TimeFlow: Temporal Conditioning for Longitudinal Brain MRI Registration and Aging Analysis
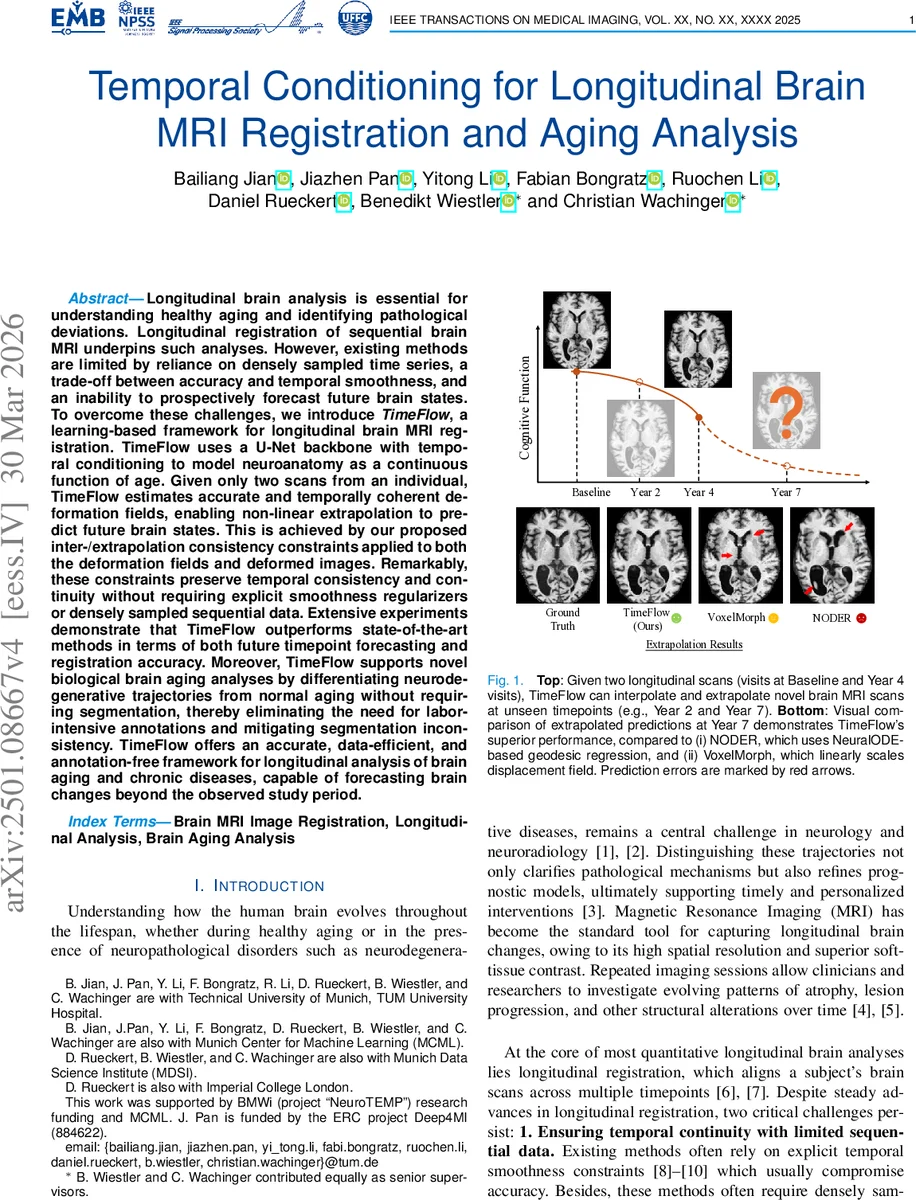
Longitudinal brain analysis is essential for understanding healthy aging and identifying pathological deviations. Longitudinal registration of sequential brain MRI underpins such analyses. However, existing methods are limited by reliance on densely sampled time series, a trade-off between accuracy and temporal smoothness, and an inability to prospectively forecast future brain states. To overcome these challenges, we introduce \emph{TimeFlow}, a learning-based framework for longitudinal brain MRI registration. TimeFlow uses a U-Net backbone with temporal conditioning to model neuroanatomy as a continuous function of age. Given only two scans from an individual, TimeFlow estimates accurate and temporally coherent deformation fields, enabling non-linear extrapolation to predict future brain states. This is achieved by our proposed inter-/extra-polation consistency constraints applied to both the deformation fields and deformed images. Remarkably, these constraints preserve temporal consistency and continuity without requiring explicit smoothness regularizers or densely sampled sequential data. Extensive experiments demonstrate that TimeFlow outperforms state-of-the-art methods in terms of both future timepoint forecasting and registration accuracy. Moreover, TimeFlow supports novel biological brain aging analyses by differentiating neurodegenerative trajectories from normal aging without requiring segmentation, thereby eliminating the need for labor-intensive annotations and mitigating segmentation inconsistency. TimeFlow offers an accurate, data-efficient, and annotation-free framework for longitudinal analysis of brain aging and chronic diseases, capable of forecasting brain changes beyond the observed study period.
💡 Research Summary
TimeFlow addresses two long‑standing challenges in longitudinal brain MRI analysis: (i) achieving temporally coherent registration when only a few scans are available, and (ii) forecasting brain anatomy at future time points that have never been observed. Existing longitudinal registration methods either require densely sampled time series or impose explicit temporal smoothness penalties that trade off spatial accuracy for temporal continuity. Moreover, most of them are limited to interpolation between observed visits and cannot extrapolate beyond the last scan.
The proposed framework solves both problems by learning a continuous deformation field φ₀→L(t) that is conditioned on a scalar age variable t. The core network is a U‑Net whose encoder‑decoder blocks are modulated through Adaptive Instance Normalization (AdaIN). A sinusoidal positional encoding followed by multilayer perceptrons (MLPs) transforms the scalar t into a latent embedding; this embedding controls the mean and variance parameters of AdaIN, thereby injecting temporal information directly into the convolutional pipeline. The model receives only two images – a baseline scan I₀ and a final scan I_L – and can generate φ₀→L(t) for any real‑valued t, enabling interpolation (0 < t < 1) and extrapolation (t > 1 or t < 0) with a single set of learned weights.
Temporal coherence is enforced not by traditional smoothness regularizers but by two novel consistency losses. Interpolation consistency consists of (a) a symmetric image similarity term that aligns both source‑warped and target‑warped images to the same intermediate time point, and (b) a flow‑consistency term that penalizes the deviation between the composition of two consecutive deformations and the direct deformation to the intermediate time. When real intermediate scans are missing, synthetic intermediate times ˆt are uniformly sampled, and the same losses are applied to the generated synthetic pairs, effectively training the network on a dense temporal grid.
Extrapolation consistency extends the same principle beyond the observed interval. By using an intermediate deformation φ₀→k¹ (the deformation from I₀ to a synthetic intermediate scan at time t_k) the model defines an extrapolated deformation φ₀→L(1) = φ₀→k¹ / t_k. An image similarity loss forces the extrapolated warped source to match the true target, while a flow‑consistency loss enforces that φ₀→L(1) and φ₀→k¹ / t_k are identical. Backward versions (t < 0) are defined analogously. These losses together guarantee that the network learns a physically plausible deformation trajectory that can be queried at any time point without additional regularization.
The authors evaluate TimeFlow on large aging and Alzheimer’s disease cohorts (OASIS‑3, ADNI). Quantitative results show superior Dice scores, lower target registration error, and higher structural similarity (SSIM) compared with NeuralODE‑based NODER, VoxelMorph, and classic diffeomorphic registration. Importantly, with only two scans per subject, TimeFlow accurately predicts brain morphology up to 5–10 years ahead, achieving lower L1 error and higher SSIM than competing methods.
Beyond registration, the framework enables annotation‑free aging analysis. By comparing deformation fields of a subject against a healthy control template, the method quantifies subject‑specific atrophy trajectories without requiring tissue segmentations. This reveals accelerated hippocampal and temporal‑lobe shrinkage in MCI and dementia groups, offering a potential early biomarker. Because no manual labels are needed, the approach reduces preprocessing workload and avoids inconsistencies arising from repeated segmentation across visits.
In summary, TimeFlow introduces (1) a temporally‑conditioned U‑Net that learns a continuous deformation function from only two time points, (2) symmetric interpolation and extrapolation consistency losses that replace explicit temporal smoothness regularization, and (3) a registration‑only pipeline for longitudinal brain analysis and future‑state forecasting. The method is data‑efficient, annotation‑free, and capable of both accurate registration and biologically meaningful prediction of brain aging, making it highly relevant for clinical research and personalized neurodegenerative disease monitoring.
Comments & Academic Discussion
Loading comments...
Leave a Comment