Recurrent Neural Networks for Multivariate Time Series with Missing Values
Multivariate time series data in practical applications, such as health care, geoscience, and biology, are characterized by a variety of missing values. In time series prediction and other related tasks, it has been noted that missing values and thei…
Authors: Zhengping Che, Sanjay Purushotham, Kyunghyun Cho
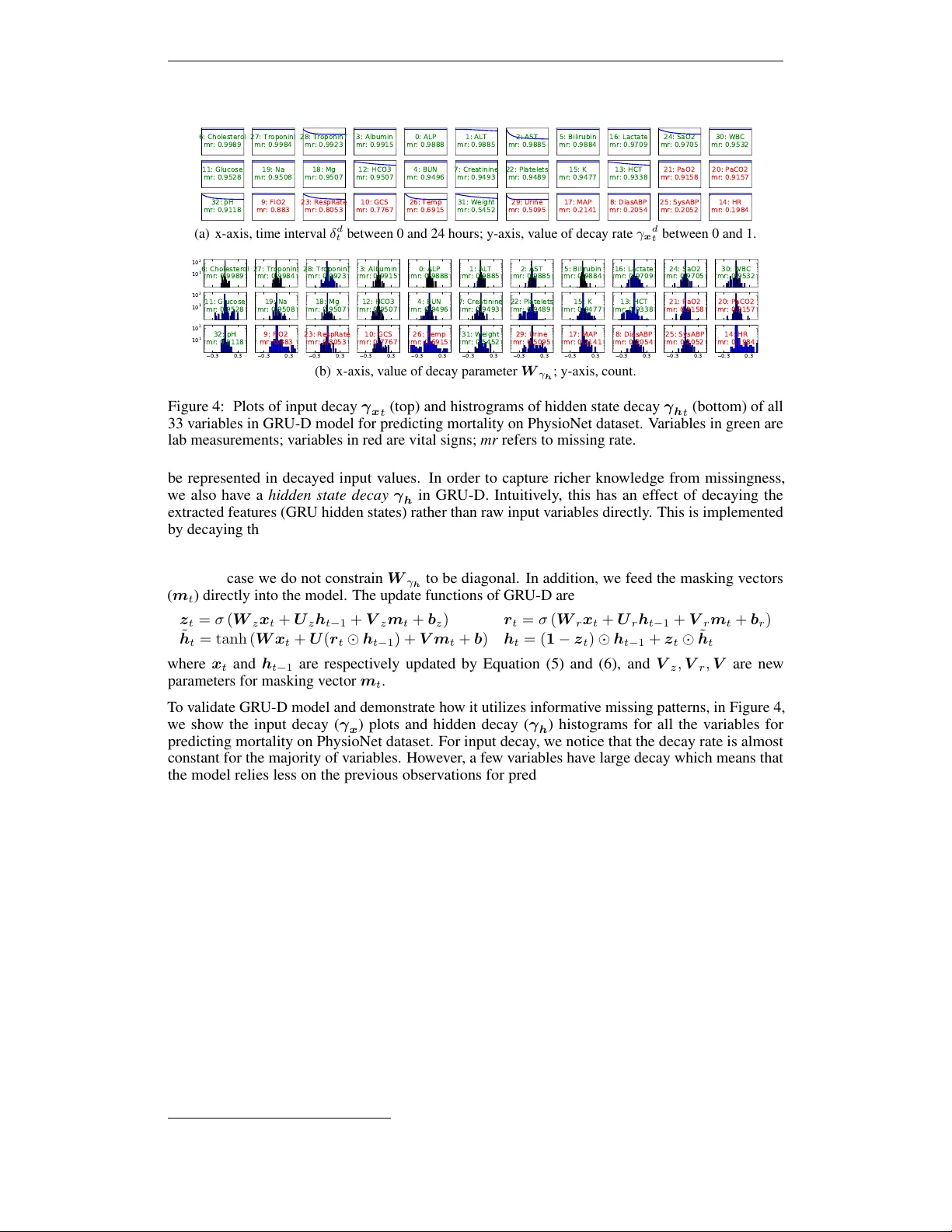
R E C U R R E N T N E U R A L N E T W O R K S F O R M U L T I V A R I - A T E T I M E S E R I E S W I T H M I S S I N G V A L U E S Zhengping Che, Sanjay Purushotham Department of Computer Science Univ ersity of Southern California Los Angeles, CA 90089, USA { zche,spurusho } @usc.edu K yunghyun Cho, Da vid Sontag Department of Computer Science New Y ork Univ ersity New Y ork, NY 10012, USA kyunghyun.cho@nyu.edu,dsontag@cs.nyu.edu Y an Liu Department of Computer Science Univ ersity of Southern California Los Angeles, CA 90089, USA yanliu.cs@usc.edu A B S T R AC T Multiv ariate time series data in practical applications, such as health care, geo- science, and biology , are characterized by a v ariety of missing v alues. In time series prediction and other related tasks, it has been noted that missing values and their missing patterns are often correlated with the target labels, a.k.a., informative miss- ingness. There is v ery limited work on exploiting the missing patterns for ef fecti ve imputation and improving prediction performance. In this paper, we de velop nov el deep learning models, namely GR U-D, as one of the early attempts. GR U-D is based on Gated Recurrent Unit (GR U), a state-of-the-art recurrent neural network. It takes two representations of missing patterns, i.e., masking and time interval , and effecti vely incorporates them into a deep model architecture so that it not only captures the long-term temporal dependencies in time series, b ut also utilizes the missing patterns to achie ve better prediction results. Experiments of time series classification tasks on real-world clinical datasets (MIMIC-III, PhysioNet) and synthetic datasets demonstrate that our models achiev e state-of-the-art performance and provides useful insights for better understanding and utilization of missing values in time series analysis. 1 I N T RO D U C T I O N Multiv ariate time series data are ubiquitous in man y practical applications ranging from health care, geoscience, astronomy , to biology and others. They often ine vitably carry missing observ ations due to various reasons, such as medical e vents, sa ving costs, anomalies, incon venience and so on. It has been noted that these missing v alues are usually informative missingness (Rubin, 1976), i.e., the missing values and patterns pro vide rich information about target labels in supervised learning tasks (e.g, time series classification). T o illustrate this idea, we show some examples from MIMIC-III, a real world health care dataset in Figure 1. W e plot the Pearson correlation coef ficient between variable missing rates, which indicates how often the variable is missing in the time series, and the labels of our interests such as mortality and ICD-9 diagnoses. W e observ e that the missing rate is correlated with the labels, and the missing rates with lo w rate values are usually highly (either positive or negativ e) correlated with the labels. These findings demonstrate the usefulness of missingness patterns in solving a prediction task. In the past decades, various approaches hav e been dev eloped to address missing v alues in time series (Schafer & Graham, 2002). A simple solution is to omit the missing data and to perform analysis only on the observed data. A v ariety of methods hav e been dev eloped to fill in the missing values, such as smoothing or interpolation (Kreindler & Lumsden, 2012), spectral analysis (Mondal & Perci val, 2010), k ernel methods (Rehfeld et al., 2011), multiple imputation (White et al., 2011), 1 0 . 8 1 80 60 40 20 20 40 60 80 -0.2 -0.1 0 0.1 0.2 5 10 15 20 20 40 60 80 -0.2 -0.1 0 0.1 0.2 Figure 1: Demonstrations of informative missingness on MIMIC-III dataset. Left figure shows variable missing rate (x-axis, missing rate; y-axis, input variable). Middle/right figures respecti vely shows the correlations between missing rate and mortality/ICD-9 diagnosis categories (x-axis, target label; y-axis, input variable; color , correlation value). Please refer to Appendix A.1 for more details. and EM algorithm (Garc ´ ıa-Laencina et al., 2010). Schafer & Graham (2002) and references therein provide excellent revie ws on related solutions. Howe ver , these solutions often result in a two- step process where imputations are disparate from prediction models and missing patterns are not effecti vely e xplored, thus leading to suboptimal analyses and predictions (W ells et al., 2013). In the meantime, Recurrent Neural Networks (RNNs), such as Long Short-T erm Memory (LSTM) (Hochreiter & Schmidhuber, 1997) and Gated Recurrent Unit (GR U) (Cho et al., 2014), hav e shown to achie ve the state-of-the-art results in many applications with time series or sequen- tial data, including machine translation (Bahdanau et al., 2014; Sutske ver et al., 2014) and speech recognition (Hinton et al., 2012). RNNs enjoy se veral nice properties such as strong prediction performance as well as the ability to capture long-term temporal dependencies and v ariable-length observations. RNNs for missing data has been studied in earlier works (Bengio & Gingras, 1996; T resp & Briegel, 1998; Parv een & Green, 2001) and applied for speech recognition and blood-glucose prediction. Recent works (Lipton et al., 2016; Choi et al., 2015) tried to handle missingness in RNNs by concatenating missing entries or timestamps with the input or performing simple imputations. Howe ver , there hav e not been works which systematically model missing patterns into RNN for time series classification problems. Exploiting the po wer of RNNs along with the informativeness of missing patterns is a ne w promising venue to effecti vely model multiv ariate time series and is the main motiv ation behind our work. In this paper , we develop a no vel deep learning model based on GRU, namely GR U-D, to effecti vely exploit two representations of informative missingness patterns, i.e., masking and time interval . Masking informs the model which inputs are observed (or missing), while time interval encapsulates the input observation patterns. Our model captures the observ ations and their dependencies by applying masking and time interv al (using a decay term) to the inputs and network states of GR U, and jointly train all model components using back-propagation. Thus, our model not only captures the long-term temporal dependencies of time series observations b ut also utilizes the missing patterns to improv e the prediction results. Empirical experiments on real-world clinical datasets as well as synthetic datasets demonstrate that our proposed model outperforms strong deep learning models built on GR U with imputation as well as other strong baselines. These experiments show that our proposed method is suitable for many time series classification problems with missing data, and in particular is readily applicable to the predicti ve tasks in emerging health care applications. Moreover , our method pro vides useful insights into more general research challenges of time series analysis with missing data beyond classification tasks, including 1) a general deep learning framework to handle time series with missing data, 2) ef fectiv e solutions to characterize the missing patterns of not missing-completely-at-random time series data such as modeling masking and time interv al, and 3) an insightful approach to study the impact of variable missingness on the prediction labels by decay analysis. 2 R N N M O D E L S F O R T I M E S E R I E S W I T H M I S S I N G V A R I A B L E S W e denote a multiv ariate time series with D variables of length T as X = ( x 1 , x 2 , . . . , x T ) T ∈ R T × D , where x t ∈ R D represents the t -th observ ations (a.k.a., measurements) of all variables and x d t denotes the measurement of d-th v ariable of x t . Let s t ∈ R denote the time-stamp when the t -th observation is obtained and we assume that the first observ ation is made at time t = 0 ( s 1 = 0 ). A 2 𝑿 : Input t ime seri es (2 v ari ables); 𝒔 : Ti mestamps f or 𝑿 ; 𝑿 = 4 7 49 𝑁𝐴 40 𝑁𝐴 43 55 𝑁𝐴 15 14 𝑁𝐴 𝑁𝐴 𝑁𝐴 15 𝒔 = 0 0 . 1 0 . 6 1 . 6 2 . 2 2 . 5 3 . 1 𝑴 : Mask in g f or 𝑿 ; 𝚫 : Ti me in t erv al f or 𝑿 . 𝑴 = 1 1 0 1 0 1 1 0 1 1 0 0 0 1 𝚫 = 0 . 0 0 . 1 0 . 5 1 . 5 0 . 6 0 . 9 0 . 6 0 . 0 0 . 1 0 . 5 1 . 0 1 . 6 1 . 9 2 . 5 Figure 2: An e xample of measurement vectors x t , time stamps s t , masking m t , and time interval δ t . time series X could hav e missing v alues. W e introduce a masking vector m t ∈ { 0 , 1 } D to denote which variables are missing at time step t . The masking vector for x t is giv en by m d t = 1 , if x d t is observed 0 , otherwise For each v ariable d , we also maintain the time interval δ d t ∈ R since its last observ ation as δ d t = s t − s t − 1 + δ d t − 1 , t > 1 , m d t − 1 = 0 s t − s t − 1 , t > 1 , m d t − 1 = 1 0 , t = 1 An example of these notations is illustrated in Figure 2. In this paper , we are interested in the time series classification problem, where we predict the labels l n giv en the time series data D , where D = { ( X n , s n , M n , ∆ n , l n ) } N n =1 , and X n = h x ( n ) 1 , . . . , x ( n ) T n i , s n = h s ( n ) 1 , . . . , s ( n ) T n i , M n = h m ( n ) 1 , . . . , m ( n ) T n i , ∆ n = h δ ( n ) 1 , . . . , δ ( n ) T n i , and l n ∈ { 1 , . . . , L } . 2 . 1 G R U - R N N F O R T I M E S E R I E S C L A S S I FI C A T I O N W e in vestigate the use of recurrent neural networks (RNN) for time-series classification, as their recursiv e formulation allow them to handle variable-length sequences naturally . Moreov er, RNN shares the same parameters across all time steps which greatly reduces the total number of parameters we need to learn. Among dif ferent variants of the RNN, we specifically consider an RNN with gated recurrent units (Cho et al., 2014; Chung et al., 2014), but similar discussion and con volutions are also valid for other RNN models such as LSTM (Hochreiter & Schmidhuber, 1997). The structure of GR U is shown in Figure 3(a). GRU has a reset gate r j t and an update gate z j t for each of the hidden state h j t to control. At each time t , the update functions are shown as follo ws: z t = σ ( W z x t + U z h t − 1 + b z ) r t = σ ( W r x t + U r h t − 1 + b r ) ˜ h t = tanh ( W x t + U ( r t h t − 1 ) + b ) h t = ( 1 − z t ) h t − 1 + z t ˜ h t where matrices W z , W r , W , U z , U r , U and vectors b z , b r , b are model parameters. W e use σ for element-wise sigmoid function, and for element-wise multiplication. This formulation assumes that all the v ariables are observed. A sigmoid or soft-max layer is then applied on the output of the GR U layer at the last time step for classification task. Existing work on handling missing values lead to three possible solutions with no modification on GR U network structure. One straightforward approach is simply replacing each missing observ ation with the mean of the variable across the training e xamples. In the context of GR U, we hav e x d t ← m d t x d t + (1 − m d t ) ˜ x d (1) 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒓 (a) GR U 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒎 𝒓 MASK 𝜸 𝒉 𝜸 𝒙 (b) GR U-D Figure 3: Graphical illustrations of the original GR U (left) and the proposed GRU-D (right) models. 3 where ˜ x d = P N n =1 P T n t =1 m d t,n x d t,n . P N n =1 P T n t =1 m d t,n . W e refer to this approach as GRU-mean . A second approach is exploiting the temporal structure in time series. For e xample, we may assume any missing v alue is same as its last measurement and use forward imputation ( GR U-forward ), i.e., x d t ← m d t x d t + (1 − m d t ) x d t 0 (2) where t 0 < t is the last time the d -th v ariable was observ ed. Instead of explicitly imputing missing v alues, the third approach simply indicates which variables are missing and ho w long they hav e been missing as a part of input, by concatenating the measurement, masking and time interval v ectors as x ( n ) t ← h x ( n ) t ; m ( n ) t ; δ ( n ) t i (3) where x ( n ) t can be either from Equation (1) or (2). W e later refer to this approach as GRU-simple . These approaches solve the missing v alue issue to a certain extent, Howe ver , it is kno wn that imputing the missing v alue with mean or forward imputation cannot distinguish whether missing values are imputed or truly observ ed. Simply concatenating masking and time interval v ectors fails to exploit the temporal structure of missing v alues. Thus none of them fully utilize missingness in data to achiev e desirable performance. 2 . 2 G R U - D : M O D E L W I T H T R A I NA B L E D E C AY S T o fundamentally address the issue of missing values in time series, we notice two important properties of the missing v alues in time series, especially in health care domain: First, the v alue of the missing variable tend to be close to some default v alue if its last observation happens a long time ago. This property usually exists in health care data for human body as homeostasis mechanisms and is considered to be critical for disease diagnosis and treatment (V odovotz et al., 2013). Second, the influence of the input variables will fade a way ov er time if the v ariable has been missing for a while. F or example, one medical feature in electronic health records (EHRs) is only significant in a certain temporal context (Zhou & Hripcsak, 2007). Therefore we propose a GR U-based model called GR U-D , in which a decay mechanism is designed for the input variables and the hidden states to capture the aforementioned properties. W e introduce decay rates in the model to control the decay mechanism by considering the following important factors. First, each input variable in health care time series has its o wn medical meaning and importance. The decay rates should be flexible to dif fer from variable to variable based on the underlying properties associated with the variables. Second, as we see lots of missing patterns are informati ve in prediction tasks, the decay rate should be indicativ e of such patterns and benefits the prediction tasks. Furthermore, since the missing patterns are unknown and possibly comple x, we aim at learning decay rates from the training data rather than being fixed a priori. That is, we model a vector of decay rates γ as γ t = exp {− max ( 0 , W γ δ t + b γ ) } (4) where W γ and b γ are model parameters that we train jointly with all the other parameters of the GR U. W e chose the exponentiated ne gativ e rectifier in order to keep each decay rate monotonically decreasing in a reasonable range between 0 and 1 . Note that other formulations such as a sigmoid function can be used instead, as long as the resulting decay is monotonic and is in the same range. Our proposed GR U-D model incorporates two dif ferent trainable decays to utilize the missingness directly with the input feature v alues and implicitly in the RNN states. First, for a missing variable, we use an input decay γ x to decay it over time toward the empirical mean (which we take as a default configuration), instead of using the last observ ation as it is. Under this assumption, the trainable decay scheme can be readily applied to the measurement vector by x d t ← m d t x d t + (1 − m d t ) γ x d t x d t 0 + (1 − m d t )(1 − γ x d t ) ˜ x d (5) where x d t 0 is the last observation of the d -th v ariable ( t 0 < t ) and ˜ x d is the empirical mean of the d -th v ariable. When decaying the input variable directly , we constrain W γ x to be diagonal, which effecti vely makes the decay rate of each v ariable independent from the others. Sometimes the input decay may not fully capture the missing patterns since not all missingness information can 4 6: Cholesterol mr: 0.9989 27: TroponinI mr: 0.9984 28: TroponinT mr: 0.9923 3: Albumin mr: 0.9915 0: ALP mr: 0.9888 1: ALT mr: 0.9885 2: AST mr: 0.9885 5: Bilirubin mr: 0.9884 16: Lactate mr: 0.9709 24: SaO2 mr: 0.9705 30: WBC mr: 0.9532 11: Glucose mr: 0.9528 19: Na mr: 0.9508 18: Mg mr: 0.9507 12: HCO3 mr: 0.9507 4: BUN mr: 0.9496 7: Creatinine mr: 0.9493 22: Platelets mr: 0.9489 15: K mr: 0.9477 13: HCT mr: 0.9338 21: PaO2 mr: 0.9158 20: PaCO2 mr: 0.9157 32: pH mr: 0.9118 9: FiO2 mr: 0.883 23: RespRate mr: 0.8053 10: GCS mr: 0.7767 26: Temp mr: 0.6915 31: Weight mr: 0.5452 29: Urine mr: 0.5095 17: MAP mr: 0.2141 8: DiasABP mr: 0.2054 25: SysABP mr: 0.2052 14: HR mr: 0.1984 (a) x-axis, time interv al δ d t between 0 and 24 hours; y-axis, value of decay rate γ x d t between 0 and 1. 1 0 1 1 0 2 6: Cholesterol mr: 0.9989 27: TroponinI mr: 0.9984 28: TroponinT mr: 0.9923 3: Albumin mr: 0.9915 0: ALP mr: 0.9888 1: ALT mr: 0.9885 2: AST mr: 0.9885 5: Bilirubin mr: 0.9884 16: Lactate mr: 0.9709 24: SaO2 mr: 0.9705 30: WBC mr: 0.9532 1 0 1 1 0 2 11: Glucose mr: 0.9528 19: Na mr: 0.9508 18: Mg mr: 0.9507 12: HCO3 mr: 0.9507 4: BUN mr: 0.9496 7: Creatinine mr: 0.9493 22: Platelets mr: 0.9489 15: K mr: 0.9477 13: HCT mr: 0.9338 21: PaO2 mr: 0.9158 20: PaCO2 mr: 0.9157 0.3 0.3 1 0 1 1 0 2 32: pH mr: 0.9118 0.3 0.3 9: FiO2 mr: 0.883 0.3 0.3 23: RespRate mr: 0.8053 0.3 0.3 10: GCS mr: 0.7767 0.3 0.3 26: Temp mr: 0.6915 0.3 0.3 31: Weight mr: 0.5452 0.3 0.3 29: Urine mr: 0.5095 0.3 0.3 17: MAP mr: 0.2141 0.3 0.3 8: DiasABP mr: 0.2054 0.3 0.3 25: SysABP mr: 0.2052 0.3 0.3 14: HR mr: 0.1984 (b) x-axis, v alue of decay parameter W γ h ; y-axis, count. Figure 4: Plots of input decay γ x t (top) and histrograms of hidden state decay γ h t (bottom) of all 33 v ariables in GRU-D model for predicting mortality on PhysioNet dataset. V ariables in green are lab measurements; variables in red are vital signs; mr refers to missing rate. be represented in decayed input v alues. In order to capture richer kno wledge from missingness, we also ha ve a hidden state decay γ h in GR U-D. Intuiti vely , this has an effect of decaying the extracted features (GR U hidden states) rather than ra w input variables directly . This is implemented by decaying the previous hidden state h t − 1 before computing the new hidden state h t as h t − 1 ← γ h t h t − 1 , (6) in which case we do not constrain W γ h to be diagonal. In addition, we feed the masking vectors ( m t ) directly into the model. The update functions of GR U-D are z t = σ ( W z x t + U z h t − 1 + V z m t + b z ) r t = σ ( W r x t + U r h t − 1 + V r m t + b r ) ˜ h t = tanh ( W x t + U ( r t h t − 1 ) + V m t + b ) h t = ( 1 − z t ) h t − 1 + z t ˜ h t where x t and h t − 1 are respecti vely updated by Equation (5) and (6), and V z , V r , V are ne w parameters for masking vector m t . T o validate GR U-D model and demonstrate how it utilizes informati ve missing patterns, in Figure 4, we show the input decay ( γ x ) plots and hidden decay ( γ h ) histograms for all the variables for predicting mortality on PhysioNet dataset. F or input decay , we notice that the decay rate is almost constant for the majority of v ariables. Ho wev er , a few v ariables hav e large decay which means that the model relies less on the previous observations for prediction. For example, the changes in the variable v alues of weight, arterial pH, temperature, and respiration rate are kno wn to impact the ICU patients health condition. The hidden decay histograms sho w the distribution of decay parameters related to each variable. W e noticed that the parameters related to v ariables with smaller missing rate are more spread out. This indicates that the missingness of those v ariables has more impact on decaying or keeping the hidden states of the models. Notice that the decay term can be generalized to LSTM straightforwardly . In practical applications, missing values i n time series may contain useful information in a variety of ways. A better model should hav e the flexibility to capture different missing patterns. In order to demonstrate the capacity of our GR U-D model, we discuss some model variations in Appendix A.2. 3 E X P E R I M E N T S 3 . 1 D A TA S E T D E S C R I P T I O N S A N D E X P E R I M E N TA L D E S I G N W e demonstrate the performance of our proposed models on one synthetic and two real-world health-care datasets 1 and compare it to sev eral strong machine learning and deep learning approaches in classification tasks. W e ev aluate our models for dif ferent settings such as early prediction and different training sizes and in vestigate the impact of informati ve missingness. 1 A summary statistics of the three datasets is shown in Appendix A.3.1. 5 Gesture phase segmentation dataset (Gestur e) This UCI dataset (Madeo et al., 2013) has multi- variate time series features, regularly sampled and with no missing v alues, for 5 different gesticula- tions. W e extracted 378 time series and generate 4 synthetic datasets for the purpose of understanding model behaviors with dif ferent missing patterns. W e treat it as multi-class classification task. Physionet Challenge 2012 dataset (PhysioNet) This dataset, from PhysioNet Challenge 2012 (Silv a et al., 2012), is a publicly a vailable collection of multiv ariate clinical time series from 8000 intensiv e care unit (ICU) records. Each record is a multiv ariate time series of roughly 48 hours and contains 33 variables such as Alb umin, heart-rate, glucose etc. W e used T raining Set A subset in our experiments since outcomes (such as in-hospital mortality labels) are publicly a vailable only for this subset. W e conduct the following two prediction tasks on this dataset: 1) Mortality task : Predict whether the patient dies in the hospital. There are 554 patients with positi ve mortality label. W e treat this as a binary classification problem. and 2) All 4 tasks : Predict 4 tasks: in-hospital mortality , length-of-stay less than 3 days, whether the patient had a cardiac condition, and whether the patient was recov ering from surgery . W e treat this as a multi-task classification problem. MIMIC-III dataset (MIMIC-III) This public dataset (Johnson et al., 2016) has deidentified clinical care data collected at Beth Israel Deaconess Medical Center from 2001 to 2012. It contains ov er 58,000 hospital admission records. W e extracted 99 time series features from 19714 admission records for 4 modalities including input-events (fluids into patient, e.g., insulin), output-ev ents (fluids out of the patient, e.g., urine), lab-ev ents (lab test results, e.g., pH values) and prescription-e vents (drugs prescribed by doctors, e.g., aspirin). These modalities are kno wn to be e xtremely useful for monitoring ICU patients. W e only use the first 48 hours data after admission from each time series. W e perform follo wing two predicti ve tasks: 1) Mortality task : Predict whether the patient dies in the hospital after 48 hours. There are 1716 patients with positive mortality label and we perform binary classification. and 2) ICD-9 Code tasks : Predict 20 ICD-9 diagnosis cate gories (e.g., respiratory system diagnosis) for each admission. W e treat this as a multi-task classification problem. 3 . 2 M E T H O D S A N D I M P L E M E N T ATI O N D E TA I L S W e categorize all ev aluated prediction models into three following groups: • Non-RNN Baselines (Non-RNN) : W e e valuate logistic regression (LR), support vector machines (SVM) and Random Forest (RF) which are widely used in health care applications. • RNN Baselines (RNN) : W e take GR U-mean, GRU-forw ard, GR U-simple, and LSTM-mean (LSTM model with mean-imputation on the missing measurements) as RNN baselines. • Pr oposed Methods (Pr oposed) : This is our proposed GR U-D model from Section 2.2. Recently RNN models have been e xplored for modeling diseases and patient diagnosis in health care domain (Lipton et al., 2016; Choi et al., 2015; Pham et al., 2016) using EHR data. These methods do not systematically handle missing values in data or are equiv alent to our RNN baselines. W e provide more detailed discussions and comparisons in Appendix A.2.3 and A.3.4. The non-RNN baselines cannot handle missing data directly . W e carefully design experiments for non- RNN models to capture the informative missingness as much as possible to ha ve fair comparison with the RNN methods. Since non-RNN models only w ork with fixed length inputs, we regularly sample the time-series data to get a fixed length input and perform imputation to fill in the missing values. Similar to RNN baselines, we can concatenate the masking vector along with the measurements and feed it to non-RNN models. For PhysioNet dataset, we sample the time series on an hourly basis and propagate measurements forward (or backward) in time to fill gaps. For MIMIC-III dataset, we consider tw o hourly samples (in the first 48 hours) and do forward (or backward) imputation. Our preliminary experiments sho wed 2-hourly samples obtains better performance than one-hourly samples for MIMIC-III. W e report results for both concatenation of input and masking vectors (i.e., SVM/LR/RF-simple) and only input vector without masking (i.e., SVM/LR/RF-forward). W e use the scikit-learn (Pedregosa et al., 2011) for the non-RNN model implementation and tune the parameters by cross-validation. W e choose RBF kernel for SVM since it performs better than other kernels. For RNN models, we use a one layer RNN to model the sequence, and then apply a soft-max re gressor on top of the last hidden state h T to do classification. W e use 100 and 64 hidden units in GR U-mean for MIMIC-III and PhysioNet datasets, respectiv ely . All the other RNN models were constructed to 6 0. 6 0 . 7 0. 8 0. 9 1 0 0 . 2 0 . 5 0 . 8 G RU-m ea n GRU- f orwa r d G RU-s im ple GRU- D Figure 5: Classification performance on Ges- ture synthetic datasets. x-axis: average Pear- son correlation of v ariable missing rates and target label in that dataset; y-axis: A UC score. T able 1: Model performances measured by average A UC score ( mean ± std ) for multi-task predictions on real datasets. Results on each class are sho wn in Appendix A.3.3 for reference. Models MIMIC-III PhysioNet ICD-9 20 tasks All 4 tasks GR U-mean 0 . 7070 ± 0 . 001 0 . 8099 ± 0 . 011 GR U-forward 0 . 7077 ± 0 . 001 0 . 8091 ± 0 . 008 GR U-simple 0 . 7105 ± 0 . 001 0 . 8249 ± 0 . 010 GR U-D 0 . 7123 ± 0 . 003 0 . 8370 ± 0 . 012 hav e a comparable number of parameters. 2 For GR U-simple, we use mean imputation for input as shown in Equation (1). Batch normalization (Iof fe & Szegedy, 2015) and dropout (Sri vasta va et al., 2014) of rate 0.5 are applied to the top regressor layer . W e train all the RNN models with the Adam optimization method (Kingma & Ba, 2014) and use early stopping to find the best weights on the validation dataset. All the input v ariables are normalized to be 0 mean and 1 standard deviation. W e report the results from 5-fold cross validation in terms of area under the R OC curve (A UC score). 3 . 3 Q U A N T I T AT I V E R E S U LT S Exploiting informati ve missingness on synthetic dataset As illustrated in Figure 1, missing pat- terns can be useful in solving prediction tasks. A robust model should exploit informative missingness properly and av oid inducing inexistent relations between missingness and predictions. T o ev aluate the impact of modeling missingness we conduct e xperiments on the synthetic Gesture datasets. W e process the data in 4 different settings with the same missing rate b ut different correlations between missing rate and the label. A higher correlation implies more informati ve missingness. Figure 5 sho ws the A UC score comparison of three GR U baseline models (GR U-mean, GRU-forw ard, GRU-simple) and the proposed GR U-D. Since GR U-mean and GR U-forward do not utilize any missingness (i.e., masking or time interval), they perform similarly across all 4 settings. GR U-simple and GR U-D benefit from utilizing the missingness, especially when the correlation is high. Our GR U-D achiev es the best performance in all settings, while GR U-simple fails when the correlation is low . The results on synthetic datasets demonstrates that our proposed model can model and distinguish useful missing patterns in data properly compared with baselines. Prediction task evaluation on r eal datasets W e ev aluate all methods in Section 3.2 on MIMIC-III and PhysioNet datasets. W e noticed that dropout in the recurrent layer helps a lot for all RNN models on both of the datasets, probably because they contain more input v ariables and training samples than synthetic dataset. Similar to Gal (2015), we apply dropout rate of 0.3 with same dropout samples at each time step on weights W , U , V . T able 2 sho ws the prediction performance of all the models on mortality task. All models except for random forest improve their performance when they feed missingness indicators along with inputs. The proposed GRU-D achiev es the best A UC score on both the datasets. W e also conduct multi-task classification e xperiments for all 4 tasks on PhysioNet and 20 ICD-9 code tasks on MIMIC-III using all the GR U models. As shown in T able 1, GR U-D performs best in terms of av erage A UC score across all tasks and in most of the single tasks. 3 . 4 D I S C U S S I O N S Online prediction in early stage Although our model is trained on the first 48 hours data and makes prediction at the last time step, it can be used directly to make predictions before it sees all the time series and can make predictions on the fly . This is v ery useful in applications such as health care, where early decision making is beneficial and critical for patient care. Figure 6 sho ws the online prediction results for MIMIC-III mortality task. As we can see, A UC is around 0.7 at first 12 hours for all the GR U models and it keeps increasing when longer time series is fed into these models. GR U-D and GR U-simple, which explicitly handle missingness, perform consistently 2 Appendix A.3.2 compares all GR U models tested in the experiments in terms of model size. 7 T able 2: Model performances measured by A UC score ( mean ± std ) for mortality prediction. Models MIMIC-III PhysioNet Non-RNN LR-forward 0 . 7589 ± 0 . 015 0 . 7423 ± 0 . 011 SVM-forward 0 . 7908 ± 0 . 006 0 . 8131 ± 0 . 018 RF-forward 0 . 8293 ± 0 . 004 0 . 8183 ± 0 . 015 LR-simple 0 . 7715 ± 0 . 015 0 . 7625 ± 0 . 004 SVM-simple 0 . 8146 ± 0 . 008 0 . 8277 ± 0 . 012 RF-simple 0 . 8294 ± 0 . 007 0 . 8157 ± 0 . 013 RNN LSTM-mean 0 . 8142 ± 0 . 014 0 . 8025 ± 0 . 013 GR U-mean 0 . 8192 ± 0 . 013 0 . 8195 ± 0 . 004 GR U-forward 0 . 8252 ± 0 . 011 0 . 8162 ± 0 . 014 GR U-simple 0 . 8380 ± 0 . 008 0 . 8155 ± 0 . 004 Pr oposed GR U-D 0 . 8527 ± 0 . 003 0 . 8424 ± 0 . 012 0 . 6 9 0 . 7 5 0 . 8 1 0 . 8 7 12 18 24 30 36 42 48 GRU-m ea n GRU-f orward GRU-si mpl e GRU-D SVM-s im ple RF-simple Figure 6: Performance for early predicting mortality on MIMIC-III dataset. x-axis, # of hours after admission; y-axis, A UC score; Dash line, RF-simple results for 48 hours. 0 . 7 3 0 . 7 8 0 . 8 3 0 . 8 8 2k 1 0k 1 9.7k SVM- si mpl e RF -s imp le GRU-mea n GRU -f orwa r d GRU -s i mpl e GRU-D Figure 7: Performance for predicting mortal- ity on subsampled MIMIC-III dataset. x-axis, subsampled dataset size; y-axis, A UC score. superior compared to the other two methods. In addition, GR U-D outperforms GR U-simple when making predictions giv en time series of more than 24 hours, and has at least 2.5% higher A UC score after 30 hours. This indicates that GR U-D is able to capture and utilize long-range temporal missing patterns. Furthermore, GR U-D achie ves similar prediction performance (i.e., same A UC) as best non-RNN baseline model with less time series data. As sho wn in the figure, GR U-D has same A UC performance at 36 hours as the best non-RNN baseline model (RF-simple) at 48 hours. This 12 hour improv ement of GRU-D o ver non-RNN baseline is highly significant in hospital settings such as ICU where time-saving critical decisions demands accurate early predictions. Model Scalability with gro wing data size In many practical applications, model scalability with large dataset size is very important. T o ev aluate the model performance with different training dataset size, we subsample three smaller datasets of 2000 and 10000 admissions from the entire MIMIC-III dataset while keeping the same mortality rate. W e compare our proposed models with all GR U baselines and two most competitiv e non-RNN baselines (SVM-simple, RF-simple). W e observe that all models can achieve impro ved performance gi ven more training samples. Ho wev er, the impro vements of non-RNN baselines are quite limited compared to GR U models, and our GRU-D model achieves the best results on the larger datasets. These results indicate the performance gap between RNN and non-RNN baselines will continue to grow as more data become a v ailable. 4 S U M M A RY In this paper , we proposed novel GR U-based model to effecti vely handle missing values in multiv ariate time series data. Our model captures the informative missingness by incorporating masking and time interv al directly inside the GR U architecture. Empirical experiments on both synthetic and real-w orld health care datasets sho wed promising results and pro vided insightful findings. In our future work, we will explore deep learning approaches to characterize missing-not-at-random data and we will conduct theoretical analysis to understand the behaviors of e xisting solutions for missing values. 8 R E F E R E N C E S Dzmitry Bahdanau, K yunghyun Cho, and Y oshua Bengio. Neural machine translation by jointly learning to align and translate. arXiv preprint , 2014. Y oshua Bengio and Francois Gingras. Recurrent neural networks for missing or asynchronous data. Advances in neural information pr ocessing systems , pp. 395–401, 1996. Zhengping Che, David Kale, W enzhe Li, Mohammad T aha Bahadori, and Y an Liu. Deep computa- tional phenotyping. In SIGKDD , 2015. K yunghyun Cho, Bart V an Merri ¨ enboer , Caglar Gulcehre, Dzmitry Bahdanau, Fethi Bougares, Holger Schwenk, and Y oshua Bengio. Learning phrase representations using rnn encoder-decoder for statistical machine translation. arXiv preprint , 2014. Edward Choi, Mohammad T aha Bahadori, and Jimeng Sun. Doctor ai: Predicting clinical e vents via recurrent neural networks. arXiv pr eprint arXiv:1511.05942 , 2015. Junyoung Chung, Caglar Gulcehre, K yungHyun Cho, and Y oshua Bengio. Empirical e valuation of gated recurrent neural networks on sequence modeling. arXiv pr eprint arXiv:1412.3555 , 2014. Y arin Gal. A theoretically grounded application of dropout in recurrent neural networks. arXiv pr eprint arXiv:1512.05287 , 2015. Pedro J Garc ´ ıa-Laencina, Jos ´ e-Luis Sancho-G ´ omez, and An ´ ıbal R Figueiras-V idal. Pattern classifica- tion with missing data: a revie w . Neural Computing and Applications , 19(2), 2010. Geoffre y Hinton, Li Deng, Dong Y u, George E Dahl, Abdel-rahman Mohamed, Navdeep Jaitly , Andrew Senior , V incent V anhoucke, Patrick Nguyen, T ara N Sainath, et al. Deep neural networks for acoustic modeling in speech recognition: The shared views of four research groups. Signal Pr ocessing Magazine, IEEE , 29(6):82–97, 2012. Sepp Hochreiter and J ¨ urgen Schmidhuber . Long short-term memory . Neural computation , 9(8): 1735–1780, 1997. Serge y Iof fe and Christian Szegedy . Batch normalization: Accelerating deep network training by reducing internal cov ariate shift. arXiv pr eprint arXiv:1502.03167 , 2015. AEW Johnson, TJ Pollard, L Shen, L Lehman, M Feng, M Ghassemi, B Moody , P Szolovits, LA Celi, and RG Mark. Mimic-iii, a freely accessible critical care database. Scientific Data , 2016. Diederik Kingma and Jimmy Ba. Adam: A method for stochastic optimization. arXiv pr eprint arXiv:1412.6980 , 2014. Diederik P Kingma and Max W elling. Auto-encoding variational bayes. In ICLR , 2013. David M Kreindler and Charles J Lumsden. The effects of the irre gular sample and missing data in time series analysis. Nonlinear Dynamical Systems Analysis for the Behavior al Sciences Using Real Data , 2012. Zachary C Lipton, David C Kale, and Randall W etzel. Directly modeling missing data in sequences with rnns: Improved classification of clinical time series. arXiv preprint , 2016. Renata CB Madeo, Clodoaldo AM Lima, and Sarajane M Peres. Gesture unit segmentation using support vector machines: segmenting gestures from rest positions. In SA C , 2013. T omas Mikolov , Martin Karafi ´ at, Lukas Burget, Jan Cernock ` y, and Sanjee v Khudanpur . Recurrent neural network based language model. In INTERSPEECH , volume 2, pp. 3, 2010. Debashis Mondal and Donald B Perciv al. W av elet variance analysis for gappy time series. Annals of the Institute of Statistical Mathematics , 62(5):943–966, 2010. Shahla Parv een and P Green. Speech recognition with missing data using recurrent neural nets. In Advances in Neural Information Pr ocessing Systems , pp. 1189–1195, 2001. 9 F . Pedregosa, G. V aroquaux, A. Gramfort, V . Michel, B. Thirion, O. Grisel, M. Blondel, P . Pretten- hofer , R. W eiss, V . Dubour g, J. V anderplas, A. Passos, D. Cournapeau, M. Brucher , M. Perrot, and E. Duchesnay . Scikit-learn: Machine learning in Python. Journal of Machine Learning Resear ch , 12:2825–2830, 2011. T rang Pham, Truyen T ran, Dinh Phung, and Svetha V enkatesh. Deepcare: A deep dynamic memory model for predictiv e medicine. In Advances in Knowledge Disco very and Data Mining , 2016. Kira Rehfeld, Norbert Marwan, Jobst Heitzig, and J ¨ urgen K urths. Comparison of correlation analysis techniques for irregularly sampled time series. Nonlinear Pr ocesses in Geophysics , 18(3), 2011. Danilo Jimenez Rezende, Shakir Mohamed, and Daan W ierstra. Stochastic backpropagation and approximate inference in deep generativ e models. In ICML , 2014. Donald B Rubin. Inference and missing data. Biometrika , 63(3):581–592, 1976. Joseph L Schafer and John W Graham. Missing data: our view of the state of the art. Psychological methods , 2002. Iv anovitch Silv a, Galan Moody , Daniel J Scott, Leo A Celi, and Roger G Mark. Predicting in-hospital mortality of icu patients: The physionet/computing in cardiology challenge 2012. In CinC , 2012. Nitish Sri vasta va, Geof frey E Hinton, Alex Krizhevsk y , Ilya Sutske ver , and Ruslan Salakhutdinov . Dropout: a simple way to prev ent neural networks from ov erfitting. JMLR , 15(1), 2014. Ilya Sutske ver , Oriol V inyals, and Quoc VV Le. Sequence to sequence learning with neural networks. In Advances in neural information pr ocessing systems , pp. 3104–3112, 2014. V olker T resp and Thomas Briegel. A solution for missing data in recurrent neural networks with an application to blood glucose prediction. NIPS , pp. 971–977, 1998. Y oram V odovotz, Gary An, and Ioannis P Androulakis. A systems engineering perspective on homeostasis and disease. F r ontiers in bioengineering and biotechnology , 1, 2013. Brian J W ells, Ke vin M Chagin, Amy S No wacki, and Michael W Kattan. Strategies for handling missing data in electronic health record deriv ed data. EGEMS , 1(3), 2013. Ian R White, P atrick Royston, and Angela M W ood. Multiple imputation using chained equations: issues and guidance for practice. Statistics in medicine , 30(4):377–399, 2011. Li Zhou and Geor ge Hripcsak. T emporal reasoning with medical dataa revie w with emphasis on medical natural language processing. Journal of biomedical informatics , 40(2):183–202, 2007. 10 A A P P E N D I X A . 1 I N V E S T I G A T I O N O F R E L A T I O N B E T W E E N M I S S I N G N E S S A N D L A B E L S In many time series applications, the pattern of missing v ariables in the time series is often informativ e and useful for prediction tasks. Here, we empirically confirm this claim on real health care dataset by in vestigating the correlation between the missingness and prediction labels (mortality and ICD- 9 diagnosis cate gories). W e denote the missing rate for a v ariable d as p d X and calculate it by p d X = 1 − 1 T P T t =1 m d t . Note that p d X is dependent on mask v ector ( m d t ) and number of time steps T . For each prediction task, we compute the Pearson correlation coefficient between p d X and label ` across all the time series. As shown in Figure 1, we observ e that on MIMIC-III dataset the missing rates with lo w rate values are usually highly (either positiv e or negati ve) correlated with the labels. The distinct correlation between missingness and labels demonstrates usefulness of missingness patterns in solving prediction tasks. A . 2 G R U - D M O D E L V A R I A T I O N S In this section, we will discuss some v ariations of GR U-D model, and also compare some related RNN models which are used for time series with missing data with the proposed model. 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒎 𝒓 MASK 𝜸 𝒙 (a) GR U-DI 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒎 𝒓 MASK 𝜸 𝒉 (b) GR U-DS 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒎 𝒓 MASK 𝜸 𝒎 (c) GR U-DM 𝒉 ෩ 𝒉 𝒛 IN OUT 𝒙 𝒎 𝒓 MASK 𝜸 𝒊 (d) GR U-IMP Figure 8: Graphical illustrations of variations of proposed GR U models. A . 2 . 1 G RU M O D E L W I T H D I FF E R E N T T R A I NA B L E D E C AY S The proposed GRU-D applies trainable decays on both the input and hidden state transitions in order to capture the temporal missing patterns explicitly . This decay idea can be straightforwardly generated to other parts inside the GR U models separately or jointly , giv en different assumptions on the impact of missingness. As comparisons, we also describe and e valuate se veral modifications of GR U-D model. GR U-DI (Figure 8(a)) and GR U-DS (Figure 8(b)) decay only the input and only the hidden state by Equation (5) and (6), respectiv ely . They can be considered as two simplified models of the proposed GR U-D. GRU-DI aims at capturing direct impact of missing values in the data, while GRU-DS captures more indirect impact of missingness. Another intuition comes from this perspecti ve: if an input v ariable is just missing, we should pay more attention to this missingness; howe ver , if an v ariable has been missing for a long time and keeps missing, the missingness becomes less important. W e can utilize this assumption by decaying the masking. This brings us the model GR U-DM shown in Figure 8(c), where we replace the masking m d t fed into GR U-D in by m d t ← m d t + (1 − m d t ) γ m d t (1 − m d t ) = m d t + (1 − m d t ) γ m d t (7) where the equality holds since m d t is either 0 or 1. W e decay the masking for each variable indepen- dently from others by constraining W γ m to be diagonal. A . 2 . 2 G RU - I M P : G OA L - O R I E N T E D I M P U T AT I O N M O D E L W e may alternati vely let the GRU-RNN predict the missing values in the next timestep on its own. When missing values occur only during test time, we simply train the model to predict the measurement vector of the ne xt time step as a language model (Mikolo v et al., 2010) and use it to fill the missing values during test time. This is unfortunately not applicable for some time series applications such as in health care domain, which also hav e missing data during training. 11 Instead, we propose goal-oriented imputation model here called GR U-IMP , and vie w missing values as latent variables in a probabilistic graphical model. Gi ven a timeseries X , we denote all the missing variables by M X and all the observed ones by O X . Then, training a time-series classifier with missing variables becomes equi valent to maximizing the mar ginalized log-conditional probability of a correct label l , i.e., log p ( l |O X ) . The exact marginalized log-conditional probability is ho wev er intractable to compute, and we instead maximize its lowerbound: log p ( l |O X ) = log X M X p ( l |M X , O X ) p ( M X |O X ) ≥ E M X ∼ p ( M X |O X ) log p ( l |M X , O X ) where we assume the distribution ov er the missing variables at each time step is only conditioned on all the previous observ ations: p ( M X |O X ) = T Y t =1 m d t =1 Y 1 ≤ d ≤ D p ( x d t | x 1:( t − 1) , m 1:( t − 1) , δ 1:( t − 1) ) (8) Although this lowerbound is still intractable to compute exactly , we can approximate it by Monte Carlo method, which amounts to sampling the missing v ariables at each time as the RNN reads the input sequence from the beginning to the end, such that x d t ← m d t x d t + (1 − m d t ) ˜ x d t (9) where ˜ x t ∼ x d t | x 1:( t − 1) , m 1:( t − 1) , δ 1:( t − 1) . By further assuming that ˜ x t ∼ N µ t , σ 2 t , µ t = γ t ( W x h t − 1 + b x ) and σ t = 1 , we can use a reparametrization technique widely used in stochastic v ariational inference (Kingma & W elling, 2013; Rezende et al., 2014) to estimate the gradient of the lowerbound ef ficiently . During the test time, we simply use the mean of the missing v ariable, i.e., ˜ x t = µ t , as we have not seen an y impro vement from Monte Carlo approximation in our preliminary experiments. W e vie w this approach as a goal-oriented imputation method and show its structure in Figure 8(d). The whole model is trained to minimize the classification cross-entropy error ` log loss and we tak e the negati ve log likelihood of the observed values as a re gularizer . ` = ` log loss + λ 1 N N X n =1 1 T n T n X t =1 P D d =1 m d t · log p ( x d t | µ d t , σ d t ) P D d =1 m d t (10) A . 2 . 3 C O M PA R I S O N S O F R E L A T E D R N N M O D E L S Sev eral recent works (Lipton et al., 2016; Choi et al., 2015; Pham et al., 2016) use RNNs on EHR data to model diseases and to predict patient diagnosis from health care time series data with irre gular time stamps or missing v alues, but none of them ha ve explicitly attempted to capture and model the missing patterns in their RNNs. Choi et al. (2015) feeds medical codes along with its time stamps into GR U model to predict the next medical event. This feeding time stamps idea is equiv alent to the baseline GR U-simple without feeding the masking, which we denote as GR U-simple (interval only) . Pham et al. (2016) tak es time stamps into LSTM model, and modify its forgetting gate by either time decay and parametric time both from time stamps. Howe ver , their non-trainable decay is not that flexible, and the parametric time also does not change RNN model structure and is similar to GR U-simple (interval only). In addition, neither of them consider missing v alues in time series medical records, and the time stamp input used in these two models is v ector for one patient, b ut not matrix for each input v ariable of one patient as ours. Lipton et al. (2016) achiev es their best performance on diagnosis prediction by feeding masking with zero-filled missing values. Their model is equi valent to GR U-simple without feeding the time interv al, and no model structure modification is made for further capturing and utilizing missingness. W e denote their best model as GR U-simple (masking only) . Conclusively , our GRU-simple baseline can be considered as a generalization from all related RNN models mentioned abov e. 12 A . 3 S U P P L E M E N TA RY E X P E R I M E N T D E TA I L S A . 3 . 1 D A TA S TA T S I T I C S For each of the three datasets used in our experiments, we list the number of samples, the number of input v ariables, the mean and max number of time steps for all the samples, and the mean of all the variable missing rates in T able 3. T able 3: Dataset statistics. MIMIC-III PhysioNet2012 Gesture # of samples ( N ) 19714 4000 378 # of variables ( D ) 99 33 23 Mean of # of time steps 35.89 68.91 21.42 Maximum of # of time steps 150 155 31 Mean of variable missing rate 0.9621 0.8225 N/A A . 3 . 2 G RU M O D E L S I Z E C O M PA R I S O N In order to fairly compare the capacity of all GR U-RNN models, we build each model in proper size so they share similar number of parameters. T able 4 sho ws the statistics of all GR U-based models for on three datasets. W e sho w the statistics for mortality prediction on the two real datasets, and it’ s almost the same for multi-task classifications tasks on these datasets. In addition, ha ving comparable number of parameters also makes all the models have number of iterations and training time close in the same scale in all the experiments. T able 4: Comparison of GR U model size in our experiments. Size refers to the number of hidden states ( h ) in GR U . Models Gesture MIMIC-III PhysioNet 18 input variables 99 input variables 33 input variables Size # of parameters Size # of parameters Size # of parameters GR U-mean&forward 64 16281 100 60105 64 18885 GR U-simple 50 16025 56 59533 43 18495 GR U-D 55 16561 67 60436 49 18838 A . 3 . 3 M U LTI - TA S K P R E D I C T I O N D E TA I L S The RNN models for multi-task learning with m tasks is almost the same as that for binary classi- fication, except that 1) the soft-max prediction layer is replaced by a fully connected layer with n sigmoid logistic functions, and 2) a data-driven prior re gularizer (Che et al., 2015), parameterized by comorbidity (co-occurrence) counts in training data, is applied to the prediction layer to improv e the 0 . 5 5 0 . 6 5 0 . 7 5 0 . 8 5 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 GRU-m ea n G RU-f or wa r d GRU-si mple GRU-D Figure 9: Performance for predicting 20 ICD-9 diagnosis cate gories on MIMIC-III dataset. x-axis, ICD-9 diagnosis category id; y-axis, A UC score. 13 0 . 6 0 . 7 0 . 8 0 . 9 1 m ortal ity lo s < 3 surg ery c ar diac GRU-m ea n GRU-f orw a r d GRU-si mple GRU-D Figure 10: Performance for predicting all 4 tasks on PhysioNet dataset. mortality , in-hospital mortality; los < 3 , length-of-stay less than 3 days; sur gery , whether the patient was recov ering from surgery; car diac , whether the patient had a cardiac condition; y-axis, A UC score. classification performance. W e show the A UC scores for predicting 20 ICD-9 diagnosis categories on MIMIC-III dataset in Figure 9, and all 4 tasks on PhysioNet dataset in Figure 10. The proposed GR U-D achie ves the best av erage A UC score on both datasets and wins 11 of the 20 ICD-9 prediction tasks. A . 3 . 4 E M P I R I C A L C O M PA R I S O N O F M O D E L V A R I A T I O N S Finally , we test all GR U model variations mentioned in Appendix A.2 along with the proposed GR U-D. These include 1) 4 models with trainable decays (GRU-DI, GR U-DS, GR U-DM, GR U-IMP), and 2) two models simplified from GR U-simple (interv al only and masking only). The results are shown in T able 5. As we can see, GR U-D performs best among these models. T able 5: Model performances of GRU variations measured by A UC score ( mean ± std ) for mortality prediction. Models MIMIC-III PhysioNet Baselines GR U-simple (masking only) 0 . 8367 ± 0 . 009 0 . 8226 ± 0 . 010 GR U-simple (interval only) 0 . 8266 ± 0 . 009 0 . 8125 ± 0 . 005 GR U-simple 0 . 8380 ± 0 . 008 0 . 8155 ± 0 . 004 Pr oposed GR U-DI 0 . 8345 ± 0 . 006 0 . 8328 ± 0 . 008 GR U-DS 0 . 8425 ± 0 . 006 0 . 8241 ± 0 . 009 GR U-DM 0 . 8342 ± 0 . 005 0 . 8248 ± 0 . 009 GR U-IMP 0 . 8248 ± 0 . 010 0 . 8231 ± 0 . 005 GR U-D 0 . 8527 ± 0 . 003 0 . 8424 ± 0 . 012 14
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment