2015. [11] S. Bakas et al . , "Advancing the cancer genome atlas glioma MRI collections with expe rt seg mentation labels and radiomic fe atures," Scientific data, vol . 4, p. 17011 7, 2017. [12] W. Xiong, C .-K. C hui, and S. -H. On g, "Focus, Segment and Erase: A n Efficient Networ k for Multi-Label Brain Tumor Segmentation," Ratio, vol. 5, no. 13.4, p . 16.8. [13] S. Ruder, "An overv iew of multi-task learning in d eep neural networks," arXiv preprint arXiv:1706.05 098, 2017. [14] Z. Chen, V. Badri narayanan, C.-Y. Lee, and A . Rabinovich, " GradNorm: Gradient Normalization for Adaptive Loss Balancing in Deep Multitask Networks," arXiv preprint arX iv:1711.0225 7, 2017. [15] G. W ang, W. L i, S. Oursel in, and T. V ercautere n, "Automatic brain tumor segme ntation using cascade d anisotro p ic co nvolutional ne ural ne tworks," in International MICCAI Brainlesio n Workshop , 2017, pp. 178-190: Springer. [16] L. Zhou, C. Zhang, and M. Wu, "D-LinkNet: LinkNet with Pretrained Encoder and Dilated Convolution for High Resolution Satellite Imagery Road Extraction," i n Proceedings of the IEEE Conference on Compu ter Vision and Patt ern Recognition W orkshops , 2018, pp. 182-186. [17] O. R onneberg er, P. Fischer, and T. Brox, "U-Net: Convolutional Networks for Biomedical Image Segme ntation," in medical image computing and computer assiste d intervention , 2015, p p. 234-241. [18] D. Shen, G. Wu, and H.-I . Suk, "De ep learning in medical image analysis," Annual review o f biomedical engi neering, vo l. 19, p p. 221-248, 2017. [19] H. Chen, Q. Dou, L. Yu, J. Qin, and P.-A. Heng, "VoxResNet: Deep voxelw ise residual networ ks for brain segmentation from 3D MR images," NeuroIm age, 2017. [20] A. V. Dalca, J. Guttag, and M. R. Sabuncu, "Anatomical Priors in Convolutional Netw orks for Unsupervised Biomedical Segme ntation," in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition , 2018, pp. 9290-9299. [21] X. Xu et al. , "Quantization of f ully convolutional networks for accurate biomedical im age seg mentation," Preprint at https://arxiv . org/abs/1803.04 907, 2018. [22] K. -L. Tseng, Y.-L . Lin, W . Hsu, a nd C.-Y. Huang, "Joi nt sequence learning and cross-modality convolution for 3d biomedical segme ntation," in Com puter Vision and Pattern Recognition (CVPR), 2017 IEEE Conf erence on , 2017, pp. 3739- 37 46: IEEE. [23] V. Badrinaray anan, A. Kendall, R. J. I. t. o. p . a. Cipolla, and m. intellige nce, "Segne t: A de ep convol utional encoder-decoder architecture for image se gmentation," vol . 39, no. 12, pp. 248 1-2495, 2017. [24] F. Mi lle tari, N. Navab, and S. -A. Ahmadi, "V-net: Fully convolutional neural networks for volumetric medical image segmentation," in 2016 Fourth I n ternationa l Conference on 3D Vis ion (3DV) , 2 016, pp. 565-571: IEEE. [25] L. -C. Chen, Y. Z hu, G. Papandreo u, F. Schroff , and H. Adam, "Enco der- decoder with atrous separable convolution for semantic image segme ntation," arXiv preprint arXiv:1802.02611, 2018. [26] Y. Zha ng a nd Q . J . a . p. a. Y ang, "A surv ey on multi-task l earning," 2017. [27] J. Dai, K. He, and J. Sun, "Instance-aw are s emantic segmentation via multi-task netw ork casca des," in Proceedi ngs of the IEE E Confere nce on Computer Visi on and Pattern Recognition , 2016, p p. 3150-3158. [28] K. He, G. Gkioxari, P. Dollá r, and R. Girshick, "Mask r -cnn," in Computer Visi on (ICCV) , 2017 IEEE International C onference o n , 2017, pp. 2980-2988: I EEE. [29] C. Zu et al. , "L abel-aligned multi-task feature learning for mu ltimodal classification of Alzheime r’s disease and mild cognitive impairment," vol. 10, no. 4, pp. 1 148-1159, 2016. [30] H. C hen, X. Qi, L . Yu, Q. Dou, J. Qin, and P.-A. J. M. i. a. Heng, " DCAN: Deep contour-awar e netwo rks for object instance seg mentation from histology images," vol . 36, pp. 135-146, 2017. [31] W. Luo, Y. Li, R . Urtasu n, an d R. Z emel, "Understanding the effective receptive field in deep convolutional neural n etwo rks," in Advances in neural informa tion processing s ystems , 2016, pp. 489 8-4906. [32] C. Szegedy et al. , "G oing deeper w ith convolutions," 2015: Cvpr. [33] J. Deng, W. Dong, R. Socher , L.-J. Li, K. Li, and L. Fei -Fei, "Imagenet: A large-scale hierarchical image database," in Computer Vision and Pattern Recognition, 2009. CVPR 2009. IEEE Conference on , 2009, pp. 248-255: IEEE. [34] A. Chaurasia and E. Culurciel lo, "Linknet: Ex ploiting encode r represe ntations for efficient se mantic segmentation," in 2017 IEEE Visual Communications and Image Pro cessing (VCIP) , 2017, pp. 1-4: I EEE. [35] G. Lin, A. Milan, C. Shen, and I. D. Reid, "RefineNet: Multi-path Refinement Networks for High-Resolution Semantic Segmentation," in Cvpr , 2017, vo l. 1, no. 2, p. 5. [36] M. Lin, Q. Chen, and S. Yan, "Network in n etwo rk," ar Xiv p reprint arXiv:1312.4400 , 2013. [37] A. Paszke et al. , "Automa tic differe ntiation in py torch," 2017. [38] B. H. Menze et al. , "The multimoda l brain tumor image segmentation benchmark ( BRATS) ," IEEE transactions on medical imaging, vol. 34, no. 10, p. 1993, 2015. [39] F. Mi lle tari, N. Navab, and S. -A. Ahmadi, "V-net: Fully convolutional neural netw orks for vol umetric medical i mage segmentatio n," in 3D Vision (3DV), 2016 Fourth International Conference on , 2016, pp. 565- 571: I EEE. 10 [40] K. Kamnitsas et al. , "DeepMedic for brain tumor segme ntation," in International Workshop on Brainlesion: Glioma, Multiple S clerosis, Stroke and Trauma tic Brain Inj uries , 2016, pp. 1 38-149: Springer .
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment