dna2vec: Consistent vector representations of variable-length k-mers
One of the ubiquitous representation of long DNA sequence is dividing it into shorter k-mer components. Unfortunately, the straightforward vector encoding of k-mer as a one-hot vector is vulnerable to the curse of dimensionality. Worse yet, the dista…
Authors: Patrick Ng
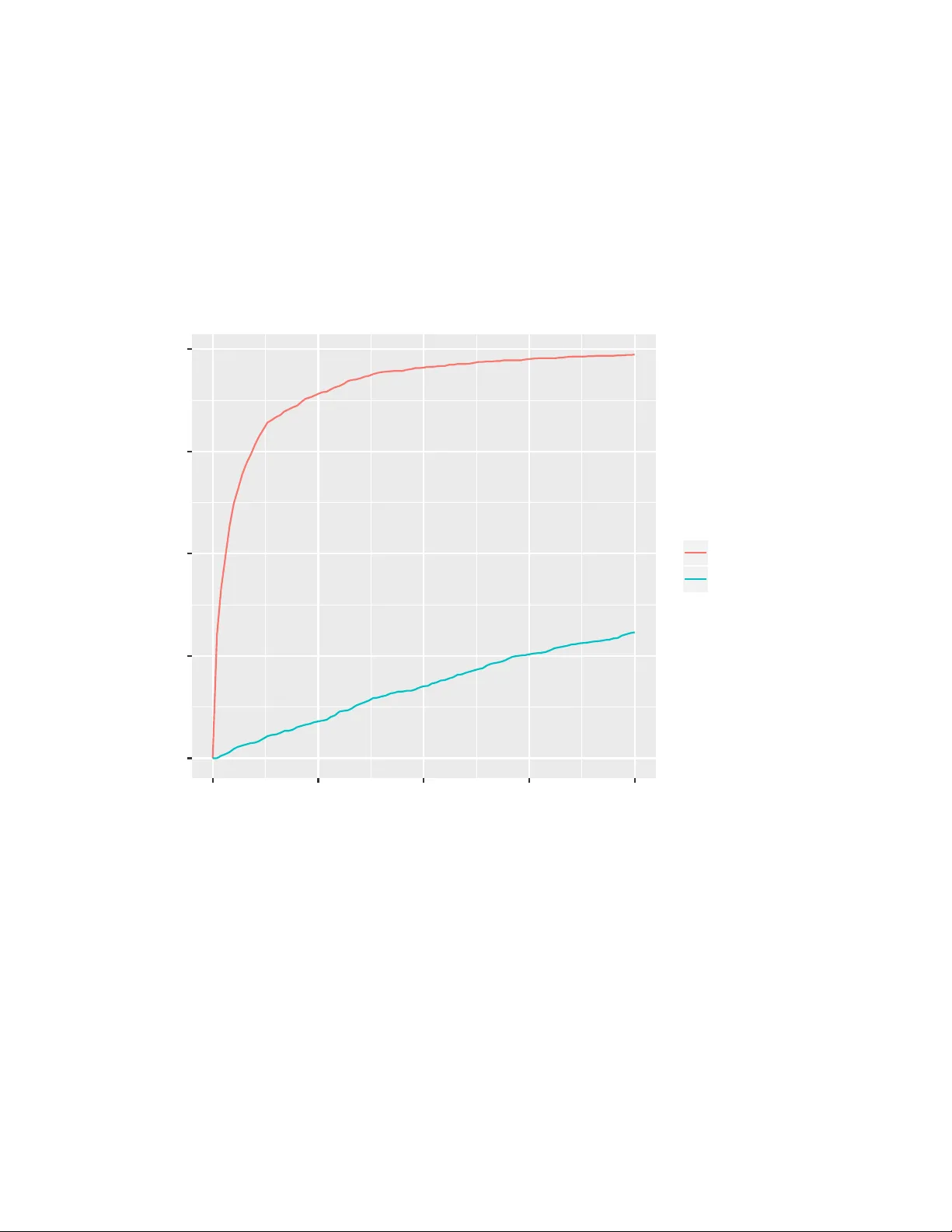
dna2v ec: Consisten t v ector represen tations of v ariable-length k-mers P atrick Ng ppn3@cs.cornell.edu Abstract One of the ubiquitous represen tation of long DNA sequence is dividing it in to shorter k-mer comp onen ts. Unfortunately , the straightforw ard vector enco ding of k-mer as a one-hot vector is vulnerable to the curse of dimensionalit y . W orse y et, the distance b etw een any pair of one-hot v ectors is equidistant. This is particularly problematic when applying the latest machine learning algorithms to solve problems in biological sequence analysis. In this pap er, we propose a nov el metho d to train distributed representations of v ariable-length k-mers. Our metho d is based on the popular word embedding mo del wor d2ve c , which is trained on a shallo w tw o-la y er neural netw ork. Our exp eriments pro vide evidence that the summing of dna2vec vectors is akin to nucleotides concatenation. W e also demonstrate that there is correlation b et w een Needleman-W unsc h similarity score and cosine similarity of dna2vec vectors. 1 In tro duction The usage of k-mer representation has b een a p opular approac h in analyzing long sequence of DNA fragments. The k-mer represen tation is simple to understand and compute. Unfortunately , its straigh tforward vector enco ding as a one-hot v ector (i.e. bit vector that consists of all zeros except for a single dimension) is vulnerable to curse of dimensionality . Sp ecifically , its one-hot v ector has dimension exp onential to the length of k . F or example, an 8-mer needs a bit v ector of dimension 4 8 = 65536 . This is problematic when applying the latest machine learning algorithms to solv e problems in biological sequence analysis, due to the fact that most of these to ols prefer lo wer-dimensional contin uous vectors as input (Suyk ens and V andewalle, 1999; Angerm ueller et al., 2016; T urian et al., 2010). W orse yet, the distance b etw een an y arbitrary pair of one-hot v ectors is equidistant, ev en though ATGGC should be closer to ATGGG than CACGA . 1.1 W ord em b eddings The Natural Language Processing (NLP) researc h communit y has a long tradition of using bag-of-words with one-hot vector, where its dimension is equal to the vocabulary size. Recently , there has b een an explosion of using wor d emb e ddings as inputs to machine learning algorithms, esp ecially in the deep learning communit y (Mik olo v et al., 2013b; LeCun et al., 2015; Bengio et al., 2013). W ord embeddings are v ectors of real num b ers that are distributed represen tations of w ords. A p opular training technique for w ord embeddings, w ord2vec (Mikolo v et al., 2013a), consists of using a 2-la y er neural netw ork that is trained on the current word and its surrounding con text words (see Section 2.3). This reconstruction of context of words is lo osely inspired by the linguistic concept of distributional hyp othesis , which states that w ords that app ear in the same context hav e similar meaning (Harris, 1954). Deep learning algorithms applied with w ord embeddings hav e had dramatic impro vemen ts in the areas of mac hine translation (Sutskev er et al., 2014; Bahdanau et al., 2014; Cho et al., 2014), summarization (Chopra et al., 2016), sentimen t analysis (Kim, 2014; Dos Santos and Gatti, 2014) and image captioning (Viny als et al., 2015). One of the most fascinating prop erties of word2v ec is that its v ector arithmetic can solve semantic and linguistic analogies (Mikolo v et al., 2013c,a). They show ed that v ec ( k ing ) − vec ( man ) + v ec ( w oman ) ≈ v ec ( q ueen ) . In particular, the analogy task man:king :: woman:??? is interpreted as finding a wor d w suc h that v ec ( k ing ) − vec ( man ) + v ec ( w oman ) is closest to v ec ( w ) under cosine distance. F urthermore, (Levy et al., 2014) show ed that analogy works for past-tense relation v ec ( captur e ) − v ec ( captur ed ) ≈ v ec ( g o ) − v ec ( w ent ) , 1 language-sp ok en-in relation v ec ( f r ance ) − vec ( f r ench ) ≈ v ec ( mexico ) − vec ( spanish ) , as w ell as geographical lo cation v ec ( B er lin ) − vec ( Ger many ) ≈ v ec ( P ar is ) − v ec ( F r ance ) . 1.2 dna2v ec: k-mer em b eddings In this pap er, we present a no vel metho d to compute distributed representations of v ariable-length k-mers. These k-mers are consistent across different lengths, i.e. they lie in the same embedding vector space. W e em b ed k-mers of length 3 ≤ k ≤ 8 , whic h is a space consists of one k-mer p er dimension ( d = P 8 k =3 4 k ), in to a contin uous vector space of 100 dimensions. The training metho d of our shallow tw o-lay er neural net w ork for dna2vec is based on word2v ec. BioV ec (Asgari and Mofrad, 2015) and seq2vec (Kimothi et al., 2016) hav e also applied the word2v ec tec hnique to biological sequences. Although both techniques used a tw o-lay er neural netw ork to train their embedding, our technique is a generalization for v ariable-length k . (Needleman and W unsch, 1970) presented a metho d, now commonly known as Needleman-W unsch algorithm, for computing similarity of k-mers using a dynamic programming scoring of global alignments. But the dynamic programming nature of the algorithm mak es the algorithm slow, with quadratic time complexit y to the length of the sequence. In Section 3.3, we show that its cosine distance, in other words angular distance, is related to Needleman-W unsc h distance of their corresponding k-mers. In Section 3.4, w e provide evidence that nucleotide concatenation analogy can b e constructed with dna2vec arithmetic. The main contribution of this work includes: • v ariable-length k-mer embedding mo del • experimental evidence that shows arithmetic of dna2vec v ectors is akin to nucleotides concatenation • relationship betw een Needleman-W unsch alignmen t and cosine similarit y of dna2v ec vectors • n ucleotide concatenation analogy can b e constructed with dna2vec arithmetic. 2 T raining dna2v ec model The training of dna2v ec consists of four stages: 1) separate genome into long non-ov erlapping DNA fragmen ts 2) con v ert long DNA fragments into ov erlapping v ariable-length k-mers 3) unsup ervised training of an aggregate embedding model using a tw o-lay er neural net work 4) decomp ose aggregated mo del b y k-mer lengths. 2.1 Stage 1: Long non-ov erlapping DNA fragments W e fragment the genome sequence based on gap characters (e.g. X , - , etc). F or our exp eriments using hg38 dataset, the fragmen ts were typically a couple of thousand nucleotides. T o introduce more entrop y , we randomly choose to use the fragment’s rev erse-complemen t. 2.2 Stage 2: Ov erlapping v ariable-length k-mers Giv en a DNA sequence S , we conv ert the sequence S in to ov erlapping fixed length k-mer by sliding a window of length k across S . F or example, we con v ert TAGACTGTC in to five 5-mers: {TAGAC, AGACT, GACTG, ACTGT, CTGTC} . In the v ariable-length case, w e sample k from the discrete uniform distribution Uniform ( k low , k hig h ) to determine the size of each windo w. F or example, a sample of k-mers of k ∈ { 3 , 4 , 5 } could b e {TAGA, AGA, GACT, ACT, CTGTC} . 2 F ormally , given a sequence of length n , S = ( S 1 , S 2 , ..., S n ) where S i ∈ { A, C, G, T } , we con vert S in to ˜ n = n − k hig h + 1 num b er of k-mers: f ( S ) = ( S 1: k 1 , S 2:2+ k 2 , ...S ˜ n : ˜ n + k ˜ n ) k i ∼ Uniform ( k low , k hig h ) where S a : b is a shorthand for ( S a , ..., S b ) . 2.3 Stage 3: T w o-lay er neural netw ork W e use a shallow tw o-lay er neural to train an aggregate DNA k-mer embedding. The metho d is based on word2v ec (Mikolo v et al., 2013a). The word2v ec algorithm has the options of con tinuous bag-of-words (CBO W) or skip-gram. CBOW predicts the targeted w ord given the context, while skip-gram predicts the con text given the targeted word. The w ord2vec homepage 1 claims that skip-gram is slow er to train than CBO W, but skip-gram is b etter for infrequen t words. W e use skip-gram for all our exp eriments. Our dna2vec algorithm is trained by predicting the “context” surrounding a given targeted k-mer. The “con text” is the set of adjacent k-mers surrounding the targeted k-mer. F or example, the context of k-mer GACT w ould b e {TAGA, AGA, ACT, CTGTC} in our previous example from Section 2.2. F or our exp eriments in this pap er, we used a context size of 10 b efore and after the targeted word, which amounts to predicting a total of 20 k-mers. During training, either negative sampling or hierarch y softmax is t ypically used to optimize the up date pro cedure ov er all w ords. W e used negative sampling for all our exp eriments. 2.4 Stage 4: Decompose aggregated mo del b y k-mer lengths W e decomp ose the aggregate mo del by k-mer length to form k hig h − k low + 1 mo dels. This decomp osition is useful for the searc hing of nearest neighbors, as w e will discuss in Section 3.1. 3 Exp erimen ts Our dna2vec 2 w as trained with hg38 human assembly c hr1 to chr22 (Rosen blo om et al., 2015). Sp ecifically , they w ere downloaded from http://hgdo wnload.cse.ucsc.edu/downloads.h tml#human. W e excluded X and Y c hromosomes, as well as mito chondrial and unlo calized sequences. 3.1 Similarit y and nearest neighbors F or each v ector arithmetic solution, we often compute its n -nearest k-mer neighbors. W e define similarity b et w een tw o dna2v ec vectors v , w ∈ R d as the cosine similarit y: sim ( v , w ) = v · w k v kk w k The ne ar est-neighb or of dna2vec v ector v ∈ R d is a k-mer computed with: Ne ar estNeighb or k ( v ) = arg max s ∈{ A,C,G,T } k sim ( v , v ec ( s )) (1) 1 https://code.google.com/archive/p/w ord2vec/ 2 pre-trained dna2vec vectors a v ailable at https://pnpnpn.gith ub.io/dna2vec/ upon publication 3 Generally , the nNe ar estNeighb ors k ( v ) are the n -nearest neigh b oring k-mers to vector v . 3.2 dna2v ec arithmetic and nucleotide concatenation W e found that summing dna2vec em b eddings is related to concatenating k-mers. In T able 1, we inv estigated this hypothesis by adding dna2v ec embeddings of t wo arbitrary k-mers and examining whether their vector sum’s neigh b ors o v erlap with their string concatenation. The 1-NN column results were tallied using Equation (1) and the other columns used nNe ar estNeighb ors from Section 3.1. In this experiment, string concatenation can come from both 5’ and 3’ ends. F or example, the follo wing condition would b e marked as a suc c ess for 1-NN : Ne ar estNeighb or 6 ( v ec ( AAC ) + v ec ( TCT )) ∈ { AACTCT , TCTAAC } Lik ewise, the following w ould b e a suc c ess for n-NN : nNe ar estNeighb ors 6 ( v ec ( AAC ) + v ec ( TCT )) ∩ { AACTCT , TCTAAC } 6 = ∅ T able 1: K-mers concatenation and dna2v ec addition. W e took 1000 samples for each op erand. F or example, the first row is aggregated from summing the dna2v ec vectors of individual pairs of arbitrary 3-mer and observing whether each of their string concatenation o v erlaps with the vector sum’s n -nearest 6-mer neighbors. Op erands Concatenated 1-NN 5-NN 10-NN 3-mer + 3-mer 6-mer 28.7% 80.3% 94.6% 3-mer + 4-mer 7-mer 49.9% 90.4% 97.4% 3-mer + 5-mer 8-mer 53.9% 94.0% 98.4% 4-mer + 4-mer 8-mer 73.5% 96.8% 99.2% 3.3 Relationship to global alignmen t similarit y All of the Needleman-W unsch similarity score in this pap er were computed using Biop ython’s align.globalxx function, which used a match score of 1, mismatch of 0 and gap penalty of 0. In Figure 1, we pro vided evidence that edit distance betw een tw o arbitrary k-mers is correlated with the cosine distance of their corresp onding dna2vec vectors. W e sampled 1000 pairs of 8-mers for each Needleman-W unsch score level and plot their Needleman-W unsch similarit y score against dna2v ec cosine similarit y . In Figure 2, we compared the Needleman-W unsch similarity distribution of k-mer and its nearest dna2vec neigh b or against distribution of t w o random k-mers. Specifically , we sampled 1000 8-mers, found each of its nearest neighbor using Equation (1) , and computed the Needleman-W unsch score for each pair. F or the null distribution, we sampled 1000 pairs of random 8-mers. Thus we found evidence that the dna2vec nearest-neigh b or exhibits alignment similarit y . 3.4 Analogy of nucleotide concatenation W e exp erimented with tw o types of nucleotide concatenation analogy: str ong and we ak concatenations. Giv en t w o k-mers of the same length, we define str ong c onc atenation as splicing n ucleotides on the same end (either 5’ or 3’ end) of the k-mers. An example of 5-mer with 3-nucleotides snipp et would b e: 4 0.00 0.25 0.50 0.75 1.00 1 2 3 4 5 6 7 8 Needleman−W unsch score dna2v ec cosine similarity Figure 1: Boxplot of Needleman-W unsch score and dna2vec cosine similarity . The lo wer and upp er hinges are the 25 and 75 quartiles, resp ectively . The Sp earman’s rank correlation coefficient is 0.831 5 0 250 500 750 2 4 6 Needleman−W unsch score count (out of 1000 samples) relation nearest_neighbor null Figure 2: Global alignment score distribution of nearest-neighbor. The nearest-neighbor distribution is generated b y computing Needleman-W unsch score b etw een 8-mer and its nearest neighbor. The null distribution is from computing the score betw een tw o random 8-mers. 6 v ec ( AC GAT ) − v ec ( GAT ) + v ec ( AT C ) ≈ v ec ( AC AT C ) W e relaxed the same end restriction in the exp eriment we ak c onc atenation , i.e. the result can come from either end: v ec ( AC GAT ) − v ec ( GAT ) + v ec ( AT C ) ∈ approx { v ec ( AC AT C ) , v ec ( AT C AC ) } Exp erimen tal samples were generated from randomly sampling t wo k-mers of equal length and a nucleotide snipp et (3 or 4 n ucleotides) for concatenation. F or both strong and weak concatenation experiments, we randomly selected either the 5’ and 3’ end to splice. T able 2 sho ws the summary of exp erimen tal results from the t wo t yp es of n ucleotide concatenations. P articularly , we get 88% accuracy with weak concatenation analogy of 8-mer and 4-nucleotides snipp et when considering 10-NN, as defined in Section 3.1. Note that considering 30-nearest neigh b ors is relatively small comparing to the space of all p ossible 6-mers, it is merely 0.73% of all possible 6-mers and 0.046% of all p ossible 8-mers. T o confirm whether the arithmetic was actually extending a k-mer by the snipp et as opp ose to similarit y comparison, we compared the analogy results with scr amble d-snipp et exp eriments, whic h concatenated a differen t random snipp ets in the answer case. As exp ected, the vector arithmetic was significan tly fav oring the correct matching snipp et ( analogy column) ov er a different random snipp et ( scrambled-snippet column) in Figure 3 and T able 2. T able 2: Analogy Exp erimen t. W e analyzed t wo types of analogies: we ak and str ong concatenation. 1000 samples w ere randomly gen- erated for eac h type. F or comparison, we generated 1000 samples using scrambled-snippet sampling strategy . dimension w eak-concat scram bled-snipp et 5 / 10 / 30-NN w eak-concat analogy 5 / 10 / 30-NN strong-concat scram bled-snipp et 5 / 10 / 30-NN strong-concat analogy 5 / 10 / 30-NN 6-mer with 3-n t snipp et 1.4 / 4 / 16% 47 / 69 / 95% 0.6 / 1.8 / 9% 43 / 62 / 88% 7-mer with 3-n t snipp et 2.4 / 6 / 16% 66 / 82 / 96% 1.5 / 3.8 / 10% 61 / 76 / 92% 8-mer with 3-n t snipp et 3 / 6 / 19% 67 / 82 / 95% 2.3 / 3.8 / 11% 62 / 77 / 91% 8-mer with 4-n t snipp et 0.7 / 1.4 / 3% 75 / 88 / 98% 0.3 / 1.0 / 2.4% 69 / 83 / 95% 3.5 Implemen tation Details W e will make our co de and data av ailable at h ttps://pnpnpn.github.io/dna2v ec/ up on publication. The t w o-lay er neural netw ork training metho d describ ed in Section 2.3 was implemented using gensim framew ork (Řehůřek and So jka, 2010). W e used gensim’s Word2vec class with parameters sg=1 and window=10 , which sp ecified the usage of skipgram mo del and the half-size of the context window as 10, respectively . All of trained dna2v ec v ectors used in this pap er has dimension size of 100. Since the window sliding step in Section 2.2 is sto c hastic in terms of v ariable-length k , w e could essen tially generate more training data by lo oping through the complete genomic sequence data with multiple passes, which we called ep o chs . The dna2vec mo del used in this pap er was trained with 10 ep o c hs. The training step to ok o ver 3 days using gensim parameter workers=4 on a 2.66 GHz Quad-Core Intel Xeon with 8GB memory . 7 0 250 500 750 1000 0 25 50 75 100 n−Nearest Neighbors count (out of 1000 samples) task analogy scramb led−snippet Figure 3: Cumulativ e mass for analogy exp eriment of 8-mer with 3-nt snipp et. 1000 samples were generated with the strong-concatenation analogy setup. W e compared it with another 1000 samples using the scr amble d- snipp et sampling pro cedure. 8 4 Discussion In this w ork, we presented a nov el metho d for training distributed representations of k-mers. W e demonstrated that our dna2vec embeddings can represen t v ariable-length k-mers in a consisten t fashion via nucleotide concatenation exp eriments. W e pro vided experimental evidence sho wing that the arithmetic of dna2vec v ectors is akin to nucleotides concatenation. W e also show ed that Needleman-W unsch similarit y score b etw een t w o arbitrary k-mers is correlated with the cosine distance of their corresp onding dna2v ec vectors. As for future work, due to the fact that man y machine learning algorithms require fixed-length con tinuous vectors as input, we will explore the application of dna2vec with mac hine learning techniques on biological sequence analysis. References Angerm ueller, C., Pärnamaa, T., Parts, L., and Stegle, O. (2016). Deep learning for computational biology . Mole cular systems biolo gy , 12(7):878. Asgari, E. and Mofrad, M. R. (2015). Contin uous distributed represen tation of biological sequences for deep proteomics and genomics. PloS one , 10(11):e0141287. Bahdanau, D., Cho, K., and Bengio, Y. (2014). Neural machine translation by jointly learning to align and translate. arXiv pr eprint arXiv:1409.0473 . Bengio, Y., Courville, A., and Vincen t, P . (2013). Representation learning: A review and new persp ectives. IEEE tr ansactions on p attern analysis and machine intel ligenc e , 35(8):1798–1828. Cho, K., V an Merriën b o er, B., Gulcehre, C., Bahdanau, D., Bougares, F., Sch wenk, H., and Bengio, Y. (2014). Learning phrase represen tations using rnn encoder-deco der for statistical mac hine translation. arXiv pr eprint arXiv:1406.1078 . Chopra, S., A uli, M., R ush, A. M., and Harv ard, S. (2016). Abstractive sen tence summarization with attentiv e recurren t neural netw orks. Pr o c e e dings of NAA CL-HL T16 , pages 93–98. Dos Santos, C. N. and Gatti, M. (2014). Deep conv olutional neural netw orks for sen timent analysis of short texts. In COLING , pages 69–78. Harris, Z. S. (1954). Distributional structure. W or d , 10(2-3):146–162. Kim, Y. (2014). Conv olutional neural netw orks for sen tence classification. arXiv pr eprint arXiv:1408.5882 . Kimothi, D., Soni, A., Biyani, P ., and Hogan, J. M. (2016). Distributed represen tations for biological sequence analysis. arXiv pr eprint arXiv:1608.05949 . LeCun, Y., Bengio, Y., and Hinton, G. (2015). Deep learning. Natur e , 521(7553):436–444. Levy , O., Goldb erg, Y., and Ramat-Gan, I. (2014). Linguistic regularities in sparse and explicit word represen tations. In CoNLL , pages 171–180. Mik olo v, T., Chen, K., Corrado, G., and Dean, J. (2013a). Efficient estimation of word representations in v ector space. arXiv pr eprint arXiv:1301.3781 . Mik olo v, T., Sutskev er, I., Chen, K., Corrado, G. S., and Dean, J. (2013b). Distributed represen tations of w ords and phrases and their comp ositionality . In A dvanc es in neur al information pr o c essing systems , pages 3111–3119. Mik olo v, T., Yih, W.-t., and Zw eig, G. (2013c). Linguistic regularities in contin uous space w ord representations. In HL T-NAA CL , volume 13, pages 746–751. 9 Needleman, S. B. and W unsch, C. D. (1970). A general method applicable to the searc h for similarities in the amino acid sequence of tw o proteins. Journal of mole cular biolo gy , 48(3):443–453. Řehůřek, R. and Sojka, P . (2010). Soft ware F ramework for T opic Modelling with Large Corpora. In Pr o c e e dings of the LREC 2010 W orkshop on New Chal lenges for NLP F r ameworks , pages 45–50, V alletta, Malta. ELRA. h ttp://is.m uni.cz/publication/884893/en . Rosen blo om, K. R., Armstrong, J., Barb er, G. P ., Casp er, J., Clawson, H., Diekhans, M., Dreszer, T. R., F ujita, P . A., Guruv ado o, L., Haeussler, M., et al. (2015). The ucsc genome browser database: 2015 update. Nucleic acids r ese ar ch , 43(D1):D670–D681. Sutsk ev er, I., Vin y als, O., and Le, Q. V. (2014). Sequence to sequence learning with neural netw orks. In A dvanc es in neur al information pr o c essing systems , pages 3104–3112. Suyk ens, J. A. and V andewalle, J. (1999). Least squares support vector mac hine classifiers. Neur al pr o c essing letters , 9(3):293–300. T urian, J., Ratino v, L., and Bengio, Y. (2010). W ord representations: a simple and general metho d for semi-sup ervised learning. In Pr o c e e dings of the 48th annual me eting of the asso ciation for c omputational linguistics , pages 384–394. Asso ciation for Computational Linguistics. Vin y als, O., T oshev, A., Bengio, S., and Erhan, D. (2015). Show and tell: A neural image caption generator. In Pr o c e e dings of the IEEE Confer enc e on Computer V ision and Pattern R e c o gnition , pages 3156–3164. 10
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment