Massively Multitask Networks for Drug Discovery
Massively multitask neural architectures provide a learning framework for drug discovery that synthesizes information from many distinct biological sources. To train these architectures at scale, we gather large amounts of data from public sources to…
Authors: Bharath Ramsundar, Steven Kearnes, Patrick Riley
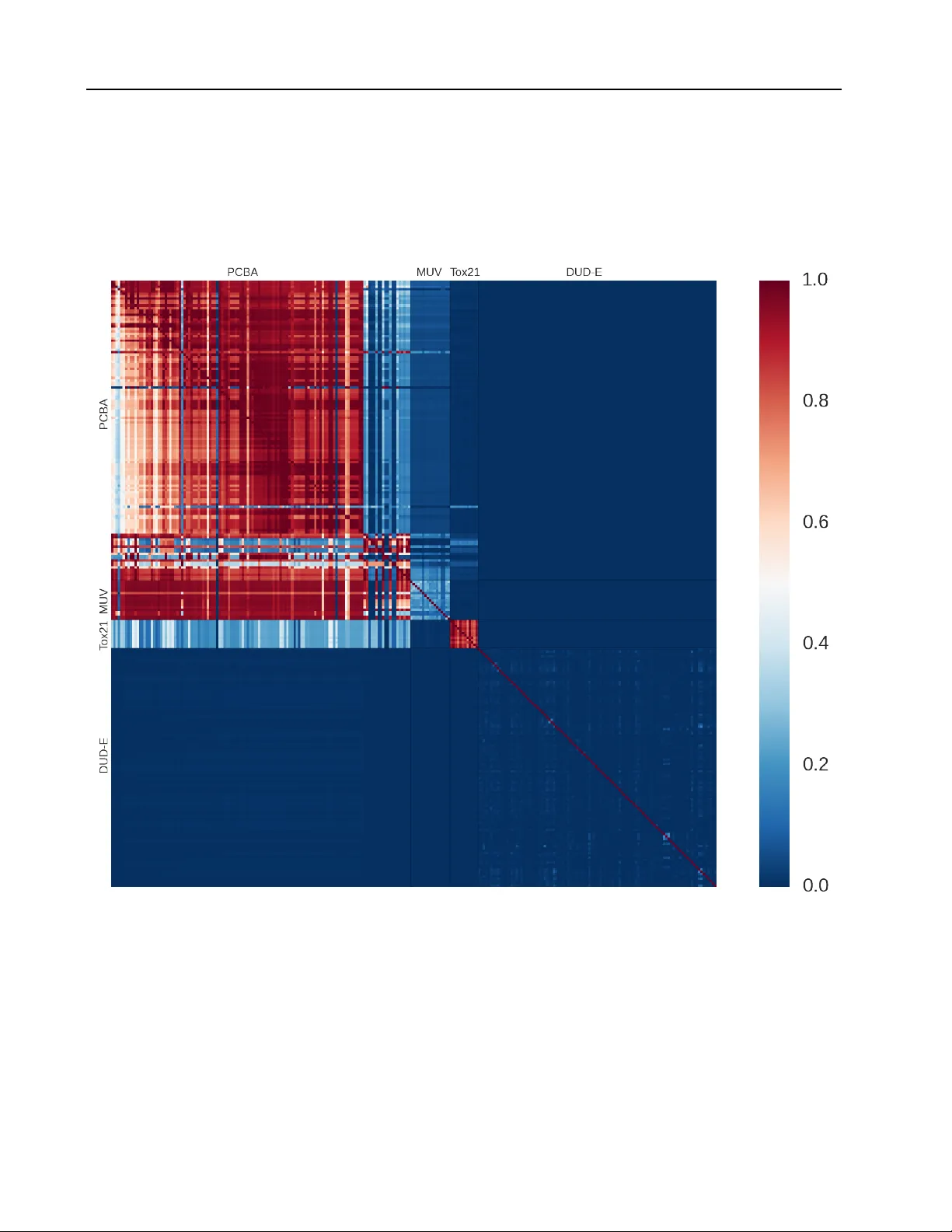
Massiv ely Multitask Networks f or Drug Disco very Bharath Ramsundar *, † , ◦ R B H A R A T H @ S T A N F O R D . E D U Steven K ear nes *, † K E A R N E S @ S T A N F O R D . E D U Patrick Riley ◦ P F R @ G O O G L E . C O M Dale W ebster ◦ D RW @ G O O G L E . C O M David K onerding ◦ D E K @ G O O G L E . C O M V ijay Pande † PA N DE @ S TAN F O R D . E D U ( * Equal contribution, † Stanford Univ ersity , ◦ Google Inc.) Abstract Massiv ely multitask neural architectures provide a learning frame work for drug discov ery that synthesizes information from many distinct bi- ological sources. T o train these architectures at scale, we gather large amounts of data from pub- lic sources to create a dataset of nearly 40 mil- lion measurements across more than 200 bio- logical targets. W e inv estigate several aspects of the multitask framew ork by performing a se- ries of empirical studies and obtain some in- teresting results: (1) massiv ely multitask net- works obtain predicti ve accuracies significantly better than single-task methods, (2) the pre- dictiv e power of multitask networks improves as additional tasks and data are added, (3) the total amount of data and the total number of tasks both contribute significantly to multitask improv ement, and (4) multitask networks afford limited transferability to tasks not in the training set. Our results underscore the need for greater data sharing and further algorithmic innov ation to accelerate the drug discov ery process. 1. Introduction Discov ering new treatments for human diseases is an im- mensely complicated challenge. Prospectiv e drugs must attack the source of an illness, but must do so while sat- isfying restrictive metabolic and toxicity constraints. Tra- ditionally , drug discovery is an e xtended process that takes years to mo ve from start to finish, with high rates of failure along the way . Preliminary w ork. Under revie w by the International Conference on Machine Learning (ICML). Copyright 2015 by the author(s). After a suitable target has been identified, the first step in the drug discovery process is “hit finding. ” Giv en some druggable target, pharmaceutical companies will screen millions of drug-like compounds in an ef fort to find a few attractiv e molecules for further optimization. These screens are often automated via robots, but are expensi ve to perform. V irtual screening attempts to replace or aug- ment the high-throughput screening process by the use of computational methods ( Shoichet , 2004 ). Machine learn- ing methods hav e frequently been applied to virtual screen- ing by training supervised classifiers to predict interactions between targets and small molecules. There are a v ariety of challenges that must be ov ercome to achiev e effecti ve virtual screening. Low hit rates in experimental screens (often only 1 – 2 % of screened com- pounds are activ e against a gi ven target) result in im- balanced datasets that require special handling for effec- tiv e learning. For instance, care must be taken to guard against unrealistic divisions between activ e and inactiv e compounds (“artificial enrichment”) and against informa- tion leakage due to strong similarity between activ e com- pounds (“analog bias”) ( Rohrer & Baumann , 2009 ). Fur- thermore, the paucity of e xperimental data means that o ver - fitting is a perennial thorn. The overall complexity of the virtual screening problem has limited the impact of machine learning in drug dis- cov ery . T o achiev e greater predictiv e power , learning al- gorithms must combine disparate sources of experimental data across multiple tar gets. Deep learning provides a flex- ible paradigm for synthesizing large amounts of data into predictiv e models. In particular, multitask networks facil- itate information sharing across different experiments and compensate for the limited data associated with an y partic- ular experiment. In this work, we in vestig ate several aspects of the multi- task learning paradigm as applied to virtual screening. W e gather a large collection of datasets containing nearly 40 Massively Multitask Netw orks for Drug Disco very million experimental measurements for ov er 200 targets. W e demonstrate that multitask networks trained on this col- lection achie ve significant improvements ov er baseline ma- chine learning methods. W e show that adding more tasks and more data yields better performance. This effect di- minishes as more data and tasks are added, but does not appear to plateau within our collection. Interestingly , we find that the total amount of data and the total number of tasks both hav e significant roles in this improvement. Fur- thermore, the features extracted by the multitask networks demonstrate some transferability to tasks not contained in the training set. Finally , we find that the presence of shared activ e compounds is moderately correlated with multitask improv ement, but the biological class of the tar get is not. 2. Related W orks Machine learning has a rich history in drug discovery . Early work combined creative featurizations of molecules with off-the-shelf learning algorithms to predict drug ac- tivity ( V arnek & Baskin , 2012 ). The state of the art has mov ed to more refined models, such as the influence rele- vance v oting method that combines low-complexity neural networks and k-nearest neighbors ( Sw amidass et al. , 2009 ), and Bayesian belief networks that repurpose textual infor- mation retrie v al methods for virtual screening ( Abdo et al. , 2010 ). Other related work uses deep recursiv e neural net- works to predict aqueous solubility by extracting features from the connectivity graphs of small molecules ( Lusci et al. , 2013 ). Deep learning has made inroads into drug discov ery in recent years, most notably in 2012 with the Merck Kag- gle competition ( Dahl , November 1, 2012 ). T eams were giv en pre-computed molecular descriptors for compounds with experimentally measured activity against 15 targets and were asked to predict the activity of molecules in a held-out test set. The winning team used ensemble models including multitask deep neural networks, Gaussian pro- cess regression, and dropout to improve the baseline test set R 2 by nearly 17%. The winners of this contest later released a technical report that discusses the use of mul- titask networks for virtual screening ( Dahl et al. , 2014 ). Additional work at Merck analyzed the choice of hyper - parameters when training single- and multitask networks and showed improvement ov er random forest models ( Ma et al. , 2015 ). The Merck Kaggle result has been received with skepticism by some in the cheminformatics and drug discov ery communities ( Lo we , December 11, 2012 , and as- sociated comments). T wo major concerns raised were that the sample size was too small (a good result across 15 sys- tems may well hav e occurred by chance) and that any gains in predictiv e accuracy were too small to justify the increase in complexity . While we were preparing this w ork, a workshop paper w as released that also used massiv ely multitask networks for virtual screening ( Unterthiner et al. ). That work curated a dataset of 1,280 biological targets with 2 million asso- ciated data points and trained a multitask network. Their network has more tasks than ours (1,280 vs. 259) but far fewer data points (2 million vs. nearly 40 million). The emphasis of our work is considerably different; while their report highlights the performance gains due to multitask networks, ours is focused on disentangling the underly- ing causes of these improv ements. Another closely related work proposed the use of collaborativ e filtering for vir- tual screening and employed both multitask networks and kernel-based methods ( Erhan et al. , 2006 ). Their multitask networks, ho wever , did not consistently outperform single- task models. W ithin the greater context of deep learning, we draw upon various strands of recent thought. Prior work has used multitask deep networks in the contexts of language understanding ( Collobert & W eston , 2008 ) and multi- language speech recognition ( Deng et al. , 2013 ). Our best-performing netw orks dra w upon design patterns intro- duced by GoogLeNet ( Szegedy et al. , 2014 ), the winner of ILSVRC 2014. 3. Methods 3.1. Dataset Construction and Design Models were trained on 259 datasets gathered from pub- licly av ailable data. These datasets were divided into four groups: PCB A, MUV , DUD-E, and T ox21. The PCB A group contained 128 experiments in the PubChem BioAssay database ( W ang et al. , 2012 ). The MUV group contained 17 challenging datasets specifically designed to av oid common pitfalls in virtual screening ( Rohrer & Bau- mann , 2009 ). The DUD-E group contained 102 datasets that were designed for the ev aluation of methods to pre- dict interactions between proteins and small molecules ( Mysinger et al. , 2012 ). The T ox21 datasets were used in the recent T ox21 Data Challenge ( https://tripod. nih.gov/tox21/challenge/ ) and contained exper- imental data for 12 targets rele v ant to drug toxicity predic- tion. W e used only the training data from this challenge because the test set had not been released when we con- structed our collection. In total, our 259 datasets contained 37 . 8 M experimental data points for 1 . 6 M compounds. De- tails for the dataset groups are giv en in T able 1 . See the Appendix for details on individual datasets and their bio- logical target cate gorization. It should be noted that we did not perform any prepro- cessing of our datasets, such as removing potential ex- perimental artifacts. Such artifacts may be due by com- Massively Multitask Netw orks for Drug Disco very T able 1. Details for dataset groups. V alues for the number of data points per dataset and the percentage of active compounds are reported as means, with standard deviations in parenthesis. Group Datasets Data Points / ea. % Activ e PCB A 128 282 K (122 K ) 1 . 8 (3 . 8) DUD-E 102 14 K (11 K ) 1 . 6 (0 . 2) MUV 17 15 K (1) 0 . 2 (0) T ox21 12 6 K (500) 7 . 8 (4 . 7) pounds whose physical properties cause interference with experimental measurements or allow for promiscuous in- teractions with many targets. A notable exception is the MUV group, which has been processed with consideration of these pathologies ( Rohrer & Baumann , 2009 ). 3.2. Small Molecule Featurization W e used extended connectivity fingerprints (ECFP4) ( Rogers & Hahn , 2010 ) generated by RDKit ( Landrum ) to featurize each molecule. The molecule is decomposed into a set of fragments—each centered at a non-hydrogen atom—where each fragment extends radially along bonds to neighboring atoms. Each fragment is assigned a unique identifier , and the collection of identifiers for a molecule is hashed into a fix ed-length bit vector to construct the molec- ular “fingerprint”. ECFP4 and other fingerprints are com- monly used in cheminformatics applications, especially to measure similarity between compounds ( Willett et al. , 1998 ). A number of molecules (especially in the T ox21 group) failed the featurization process and were not used in training our networks. See the Appendix for details. 3.3. V alidation Scheme and Metrics The traditional approach for model e v aluation is to have fixed training, validation, and test sets. Howe ver , the im- balance present in our datasets means that performance varies widely depending on the particular training/test split. T o compensate for this variability , we used stratified K - fold cross-validation; that is, each fold maintains the ac- tiv e/inactive proportion present in the unsplit data. For the remainder of the paper , we use K = 5 . Note that we did not choose an explicit v alidation set. Sev- eral datasets in our collection have very few actives ( ∼ 30 each for the MUV group), and we feared that selecting a specific validation set would skew our results. As a conse- quence, we suspect that our choice of hyperparameters may be affected by information leakage across folds. Ho wev er , our networks do not appear to be highly sensiti ve to hyper - parameter choice (see Section 4.1 ), so we do not consider leakage to be a serious issue. Follo wing recommendations from the cheminformatics community ( Jain & Nicholls , 2008 ), we used metrics de- riv ed from the receiver operating characteristic (R OC) curve to e valuate model performance. Recall that the ROC curve for a binary classifier is the plot of true positiv e rate (TPR) vs. false positive rate (FPR) as the discrimination threshold is varied. For individual datasets, we are inter- ested in the area under the R OC curve (A UC), which is a global measure of classification performance (note that A UC must lie in the range [0 , 1] ). More generally , for a collection of N datasets, we consider the mean and median K -fold-average A UC: Mean / Median ( 1 K K X k =1 A UC k ( D n ) n = 1 , . . . , N ) , where A UC k ( D n ) is defined as the A UC of a classifier trained on folds { 1 , . . . , K } \ k of dataset D n and tested on fold k . F or completeness, we include in the Appendix an alternativ e metric called “enrichment” that is widely used in the cheminformatics literature ( Jain & Nicholls , 2008 ). W e note that many other performance metrics exist in the literature; the lack of standard metrics makes it difficult to do direct comparisons with previous w ork. 3.4. Multitask Networks A neural network is a nonlinear classifier that performs re- peated linear and nonlinear transformations on its input. Let x i represent the input to the i -th layer of the network (where x 0 is simply the feature vector). The transformation performed is x i +1 = σ ( W i x i + b i ) where W i and b i are respectiv ely the weight matrix and bias for the i -th layer, and σ is a nonlinearity (in our work, the rectified linear unit ( Nair & Hinton , 2010 )). After L such transformations, the final layer of the network x L is then fed to a simple linear classifier , such as the softmax, which predicts the probability that the input x 0 has label j : P ( y = j | x 0 ) = e ( w j ) T x L P M m =1 e ( w m ) T x L , where M is the number of possible labels (here M = 2 ) and w 1 , · · · , w M are weight vectors. W i , b i , and w m are learned during training by the backpropagation algorithm ( Rumelhart et al. , 1988 ). A multitask network attaches N softmax classifiers, one for each task, to the final layer x L . (A “task” corresponds to the classifier associated with a particular dataset in our collection, although we often use “task” and “dataset” interchangeably . See Figure 1 .) 4. Experimental Section In this section, we seek to answer a number of questions about the performance, capabilities, and limitations of mas- Massively Multitask Netw orks for Drug Disco very Input Layer 1024 binary nodes Hidden layers 1-4 layers with 50-3000 nodes Fully connected to layer below, rectified linear activation Softmax nodes, one per dataset Figure 1. Multitask neural network. siv ely multitask neural networks: 1. Do massiv ely multitask networks provide a perfor - mance boost ov er simple machine learning methods? If so, what is the optimal architecture for massively multitask networks? 2. Ho w does the performance of a multitask netw ork de- pend on the number of tasks? How does the perfor- mance depend on the total amount of data? 3. Do massiv ely multitask networks extract generaliz- able information about chemical space? 4. When do datasets benefit from multitask training? The following subsections detail a series of experiments that seek to answer these questions. 4.1. Experimental Exploration of Massively Multitask Networks W e in vestigate the performance of multitask networks with various hyperparameters and compare to sev eral standard machine learning approaches. T able 2 shows some of the highlights of our experiments. Our best multitask archi- tecture (pyramidal multitask networks) significantly out- performed simpler models, including a hypothetical model whose performance on each dataset matches that of the best single-task model (Max { LR, RF , STNN, PSTNN } ). Every model we trained performed extremely well on the DUD-E datasets (all models in T able 2 had median 5 - fold-av erage A UCs ≥ 0 . 99 ), making comparisons between models on DUD-E uninformati ve. For that reason, we exclude DUD-E from our subsequent statistical analysis. Howe ver , we did not remo ve DUD-E from the training alto- gether because doing so adversely af fected performance on the other datasets (data not sho wn); we theorize that DUD- E helped to regularize the classifier and a v oid ov erfitting. During our first explorations, we had consistent problems with the networks o verfitting the data. As discussed in Sec- tion 3.1 , our datasets had a very small fraction of positive examples. For the single hidden layer multitask network in T able 2 , each dataset had 1200 associated parameters. W ith a total number of positives in the tens or hundreds, ov erfitting this number of parameters is a major issue in the absence of strong regularization. Reducing the number of parameters specific to each dataset is the motiv ation for the pyramidal architecture. In our pyramidal networks, the first hidden layer is very wide ( 2000 nodes) with a second narro w hidden layer ( 100 nodes). This dimensionality reduction is similar in moti- vation and implementation to the 1 x 1 con volutions in the GoogLeNet architecture ( Szegedy et al. , 2014 ). The wide lower layer allows for complex, expressi ve features to be learned while the narrow layer limits the parameters spe- cific to each task. Adding dropout of 0 . 25 to our pyramidal networks improved performance. W e also trained single- task versions of our best pyramidal network to understand whether this design pattern works well with less data. T a- ble 2 indicates that these models outperform v anilla single- task networks but do not substitute for multitask training. Results for a variety of alternate models are presented in the Appendix. W e in vestigated the sensitivity of our results to the sizes of the pyramidal layers by running networks with all com- binations of hidden layer sizes: (1000 , 2000 , 3000) and (50 , 100 , 150) . Across the architectures, means and medi- ans shifted by ≤ . 01 A UC with only MUV showing larger changes with a range of . 038 . W e note that performance is sensitiv e to the choice of learning rate and the number of training steps. See the Appendix for details and data. 4.2. Relationship between performance and number of tasks The pre vious section demonstrated that massively multi- task networks improv e performance over single-task mod- els. In this section, we seek to understand how multitask performance is affected by increasing the number of tasks. A priori , there are three reasonable “growth curves” (visu- ally represented in Figure 2 ): Over the hill: performance initially improves, hits a max- imum, then falls. Plateau: performance initially improv es, then plateaus. Still climbing: performance improves throughout, but with a diminishing rate of return. W e constructed and trained a series of multitask networks on datasets containing 10 , 20 , 40 , 80 , 160 , and 249 tasks. These datasets all contain a fixed set of ten “held-in” tasks, which consists of a randomly sampled collection of five Massively Multitask Netw orks for Drug Disco very T able 2. Median 5 -fold-average A UCs for various models. For each model, the sign test in the last column estimates the fraction of datasets (excluding the DUD-E group, for reasons discussed in the text) for which that model is superior to the PMTNN (bottom row). W e use the W ilson score interval to derive a 95% confidence interv al for this fraction. Non-neural network methods were trained using scikit-learn ( Pedregosa et al. , 2011 ) implementations and basic hyperparameter optimization. W e also include results for a hypothetical “best” single-task model (Max { LR, RF , STNN, PSTNN } ) to provide a stronger baseline. Details for our cross-validation and training procedures are giv en in the Appendix. Model PCB A ( n = 128) MUV ( n = 17) T ox21 ( n = 12) Sign T est CI Logistic Regression (LR) . 801 . 752 . 738 [ . 04 , . 13] Random Forest (RF) . 800 . 774 . 790 [ . 06 , . 16] Single-T ask Neural Net (STNN) . 795 . 732 . 714 [ . 04 , . 12] Pyramidal (2000 , 100) STNN (PSTNN) .809 .745 .740 [ . 06 , . 16] Max { LR, RF , STNN, PSTNN } . 824 . 781 . 790 [ . 12 , . 24] 1 -Hidden (1200) Layer Multitask Neural Net (MTNN) . 842 . 797 . 785 [ . 08 , . 18] Pyramidal (2000 , 100) Multitask Neural Net (PMTNN) . 873 . 841 . 818 Figure 2. Potential multitask growth curv es PCB A, three MUV , and tw o T ox21 datasets. These datasets correspond to unique targets that do not have any obvious analogs in the remaining collection. (W e also excluded a similarly chosen set of ten “held-out” tasks for use in Sec- tion 4.4 ). Each training collection is a superset of the pre- ceding collection, with tasks added randomly . For each net- work in the series, we computed the mean 5 -fold-average- A UC for the tasks in the held-in collection. W e repeated this experiment ten times with different choices of random seed. Figure 3 plots the results of our experiments. The shaded region emphasizes the average growth curve, while black dots indicate av erage results for different experimental runs. The figure also displays lines associated with each held-in dataset. Note that several datasets sho w initial dips in performance. Howe ver , all datasets sho w subsequent im- prov ement, and all but one achiev es performance superior to the single-task baseline. Within the limits of our current dataset collection, the distribution in Figure 3 agrees with either plateau or still climbing. The mean performance on the held-in set is still increasing at 249 tasks, so we hypoth- ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● MUV 466 MUV 852 MUV 859 PCBA 485297 PCBA 651644 PCBA 651768 PCBA 743266 PCBA 899 T ox21 NR−Aromatase T ox21 SR−MMP −0.08 −0.04 0.00 0.04 0.08 0.12 10 20 40 80 160 249 # of T asks ∆ A UC from single task NN Figure 3. Held-in growth curves. The y -axis shows the change in A UC compared to a single-task neural network with the same architecture (PSTNN). Each colored curve is the multitask im- prov ement for a given held-in dataset. Black dots represent means across the 10 held-in datasets for each experimental run, where additional tasks were randomly selected. The shaded curve is the mean across the 100 combinations of datasets and experimental runs. esize that performance is still climbing . It is possible that our collection is too small and that an alternate pattern may ev entually emerge. 4.3. More tasks or mor e data? In the previous section we studied the effects of adding more tasks, but here we inv estigate the relativ e importance of the total amount of data vs. the total number of tasks. Namely , is it better to ha ve many tasks with a small amount of associated data, or a small number of tasks with a large amount of associated data? Massively Multitask Netw orks for Drug Disco very W e constructed a series of multitask networks with 10 , 15 , 20 , 30 , 50 and 82 tasks. As in the previous section, the tasks are randomly associated with the networks in a cumulativ e manner ( i.e. , the 82 -task network contained all tasks present in the 50 -task network, and so on). All net- works contained the ten held-in tasks described in the pre- vious section. The 82 tasks chosen were associated with the largest datasets in our collection, each containing 300 K- 500 K data points. Note that all of these tasks belonged to the PCB A group. W e then trained this series of netw orks multiple times with 1 . 6 M, 3 . 3 M, 6 . 5 M, 13 M, and 23 M data points sampled from the non-held-in tasks. W e perform the sampling such that for a given task, all data points present in the first stage ( 1 . 6 M) appeared in the second ( 3 . 3 M), all data points present in the second stage appeared in the third ( 6 . 5 M), and so on. W e decided to use larger datasets so we could sample meaningfully across this entire range. Some com- binations of tasks and data points were not realized; for instance, we did not hav e enough data to train a 20 -task network with 23 M additional data points. W e repeated this experiment ten times using dif ferent random seeds. Figure 4 shows the results of our experiments. The x -axis tracks the number of additional tasks, while the y -axis dis- plays the improvement in performance for the held-in set relativ e to a multitask network trained only on the held-in data. When the total amount of data is fixed, having more tasks consistently yields impro vement. Similarly , when the number of tasks is fixed, adding additional data consis- tently improves performance. Our results suggest that the total amount of data and the total number of tasks both con- tribute significantly to the multitask ef fect. 4.4. Do massively multitask netw orks extract generalizable features? The features extracted by the top layer of the network rep- resent information useful to many tasks. Consequently , we sought to determine the transferability of these features to tasks not in the training set. W e held out ten data sets from the growth curves calculated in Section 4.2 and used the learned weights from points along the growth curv es to ini- tialize single-task networks for the held-out datasets, which we then fine-tuned. The results of training these networks (with 5 -fold strat- ified cross-validation) are shown in Figure 5 . First, note that many of the datasets performed worse than the baseline when initialized from the 10-held-in-task networks. Fur- ther , some datasets nev er exhibited any positiv e effect due to multitask initialization. T ransfer learning can be nega- tiv e. Second, note that the transfer learning effect became ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.00 0.01 0.02 0.03 5 10 20 40 72 # Additional T asks ∆ Mean A UC # Additional Input Examples 23M 13M 6.5M 3.3M 1.6M Figure 4. Multitask benefit from increasing tasks and data inde- pendently . As in Figure 2 , we added randomly selected tasks ( x - axis) to a fixed held-in set. A stratified random sampling scheme was applied to the additional tasks in order to achieve fixed total numbers of additional input examples (color , line type). White points indicate the mean over 10 experimental runs of ∆ mean- A UC over the initial network trained on the 10 held-in datasets. Color-filled areas and error bars describe the smoothed 95% con- fidence intervals. stronger as multitask networks were trained on more data. Large multitask networks exhibited better transferability , but the average ef fect even with 249 datasets was only ∼ . 01 A UC. W e hypothesize that the extent of this gen- eralizability is determined by the presence or absence of relev ant data in the multitask training set. 4.5. When do datasets benefit from multitask training? The results in Sections 4.2 and 4.4 indicate that some datasets benefit more from multitask training than others. In an effort to explain these differences, we consider three specific questions: 1. Do shared activ e compounds explain multitask im- prov ement? 2. Do some biological target classes realize greater mul- titask improv ement than others? 3. Do tasks associated with duplicated targets hav e arti- ficially high multitask performance? 4 . 5 . 1 . S H A R E D A C T I V E C O M P O U N D S The biological context of our datasets implies that active compounds contain more information than inactive com- pounds; while an inactive compound may be inactiv e for many reasons, activ e compounds often rely on similar physical mechanisms. Hence, shared activ e compounds should be a good measure of dataset similarity . Massively Multitask Netw orks for Drug Disco very ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● MUV 548 MUV 832 MUV 846 PCBA 1461 PCBA 2675 PCBA 602233 PCBA 624417 PCBA 652106 T ox21 NR−AhR T ox21 SR−A T AD5 −0.10 −0.05 0.00 0.05 10 20 40 80 160 249 # of T asks/Datasets ∆ A UC from single task NN Figure 5. Held-out growth curves. The y -axis shows the change in A UC compared to a single-task neural network with the same architecture (PSTNN). Each colored curve is the result of initializ- ing a single-task neural network from the weights of the networks from Section 4.2 and computing the mean across the 10 experi- mental runs. These datasets were not included in the training of the original networks. The shaded curve is the mean across the 100 combinations of datasets and experimental runs, and black dots represent means across the 10 held-out datasets for each ex- perimental run, where additional tasks were randomly selected. Figure 6 plots multitask impro vement ag ainst a measure of dataset similarity we call “active occurrence rate” (A OR). For each activ e compound α in dataset D i , A OR i,α is de- fined as the number of additional datasets in which this compound is also activ e: A OR i,α = X d 6 = i 1 ( α ∈ Activ es ( D d )) . Each point in Figure 6 corresponds to a single dataset D i . The x -coordinate is A OR i = Mean α ∈ Activ es ( D i ) ( A OR i,α ) , and the y -coordinate ( ∆ log-odds-mean-A UC) is logit 1 K K X k =1 A UC ( M ) k ( D i ) ! − logit 1 K K X k =1 A UC ( S ) k ( D i ) ! , where A UC ( M ) k ( D i ) and A UC ( S ) k ( D i ) are respectiv ely the A UC values for the k -th fold of dataset i in the multitask and single-task models, and logit ( p ) = log ( p/ (1 − p )) . The use of log-odds reduces the effect of outliers and em- phasizes changes in A UC when the baseline is high. Note that for reasons discussed in Section 4.1 , DUD-E was ex- cluded from this analysis. There is a moderate correlation between A OR and ∆ log- odds-mean-A UC ( r 2 = . 33 ); we note that this correlation is not present when we use ∆ mean-A UC as the y -coordinate ( r 2 = . 09 ). W e hypothesize that some portion of the multi- task ef fect is determined by shared acti ve compounds. That is, a dataset is most likely to benefit from multitask training when it shares many active compounds with other datasets in the collection. Figure 6. Multitask improvement compared to active occurrence rate (A OR). Each point in the figure represents a particular dataset D i . The x -coordinate is the mean A OR across all active com- pounds in D i , and the y -coordinate is the difference in log-odds- mean-A UC between multitask and single-task models. The gray bars indicate standard deviations around the A OR means. There is a moderate correlation ( r 2 = . 33 ). For reasons discussed in Section 4.1 , we excluded DUD-E from this analysis. (Including DUD-E results in a similar correlation, r 2 = . 22 .) 4 . 5 . 2 . T A R G E T C L A S S E S Figure 7 shows the relationship between multitask im- prov ement and target classes. As before, we report mul- titask improv ement in terms of log-odds and exclude the DUD-E datasets. Qualitativ ely , no target class benefited more than any other from multitask training. Nearly ev- ery target class realized gains, suggesting that the multitask framew ork is applicable to experimental data from multiple target classes. 4 . 5 . 3 . D U P L I C ATE T A R G E T S As mentioned in Section 3.1 , there are many cases of tasks with identical targets. W e compared the multitask improve- ment of duplicate vs. unique tasks. The distributions have substantial ov erlap (see the Appendix), but the a verage log- odds improvement was slightly higher for duplicated tasks ( . 531 vs. . 372 ; a one-sided t -test between the duplicate and unique distributions gave p = . 016 ). Since duplicated targets are likely to share man y acti ve compounds, this im- prov ement is consistent with the correlation seen in Sec- Massively Multitask Netw orks for Drug Disco very Figure 7. Multitask improvement across target classes. The x - coordinate lists a series of biological target classes represented in our dataset collection, and the y -coordinate is the dif ference in log-odds-mean-A UC between multitask and single-task models. Note that the DUD-E datasets are excluded. Classes are ordered by total number of tar gets (in parenthesis), and tar get classes with fewer than fi ve members are merged into “miscellaneous. ” tion 4.5.1 . Howe ver , sign tests for single-task vs. multitask models for duplicate and unique tar gets gave significant and highly overlapping confidence intervals ( [0 . 04 , 0 . 24] and [0 . 06 , 0 . 17] , respecti vely; recall that the meaning of these intervals is given in the caption for T able 2 ). T ogether, these results suggest that there is not significant informa- tion leakage within multitask networks. Consequently , the results of our analysis are unlikely to be significantly af- fected by the presence of duplicate targets in our dataset collection. 5. Discussion and Conclusion In this work, we in vestigated the use of massively multitask networks for virtual screening. W e gathered a large collec- tion of publicly av ailable experimental data that we used to train massiv ely multitask neural networks. These net- works achiev ed significant improvement over simple ma- chine learning algorithms. W e explored several aspects of the multitask frame work. First, we demonstrated that multitask performance im- prov ed with the addition of more tasks; our performance was still climbing at 259 tasks. Next, we considered the rel- ativ e importance of introducing more data vs. more tasks. W e found that additional data and additional tasks both contributed significantly to the multitask effect. W e next discov ered that multitask learning afforded limited trans- ferability to tasks not contained in the training set. This ef- fect was not univ ersal, and required large amounts of data ev en when it did apply . W e observed that the multitask effect was stronger for some datasets than others. Consequently , we in vestigated possible explanations for this discrepancy and found that the presence of shared activ e compounds was moderately correlated with multitask improvement, but the biological class of the tar get was not. It is also possible that multitask improv ement results from accurately modeling e xperimen- tal artifacts rather than specific interactions between tar gets and small molecules. W e do not belie ve this to be the case, as we demonstrated strong improvement on the thoroughly- cleaned MUV datasets. The ef ficacy of multitask learning is directly related to the av ailability of relev ant data. Hence, obtaining greater amounts of data is of critical importance for improving the state of the art. Major pharmaceutical companies pos- sess vast priv ate stores of experimental measurements; our work provides a strong argument that increased data shar- ing could result in benefits for all. More data will maximize the benefits achie v able using cur - rent architectures, but in order for algorithmic progress to occur , it must be possible to judge the performance of pro- posed models against previous work. It is disappointing to note that all published applications of deep learning to vir- tual screening (that we are aware of) use distinct datasets that are not directly comparable. It remains to future re- search to establish standard datasets and performance met- rics for this field. Another direction for future work is the further study of small molecule featurization. In this work, we use only one possible featurization (ECFP4), but there exist many others. Additional performance may also be realized by considering targets as well as small molecules in the fea- turization. Y et another line of research could improve per- formance by using unsupervised learning to explore much larger se gments of chemical space. Although deep learning offers interesting possibilities for virtual screening, the full drug discov ery process remains immensely complicated. Can deep learning—coupled with large amounts of experimental data—trigger a re volution in this field? Considering the transformational effect that these methods have had on other fields, we are optimistic about the future. Acknowledgments B.R. was supported by the Fannie and John Hertz Foun- dation. S.K. was supported by a Smith Stanford Graduate Fellowship. W e also acknowledge support from NIH and NSF , in particular NIH U54 GM072970 and NSF 0960306. The latter award was funded under the American Reco very and Rein vestment Act of 2009 (Public Law 111-5). Massively Multitask Netw orks for Drug Disco very References Abdo, Ammar, Chen, Beining, Mueller , Christoph, Salim, Naomie, and W illett, Peter . Ligand-based virtual screen- ing using bayesian networks. J ournal of c hemical infor- mation and modeling , 50(6):1012–1020, 2010. Collobert, Ronan and W eston, Jason. A unified architecture for natural language processing: Deep neural networks with multitask learning. In Pr oceedings of the 25th inter- national conference on Machine learning , pp. 160–167. A CM, 2008. Dahl, George. Deep Learning How I Did It: Merck 1st place interview. No F ree Hunch , Nov ember 1, 2012. Dahl, George E, Jaitly , Navdeep, and Salakhutdinov , Rus- lan. Multi-task neural networks for QSAR predictions. arXiv pr eprint arXiv:1406.1231 , 2014. Deng, Li, Hinton, Geoffrey , and Kingsbury , Brian. New types of deep neural network learning for speech recog- nition and related applications: An overvie w . In Acoustics, Speech and Signal Pr ocessing (ICASSP), 2013 IEEE International Confer ence on , pp. 8599–8603. IEEE, 2013. Erhan, Dumitru, L ’Heureux, Pierre-Jean, Y ue, Shi Y i, and Bengio, Y oshua. Collaborati ve filtering on a family of biological targets. Journal of chemical information and modeling , 46(2):626–635, 2006. Jain, Ajay N and Nicholls, Anthony . Recommenda- tions for ev aluation of computational methods. Journal of computer-aided molecular design , 22(3-4):133–139, 2008. Landrum, Greg. RDKit: Open-source cheminformatics. URL http://www.rdkit.org . Lowe, Derek. Did Kaggle Predict Drug Candidate Activi- ties? Or Not? In the Pipeline , December 11, 2012. Lusci, Alessandro, Pollastri, Gianluca, and Baldi, Pierre. Deep architectures and deep learning in chemoinformat- ics: the prediction of aqueous solubility for drug-like molecules. Journal of chemical information and mod- eling , 53(7):1563–1575, 2013. Ma, Junshui, Sheridan, Robert P , Liaw , Andy , Dahl, George, and Svetnik, Vladimir . Deep neural nets as a method for quantitativ e structure-activity relationships. Journal of Chemical Information and Modeling , 2015. Mysinger , Michael M, Carchia, Michael, Irwin, John J, and Shoichet, Brian K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better bench- marking. J ournal of medicinal chemistry , 55(14):6582– 6594, 2012. Nair , V inod and Hinton, Geoffre y E. Rectified linear units improv e restricted boltzmann machines. In Pr oceedings of the 27th International Conference on Machine Learn- ing (ICML-10) , pp. 807–814, 2010. Pedregosa, Fabian, V aroquaux, Ga ¨ el, Gramfort, Alexan- dre, Michel, V incent, Thirion, Bertrand, Grisel, Olivier , Blondel, Mathieu, Prettenhofer , Peter , W eiss, Ron, Dubourg, V incent, et al. Scikit-learn: Machine learning in python. The Journal of Machine Learning Researc h , 12:2825–2830, 2011. Rogers, David and Hahn, Mathew . Extended-connecti vity fingerprints. J ournal of chemical information and mod- eling , 50(5):742–754, 2010. Rohrer , Sebastian G and Baumann, Knut. Maximum un- biased validation (MUV) data sets for virtual screening based on pubchem bioactivity data. Journal of chemical information and modeling , 49(2):169–184, 2009. Rumelhart, David E, Hinton, Geof frey E, and W illiams, Ronald J. Learning representations by back-propagating errors. Cognitive modeling , 1988. Shoichet, Brian K. V irtual screening of chemical libraries. Natur e , 432(7019):862–865, 2004. Swamidass, S Joshua, Azencott, Chlo ´ e-Agathe, Lin, Ting- W an, Gramajo, Hugo, Tsai, Shiou-Chuan, and Baldi, Pierre. Influence rele vance voting: an accurate and inter - pretable virtual high throughput screening method. Jour - nal of chemical information and modeling , 49(4):756– 766, 2009. Szegedy , Christian, Liu, W ei, Jia, Y angqing, Sermanet, Pierre, Reed, Scott, Anguelov , Dragomir , Erhan, Du- mitru, V anhoucke, V incent, and Rabinovich, Andrew . Going deeper with con volutions. arXiv pr eprint arXiv:1409.4842 , 2014. Unterthiner , Thomas, Mayr , Andreas, ¨ unter Klambauer , G, Steijaert, Marvin, W enger , J ¨ org, Ceulemans, Hugo, and Hochreiter , Sepp. Deep learning as an opportunity in virtual screening. V arnek, Alexandre and Baskin, Igor . Machine learning methods for property prediction in chemoinformatics: quo vadis? Journal of chemical information and model- ing , 52(6):1413–1437, 2012. W ang, Y anli, Xiao, Jewen, Suzek, T ugba O, Zhang, Jian, W ang, Jiyao, Zhou, Zhigang, Han, Lianyi, Karapetyan, Karen, Drachev a, Sv etlana, Shoemaker , Benjamin A, et al. PubChem’ s BioAssay database. Nucleic acids r e- sear ch , 40(D1):D400–D412, 2012. Massively Multitask Netw orks for Drug Disco very W illett, Peter , Barnard, John M, and Downs, Geoffrey M. Chemical similarity searching. Journal of chemical in- formation and computer sciences , 38(6):983–996, 1998. Massi vely Multitask Networks for Drug Disco very: Appendix February 10, 2015 A. Dataset Construction and Design The PCBA datasets are dose-response assays performed by the NCA TS Chemical Genomics Center (NCGC) and down- loaded from PubChem BioAssay using the following search limits: T otalSidCount from 10000, ActiveSidCount from 30, Chemical, Confirmatory , Dose-Response, T arget: Single, NCGC. These limits correspond to the search query: (10000[T otalSidCount] : 1000000000[T otalSidCount]) AND (30[ActiveSidCount] : 1000000000[Activ eSidCount]) AND “small molecule”[filt] AND “doseresponse”[filt] AND 1[T ar getCount] AND “NCGC”[SourceName]. W e note that the DUD-E datasets are especially susceptible to “artificial enrichment” (unrealistic divisions between activ e and inactiv e compounds) as an artifact of the dataset construction procedure. Each data point in our collection was associated with a binary label classifying it as either activ e or inactive. A description of each of our 259 datasets is giv en in T able A1. These datasets cover a wide range of target classes and assay types, including both cell-based and in vitro experiments. Datasets with duplicated tar gets are marked with an asterisk (note that only the non-DUD-E duplicate tar get datasets were used in the analysis described in the text). For the PCB A datasets, compounds not labeled “ Acti ve” were considered inacti ve (including compounds marked “Inconclusi ve”). Due to missing data in PubChem BioAssay and/or featurization errors, some data points and compounds were not used for ev aluation of our models; failure rates for each dataset group are shown in T able A.2 . The T ox21 group suffered especially high failure rates, likely due to the relativ ely large number of metallic or otherwise abnormal compounds that are not supported by the RDKit package. The counts given in T able A1 do not include these missing data. A graphical breakdown of the datasets by target class is shown in Figure A.1 . The datasets used for the held-in and held-out analyses are repeated in T able A.3 and T able A.4 , respectively . As an extension of our treatment of task similarity in the text, we generated the heatmap in Figure A.2 to sho w the pairwise intersection between all datasets in our collection. A few characteristics of our datasets are immediately apparent: • The datasets in the DUD-E group ha ve v ery little intersection with any other datasets. • The PCB A and T ox21 datasets have substantial self-overlap. In contrast, the MUV datasets have relatively little self-ov erlap. • The MUV datasets ha ve substantial o verlap with the datasets in the PCB A group. • The T ox21 datasets have v ery small intersections with datasets in other groups. Figure A.3 shows the ∆ log-odds-mean-A UC for datasets with duplicate and unique targets. Dataset Actives Inactives T arget Class T arget pcba-aid411* 1562 69 734 other enzyme luciferase pcba-aid875 32 73 870 protein-protein interaction brca1-bach1 pcba-aid881 589 106 656 other enzyme 15hLO-2 pcba-aid883 1214 8170 other enzyme CYP2C9 pcba-aid884 3391 9676 other enzyme CYP3A4 pcba-aid885 163 12 904 other enzyme CYP3A4 Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget pcba-aid887 1024 72 140 other enzyme 15hLO-1 pcba-aid891 1548 7836 other enzyme CYP2D6 pcba-aid899 1809 7575 other enzyme CYP2C19 pcba-aid902* 1872 123 512 viability H1299-p53A138V pcba-aid903* 338 54 175 viability H1299-neo pcba-aid904* 528 53 981 viability H1299-neo pcba-aid912 445 68 506 miscellaneous anthrax LF-P A internalization pcba-aid914 218 10 619 transcription fac- tor HIF-1 pcba-aid915 436 10 401 transcription fac- tor HIF-1 pcba-aid924* 1146 122 867 viability H1299-p53A138V pcba-aid925 39 64 358 miscellaneous EGFP-654 pcba-aid926 350 71 666 GPCR TSHR pcba-aid927* 61 59 108 protease USP2a pcba-aid938 1775 70 241 ion channel CNG pcba-aid995* 699 70 189 signalling path- way ERK1/2 cascade pcba-aid1030 15 963 200 920 other enzyme ALDH1A1 pcba-aid1379* 562 198 500 other enzyme luciferase pcba-aid1452 177 151 634 other enzyme 12hLO pcba-aid1454* 536 130 788 signalling path- way ERK1/2 cascade pcba-aid1457 722 204 859 other enzyme IMPase pcba-aid1458 5805 202 680 miscellaneous SMN2 pcba-aid1460* 5662 261 757 protein-protein interaction K18 pcba-aid1461 2305 218 561 GPCR NPSR pcba-aid1468* 1039 270 371 protein-protein interaction K18 pcba-aid1469 169 276 098 protein-protein interaction TRb-SRC2 pcba-aid1471 288 223 321 protein-protein interaction huntingtin pcba-aid1479 788 275 479 miscellaneous TRb-SRC2 pcba-aid1631 892 262 774 other enzyme hPK-M2 pcba-aid1634 154 263 512 other enzyme hPK-M2 pcba-aid1688 2374 218 200 protein-protein interaction HTTQ103 pcba-aid1721 1087 291 649 other enzyme LmPK pcba-aid2100* 1159 301 145 other enzyme alpha-glucosidase pcba-aid2101* 285 321 268 other enzyme glucocerebrosidase pcba-aid2147 3477 223 441 other enzyme JMJD2E pcba-aid2242* 715 198 459 other enzyme alpha-glucosidase pcba-aid2326 1069 268 500 miscellaneous influenza A NS1 pcba-aid2451 2008 285 737 other enzyme FBP A pcba-aid2517 1136 344 762 other enzyme APE1 pcba-aid2528 660 347 283 other enzyme BLM pcba-aid2546 10 550 293 509 transcription fac- tor VP16 pcba-aid2549 1210 233 706 other enzyme RECQ1 Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget pcba-aid2551 16 666 288 772 transcription fac- tor R OR gamma pcba-aid2662 110 293 953 miscellaneous MLL-HO X-A pcba-aid2675 99 279 333 miscellaneous MBNL1-CUG pcba-aid2676 1081 361 124 GPCR RXFP1 pcba-aid463254* 41 330 640 protease USP2a pcba-aid485281 254 341 253 miscellaneous apoferritin pcba-aid485290 942 343 503 other enzyme TDP1 pcba-aid485294* 148 362 056 other enzyme AmpC pcba-aid485297 9126 311 481 promoter Rab9 pcba-aid485313 7567 313 119 promoter NPC1 pcba-aid485314 4491 329 974 other enzyme DN A polymerase beta pcba-aid485341* 1729 328 952 other enzyme AmpC pcba-aid485349 618 321 745 protein kinase A TM pcba-aid485353 603 328 042 protease PLP pcba-aid485360 1485 223 830 protein-protein interaction L3MBTL1 pcba-aid485364 10 700 345 950 other enzyme TGR pcba-aid485367 557 330 124 other enzyme PFK pcba-aid492947 80 330 601 GPCR beta2-AR pcba-aid493208 342 43 647 protein kinase mTOR pcba-aid504327 759 380 820 other enzyme GCN5L2 pcba-aid504332 30 586 317 753 other enzyme G9a pcba-aid504333 15 670 341 165 protein-protein interaction B AZ2B pcba-aid504339 16 857 367 661 protein-protein interaction JMJD2A pcba-aid504444 7390 353 475 transcription fac- tor Nrf2 pcba-aid504466 4169 325 944 viability HEK293T -ELG1-luc pcba-aid504467 7647 322 464 promoter ELG1 pcba-aid504706 201 321 230 miscellaneous p53 pcba-aid504842 101 329 517 other enzyme Mm-CPN pcba-aid504845 104 385 400 miscellaneous RGS4 pcba-aid504847 3515 390 525 transcription fac- tor VDR pcba-aid504891 34 383 652 other enzyme Pin1 pcba-aid540276* 4494 279 673 miscellaneous Marb urg virus pcba-aid540317 2126 381 226 protein-protein interaction HP1-beta pcba-aid588342* 25 034 335 826 other enzyme luciferase pcba-aid588453* 3921 382 731 other enzyme T rxR1 pcba-aid588456* 51 386 206 other enzyme T rxR1 pcba-aid588579 1987 393 298 other enzyme DN A polymerase kappa pcba-aid588590 3936 382 117 other enzyme DN A polymerase iota pcba-aid588591 4715 383 994 other enzyme DN A polymerase eta pcba-aid588795 1308 384 951 other enzyme FEN1 pcba-aid588855 4894 398 438 transcription fac- tor Smad3 pcba-aid602179 364 387 230 other enzyme IDH1 pcba-aid602233 165 380 904 other enzyme PGK Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget pcba-aid602310 310 402 026 protein-protein interaction V if-A3G pcba-aid602313 762 383 076 protein-protein interaction V if-A3F pcba-aid602332 70 415 773 promoter GRP78 pcba-aid624170 837 404 440 other enzyme GLS pcba-aid624171 1239 402 621 transcription fac- tor Nrf2 pcba-aid624173 488 406 224 other enzyme PYK pcba-aid624202 3968 372 045 promoter BRCA1 pcba-aid624246 101 367 273 miscellaneous ERG pcba-aid624287 423 334 388 signalling path- way Gsgsp pcba-aid624288 1356 336 077 signalling path- way Gsgsp pcba-aid624291 222 345 619 promoter a7 pcba-aid624296* 9841 333 378 miscellaneous DN A re-replication pcba-aid624297* 6214 336 050 miscellaneous DN A re-replication pcba-aid624417 6388 398 731 GPCR GLP-1 pcba-aid651635 3784 387 779 promoter A TXN pcba-aid651644 748 361 115 miscellaneous Vpr pcba-aid651768 1677 362 320 other enzyme WRN pcba-aid651965 6422 331 953 protease ClpP pcba-aid652025 238 364 365 signalling path- way IL-2 pcba-aid652104 7126 396 566 miscellaneous TDP-43 pcba-aid652105 4072 324 774 other enzyme PI5P4K pcba-aid652106 496 368 281 miscellaneous alpha-synuclein pcba-aid686970 5949 358 501 viability HT -1080-NT pcba-aid686978* 62 746 354 086 viability DT40-hTDP1 pcba-aid686979* 48 816 368 048 viability DT40-hTDP1 pcba-aid720504 10 170 353 881 protein kinase Plk1 PBD pcba-aid720532* 945 14 532 miscellaneous Marb urg virus pcba-aid720542 733 363 349 protein-protein interaction AMA1-R ON2 pcba-aid720551* 1265 342 387 ion channel KCHN2 3.1 pcba-aid720553* 3260 338 810 ion channel KCHN2 3.1 pcba-aid720579* 1913 304 815 miscellaneous orthopoxvirus pcba-aid720580* 1508 324 844 miscellaneous orthopoxvirus pcba-aid720707 268 364 332 other enzyme EP A C1 pcba-aid720708 661 363 939 other enzyme EP A C2 pcba-aid720709 516 364 084 other enzyme EP A C1 pcba-aid720711 290 364 310 other enzyme EP A C2 pcba-aid743255 902 388 656 protease USP1/U AF1 pcba-aid743266 306 405 368 GPCR PTHR1 muv-aid466 30 14 999 GPCR S1P1 receptor muv-aid548 30 15 000 protein kinase PKA muv-aid600 30 14 999 transcription fac- tor SF1 muv-aid644 30 14 998 protein kinase Rho-Kinase2 muv-aid652 30 15 000 other enzyme HIV R T -RNase muv-aid689 30 14 999 other receptor Eph rec. A4 Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget muv-aid692 30 15 000 transcription fac- tor SF1 muv-aid712* 30 14 997 miscellaneous HSP90 muv-aid713* 30 15 000 protein-protein interaction ER-a-coact. bind. muv-aid733 30 15 000 protein-protein interaction ER-b-coact. bind. muv-aid737* 30 14 999 protein-protein interaction ER-a-coact. bind. muv-aid810* 30 14 999 protein kinase F AK muv-aid832 30 15 000 protease Cathepsin G muv-aid846 30 15 000 protease FXIa muv-aid852 30 15 000 protease FXIIa muv-aid858 30 14 999 GPCR D1 receptor muv-aid859 30 15 000 GPCR M1 receptor tox-NR-AhR 768 5780 transcription fac- tor Aryl hydrocarbon receptor tox-NR-AR-LBD* 237 6520 transcription fac- tor Androgen receptor tox-NR-AR* 309 6955 transcription fac- tor Androgen receptor tox-NR-Aromatase 300 5521 other enzyme Aromatase tox-NR-ER-LBD* 350 6604 transcription fac- tor Estrogen receptor alpha tox-NR-ER* 793 5399 transcription fac- tor Estrogen receptor alpha tox-NR-PP AR-gamma* 186 6263 transcription fac- tor PP ARg tox-SR-ARE 942 4889 miscellaneous ARE tox-SR-A T AD5 264 6807 promoter A T AD5 tox-SR-HSE 372 6094 miscellaneous HSE tox-SR-MMP 919 4891 miscellaneous mitochondrial membrane potential tox-SR-p53 423 6351 miscellaneous p53 signalling dude-aa2ar 482 31 546 GPCR Adenosine A2a receptor dude-abl1 182 10 749 protein kinase T yrosine-protein kinase ABL dude-ace 282 16 899 protease Angiotensin-con verting enzyme dude-aces 453 26 240 other enzyme Acetylcholinesterase dude-ada 93 5450 other enzyme Adenosine deaminase dude-ada17 532 35 900 protease AD AM17 dude-adrb1 247 15 848 GPCR Beta-1 adrenergic receptor dude-adrb2 231 14 997 GPCR Beta-2 adrenergic receptor dude-akt1 293 16 441 protein kinase Serine/threonine-protein kinase AKT dude-akt2 117 6899 protein kinase Serine/threonine-protein kinase AKT2 dude-aldr 159 8999 other enzyme Aldose reductase dude-ampc 48 2850 other enzyme Beta-lactamase dude-andr* 269 14 350 transcription fac- tor Androgen Receptor dude-aofb 122 6900 other enzyme Monoamine oxidase B dude-bace1 283 18 097 protease Beta-secretase 1 Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget dude-braf 152 9950 protein kinase Serine/threonine-protein kinase B- raf dude-cah2 492 31 168 other enzyme Carbonic anhydrase II dude-casp3 199 10 700 protease Caspase-3 dude-cdk2 474 27 850 protein kinase Cyclin-dependent kinase 2 dude-comt 41 3850 other enzyme Catechol O-methyltransferase dude-cp2c9 120 7449 other enzyme Cytochrome P450 2C9 dude-cp3a4 170 11 800 other enzyme Cytochrome P450 3A4 dude-csf1r 166 12 149 other receptor Macrophage colony stimulating factor receptor dude-cxcr4 40 3406 GPCR C-X-C chemokine receptor type 4 dude-def 102 5700 other enzyme Peptide deformylase dude-dhi1 330 19 350 other enzyme 11-beta-hydroxysteroid dehydro- genase 1 dude-dpp4 533 40 943 protease Dipeptidyl peptidase IV dude-drd3 480 34 037 GPCR Dopamine D3 receptor dude-dyr 231 17 192 other enzyme Dihydrofolate reductase dude-egfr 542 35 047 other receptor Epidermal growth factor receptor erbB1 dude-esr1* 383 20 675 transcription fac- tor Estrogen receptor alpha dude-esr2 367 20 190 transcription fac- tor Estrogen receptor beta dude-fa10 537 28 315 protease Coagulation factor X dude-fa7 114 6250 protease Coagulation factor VII dude-fabp4 47 2750 miscellaneous Fatty acid binding protein adipocyte dude-fak1* 100 5350 protein kinase F AK dude-fgfr1 139 8697 other receptor Fibroblast growth factor receptor 1 dude-fkb1a 111 5800 other enzyme FK506-binding protein 1A dude-fnta 592 51 481 other enzyme Protein farnesyltrans- ferase/geranylgeran yltransferase type I alpha subunit dude-fpps 85 8829 other enzyme Farnesyl diphosphate synthase dude-gcr 258 14 999 transcription fac- tor Glucocorticoid receptor dude-glcm* 54 3800 other enzyme glucocerebrosidase dude-gria2 158 11 842 ion channel Glutamate receptor ionotropic dude-grik1 101 6549 ion channel Glutamate receptor ionotropic kainate 1 dude-hdac2 185 10 299 other enzyme Histone deacetylase 2 dude-hdac8 170 10 449 other enzyme Histone deacetylase 8 dude-hivint 100 6650 other enzyme Human immunodeficienc y virus type 1 integrase dude-hivpr 536 35 746 protease Human immunodeficiency virus type 1 protease dude-hivrt 338 18 891 other enzyme Human immunodeficiency virus type 1 rev erse transcriptase dude-hmdh 170 8748 other enzyme HMG-CoA reductase dude-hs90a* 88 4849 miscellaneous HSP90 dude-hxk4 92 4700 other enzyme Hexokinase type IV Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget dude-igf1r 148 9298 other receptor Insulin-like growth factor I recep- tor dude-inha 43 2300 other enzyme Enoyl-[acyl-carrier-protein] reductase dude-ital 138 8498 miscellaneous Leukoc yte adhesion glycoprotein LF A-1 alpha dude-jak2 107 6499 protein kinase T yrosine-protein kinase J AK2 dude-kif11 116 6849 miscellaneous Kinesin-like protein 1 dude-kit 166 10 449 other receptor Stem cell growth factor receptor dude-kith 57 2849 other enzyme Thymidine kinase dude-kpcb 135 8700 protein kinase Protein kinase C beta dude-lck 420 27 397 protein kinase T yrosine-protein kinase LCK dude-lkha4 171 9450 protease Leukotriene A4 hydrolase dude-mapk2 101 6150 protein kinase MAP kinase-activ ated protein ki- nase 2 dude-mcr 94 5150 transcription fac- tor Mineralocorticoid receptor dude-met 166 11 247 other receptor Hepatocyte growth f actor receptor dude-mk01 79 4549 protein kinase MAP kinase ERK2 dude-mk10 104 6600 protein kinase c-Jun N-terminal kinase 3 dude-mk14 578 35 848 protein kinase MAP kinase p38 alpha dude-mmp13 572 37 195 protease Matrix metalloproteinase 13 dude-mp2k1 121 8149 protein kinase Dual specificity mitogen-acti v ated protein kinase kinase 1 dude-nos1 100 8048 other enzyme Nitric-oxide synthase dude-nram 98 6199 other enzyme Neuraminidase dude-pa2ga 99 5150 other enzyme Phospholipase A2 group IIA dude-parp1 508 30 049 other enzyme Poly [ADP-ribose] polymerase-1 dude-pde5a 398 27 547 other enzyme Phosphodiesterase 5A dude-pgh1 195 10 800 other enzyme Cyclooxygenase-1 dude-pgh2 435 23 149 other enzyme Cyclooxygenase-2 dude-plk1 107 6800 protein kinase Serine/threonine-protein kinase PLK1 dude-pnph 103 6950 other enzyme Purine nucleoside phosphorylase dude-ppara 373 19 397 transcription fac- tor PP ARa dude-ppard 240 12 247 transcription fac- tor PP ARd dude-pparg* 484 25 296 transcription fac- tor PP ARg dude-prgr 293 15 648 transcription fac- tor Progesterone receptor dude-ptn1 130 7250 other enzyme Protein-tyrosine phosphatase 1B dude-pur2 50 2698 other enzyme GAR transformylase dude-pygm 77 3948 other enzyme Muscle glycogen phosphorylase dude-pyrd 111 6450 other enzyme Dihydroorotate dehydrogenase dude-reni 104 6958 protease Renin dude-rock1 100 6299 protein kinase Rho-associated protein kinase 1 dude-rxra 131 6948 transcription fac- tor Retinoid X receptor alpha dude-sahh 63 3450 other enzyme Adenosylhomocysteinase dude-src 524 34 491 protein kinase T yrosine-protein kinase SRC Massively Multitask Netw orks for Drug Disco very Dataset Actives Inactives T arget Class T arget dude-tgfr1 133 8500 other receptor TGF-beta receptor type I dude-thb 103 7448 transcription fac- tor Thyroid hormone receptor beta-1 dude-thrb 461 26 999 protease Thrombin dude-try1 449 25 967 protease T rypsin I dude-tryb1 148 7648 protease T ryptase beta-1 dude-tysy 109 6748 other enzyme Thymidylate synthase dude-urok 162 9850 protease Urokinase-type plasminogen acti- vator dude-vgfr2 409 24 946 other receptor V ascular endothelial gro wth f actor receptor 2 dude-wee1 102 6150 protein kinase Serine/threonine-protein kinase WEE1 dude-xiap 100 5149 miscellaneous Inhibitor of apoptosis protein 3 T able A.2. Featurization failures. Group Original Featurized Failure Rate (%) PCB A 439 879 437 928 0 . 44 DUD-E 1 200 966 1 200 406 0 . 05 MUV 95 916 95 899 0 . 02 T ox21 11 764 7830 33 . 44 Massively Multitask Netw orks for Drug Disco very Figure A.1. T arget class breakdo wn. Classes with fewer than fi ve members were mer ged into the “miscellaneous” class. Massively Multitask Netw orks for Drug Disco very T able A.3. Held-in datasets. Dataset Actives Inactives T arget Class T arget pcba-aid899 1809 7575 other enzyme CYP2C19 pcba-aid485297 9126 311 481 promoter Rab9 pcba-aid651644 748 361 115 miscellaneous Vpr pcba-aid651768 1677 362 320 other enzyme WRN pcba-aid743266 306 405 368 GPCR PTHR1 muv-aid466 30 14 999 GPCR S1P1 receptor muv-aid852 30 15 000 protease FXIIa muv-aid859 30 15 000 GPCR M1 receptor tox-NR-Aromatase 300 5521 other enzyme Aromatase tox-SR-MMP 919 4891 miscellaneous mitochondrial membrane potential T able A.4. Held-out datasets. Dataset Acti ves Inactives T arget Class T arget pcba-aid1461 2305 218 561 GPCR NPSR pcba-aid2675 99 279 333 miscellaneous MBNL1-CUG pcba-aid602233 165 380 904 other enzyme PGK pcba-aid624417 6388 398 731 GPCR GLP-1 pcba-aid652106 496 368 281 miscellaneous alpha-synuclein muv-aid548 30 15 000 protein kinase PKA muv-aid832 30 15 000 protease Cathepsin G muv-aid846 30 15 000 protease FXIa tox-NR-AhR 768 5780 transcription factor Aryl hydrocarbon receptor tox-SR-A T AD5 264 6807 promoter A T AD5 Massively Multitask Netw orks for Drug Disco very Figure A.2. Pairwise dataset intersections. The value of the element at position ( x, y ) corresponds to the fraction of dataset x that is contained in dataset y . Thin black lines are used to indicate divisions between dataset groups. Massively Multitask Netw orks for Drug Disco very Figure A.3. Multitask performance of duplicate and unique targets. Outliers are omitted for clarity . Notches indicate a confidence interval around the median, computed as ± 1 . 57 × IQR / √ N ( McGill et al. , 1978 ). Massively Multitask Netw orks for Drug Disco very B. Perf ormance metrics T able B.1. Sign test CIs for each group of datasets. Each model is compared to the Pyramidal (2000 , 100) Multitask Neural Net, .25 Dropout model. Model PCB A ( n = 128) MUV ( n = 17) T ox21 ( n = 12) Logistic Regression (LR) [ . 3 , . 11] [ . 13 , . 53] [ . 00 , . 24] Random Forest (RF) [ . 05 , . 16] [ . 00 , . 18] [ . 14 , . 61] Single-T ask Neural Net (STNN) [ . 02 , . 10] [ . 13 , . 53] [ . 00 , . 24] Pyramidal (2000 , 100) STNN, .25 Dropout (PSTNN) [ . 05 , . 15] [ . 13 , . 53] [ . 00 , . 24] Max { LR, RF , STNN, PSTNN } [ . 09 , . 21] [ . 13 , . 53] [ . 14 , . 61] 1 -Hidden (1200) Layer Multitask Neural Net (MTNN) [ . 05 , . 15] [ . 22 , . 64] [ . 01 , . 35] T able B.2. Enrichment scores for all models reported in T able 2. Each value is the median across the datasets in a group of the mean k -fold enrichment values. Enrichment is an alternate measure of model performance common in virtual drug screening. W e use the “R OC enrichment” definition from ( Jain & Nicholls , 2008 ), but roughly enrichment is the factor better than random that a model’ s top X % predictions are. Model PCB A MUV T ox21 0.5% 1% 2% 5% 0.5% 1% 2% 5% 0.5% 1% 2% 5% LR 19.4 16.5 12.1 7.9 20.0 23.3 15.0 8.0 23.9 18.3 10.6 6.7 RF 40.0 27.4 17.4 9.1 40.0 26.7 16.7 7.3 23.2 19.5 13.6 7.8 STNN 19.0 15.6 11.8 7.7 26.7 20.0 11.7 8.0 16.2 14.4 9.8 6.1 PSTNN 21.8 16.9 12.4 7.9 26.7 16.7 13.3 8.0 23.8 16.1 10.0 6.7 MTNN 33.8 23.6 16.9 9.8 26.7 16.7 16.7 8.7 24.5 18.0 11.4 6.9 PMTNN 43.8 29.6 19.7 11.2 40.0 23.3 16.7 10.0 23.5 18.5 13.7 8.1 Massively Multitask Netw orks for Drug Disco very Figure B.1. Graphical representation of data from T able 2 in the text. Notches indicate a confidence interval around the median, computed as ± 1 . 57 × IQR / √ N ( McGill et al. , 1978 ). Occasionally the notch limits go beyond the quartile markers, producing a “folded down” effect on the boxplot. Paired t -tests (2-sided) relati ve to the PMTNN across all non-DUD-E datasets g av e p ≤ 1 . 86 × 10 − 15 . Massively Multitask Netw orks for Drug Disco very C. T raining Details The multitask networks in T able 2 were trained with learning rate . 0003 and batch size 128 for 50 M steps using stochastic gradient descent. W eights were initialized from a zero-mean Gaussian with standard de viation . 01 . The bias was initialized at . 5 . W e e xperimented with higher learning rates, b ut found that the pyramidal networks sometimes failed to train (the top hidden layer zeroed itself out). Howe ver , this ef fect v anished with the lo wer learning rate. Most of the models were trained with 64 simultaneous replicas sharing their gradient updates, but in some cases we used as man y as 256. The pyramidal single-task networks were trained with the same settings, but for 100 K steps. The vanilla single-task networks were trained with learning rate . 001 for 100 K steps. The networks used in Figure 3 and Figure 4 were trained with learning rate 0 . 003 for 500 epochs plus a constant 3 million steps. The constant factor was introduced after we observed that the smaller multitask networks required more epochs than the lar ger networks to stabilize. The networks in Figure 5 were trained with a Pyramidal (1000, 50) Single T ask architecture (matching the networks in Figure 3). The weights were initialized with the weights from the networks represented in Figure 3 and then trained for 100K steps with a learning rate of 0.0003. As we noted in the main text, the datasets in our collection contained many more inactive than active compounds. T o ensure the actives were giv en adequate importance during training, we weighted the activ es for each dataset to hav e total weight equal to the number of inactiv es for that dataset (inactives were gi ven unit weight). T able C.1 contains the results of our pyramidal model sensitivity analysis. T ables C.2 and C.3 giv e results for a variety of additional models not reported in T able 2. T able C.1. Pyramid sensitivity analysis. Median 5-fold-av erage-A UC values are giv en for se veral variations of the pyramidal architec- ture. In an attempt to avoid the problem of training failures due to the top layer becoming all zero early in the training, the learning rate was set to 0.0001 for the first 2M steps then to 0.0003 for 28M steps. Model PCB A ( n = 128) MUV ( n = 17) T ox21 ( n = 12) Pyramidal (1000 , 50) MTNN . 846 . 825 . 799 Pyramidal (1000 , 100) MTNN . 845 . 818 . 796 Pyramidal (1000 , 150) MTNN . 842 . 812 . 798 Pyramidal (2000 , 50) MTNN . 846 . 819 . 794 Pyramidal (2000 , 100) MTNN . 846 . 821 . 798 Pyramidal (2000 , 150) MTNN . 845 . 839 . 792 Pyramidal (3000 , 50) MTNN . 848 . 801 . 796 Pyramidal (3000 , 100) MTNN . 844 . 804 . 799 Pyramidal (3000 , 150) MTNN . 843 . 810 . 789 Massively Multitask Netw orks for Drug Disco very T able C.2. Descriptions for additional models. MTNN: multitask neural net. “ Auxiliary heads” refers to the attachment of independent softmax units for each task to hidden layers (see Szegedy et al. , 2014 ). Unless otherwise marked, assume 10M training steps. A 8 -Hidden (300) Layer MTNN, auxiliary heads attached to hidden layers 3 and 6, 6M steps B 1 -Hidden (3000) Layer MTNN, 1M steps C 1 -Hidden (3000) Layer MTNN, 1.5M steps D Pyramidal (1800 , 100) , 2 deep, reconnected (original input concatenated to first pyramid output) E Pyramidal (1800 , 100) , 3 deep F 4 -Hidden (1000) Layer MTNN, auxiliary heads attached to hidden layer 2, 4.5M steps G Pyramidal (2000 , 100) MTNN, 10% connected H Pyramidal (2000 , 100) MTNN, 50% connected I Pyramidal (2000 , 100) MTNN, . 001 learning rate J Pyramidal (2000 , 100) MTNN, 50M steps, . 0003 learning rate K Pyramidal (2000 , 100) MTNN, . 25 Dropout (first layer only), 50M steps L Pyramidal (2000 , 100) MTNN, . 25 Dropout, . 001 learning rate T able C.3. Median 5-fold-average A UC values for additional models. Sign test confidence intervals and paired t -test (2-sided) p -values are relativ e to the PMTNN from T able 2 and were calculated across all non-DUD-E datasets. Model PCB A ( n = 128) MUV ( n = 17) T ox21 ( n = 12) Sign T est CI Paired t -T est A . 836 . 793 . 786 [ . 01 , . 06] 9 . 37 × 10 − 43 B . 835 . 855 . 769 [ . 11 , . 22] 1 . 17 × 10 − 17 C . 837 . 851 . 765 [ . 12 , . 24] 2 . 60 × 10 − 16 D . 842 . 842 . 816 [ . 08 , . 18] 1 . 89 × 10 − 21 E . 842 . 808 . 789 [ . 02 , . 08] 9 . 25 × 10 − 43 F . 858 . 836 . 810 [ . 10 , . 22] 4 . 85 × 10 − 13 G . 831 . 795 . 774 [ . 03 , . 11] 1 . 15 × 10 − 31 H . 856 . 827 . 796 [ . 04 , . 13] 5 . 34 × 10 − 21 I . 860 . 862 . 824 [ . 07 , . 17] 6 . 23 × 10 − 14 J . 830 . 810 . 801 [ . 05 , . 14] 9 . 25 × 10 − 25 K . 859 . 843 . 803 [ . 24 , . 38] 3 . 25 × 10 − 9 L . 872 . 837 . 802 [ . 35 , . 50] 2 . 74 × 10 − 2 Massively Multitask Netw orks for Drug Disco very References Jain, Ajay N and Nicholls, Anthony . Recommendations for ev aluation of computational methods. Journal of computer- aided molecular design , 22(3-4):133–139, 2008. McGill, Robert, T uke y , John W , and Larsen, W ayne A. V ariations of box plots. The American Statistician , 32(1):12–16, 1978. Szegedy , Christian, Liu, W ei, Jia, Y angqing, Sermanet, Pierre, Reed, Scott, Anguelov , Dragomir, Erhan, Dumitru, V an- houcke, V incent, and Rabinovich, Andre w . Going deeper with conv olutions. arXiv pr eprint arXiv:1409.4842 , 2014.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment