RAP: Retrieve, Adapt, and Prompt-Fit for Training-Free Few-Shot Medical Image Segmentation
Few-shot medical image segmentation (FSMIS) has achieved notable progress, yet most existing methods mainly rely on semantic correspondences from scarce annotations while under-utilizing a key property of medical imagery: anatomical targets exhibit r…
Authors: Zhihao Mao, Bangpu Chen
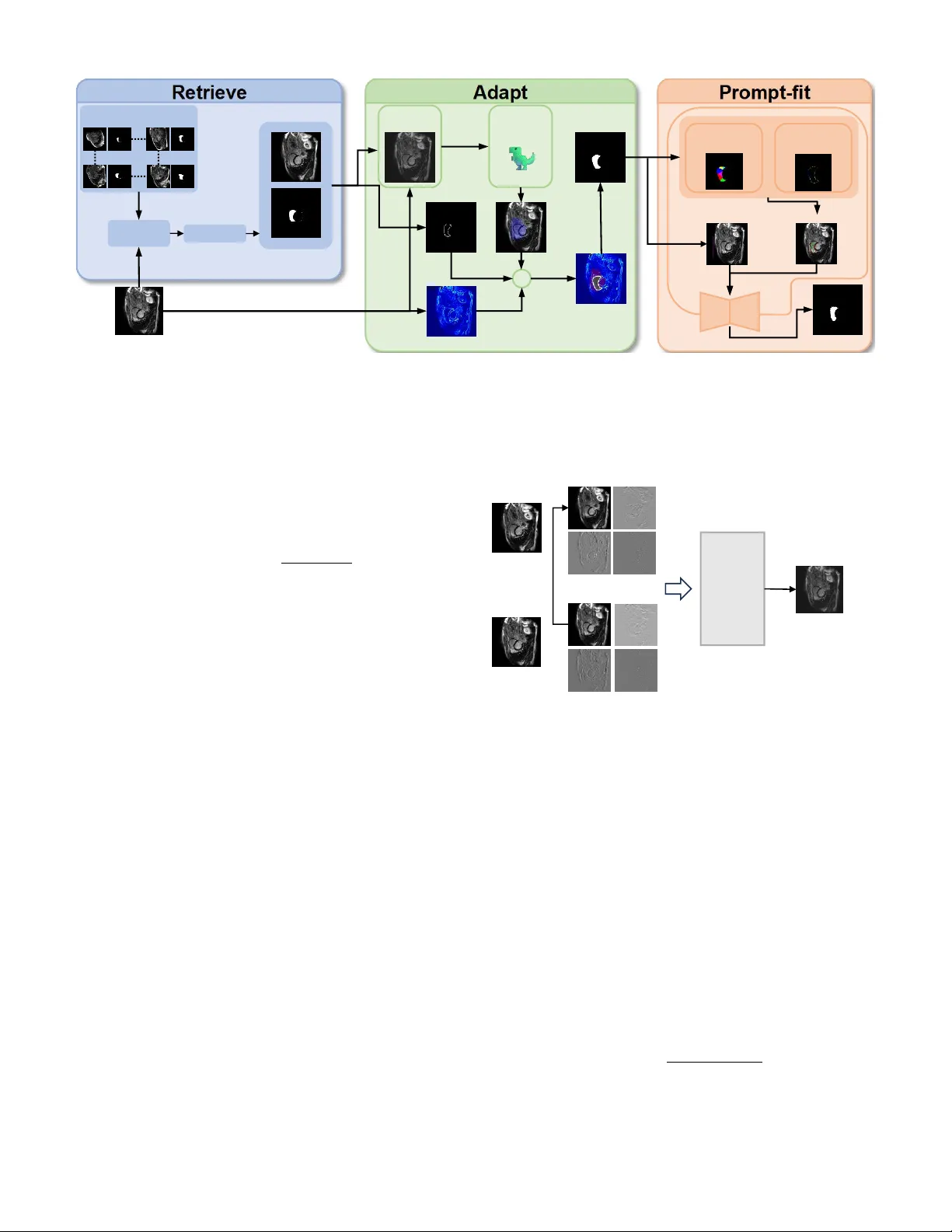
RAP: Retrie v e, Adapt, and Prompt-Fit for T raining-Free Fe w-Shot Medical Image Se gmentation *Zhihao Mao 1 , *Bangpu Chen 1 Abstract —Few-shot medical image segmentation (FSMIS) has achieved notable progress, yet most existing methods mainly rely on semantic correspondences from scarce annotations while under -utilizing a key property of medical imagery: anatomi- cal targets exhibit repeatable high-frequency morphology (e.g., boundary geometry and spatial layout) acr oss patients and acquisitions. W e propose RAP , a training-free framework that Retrieves, Adapts, and Prompt-Fits Segment Anything Model 2 (SAM2) for FSMIS. First, RAP r etrieves morphologically compatible supports from an archive using DINOv3 features to reduce brittleness to single-support choice. Second, it adapts the retrie ved support mask to the query by fitting boundary-aware structural cues, yielding an anatomy-consistent pre-mask under domain shifts. Third, RAP conv erts the pre-mask into prompts by sampling positive points via V or onoi partitioning and negative points via sector-based sampling, and feeds them into SAM2 for final refinement—without any fine-tuning. Extensive experiments on multiple medical segmentation benchmarks show that RAP consistently surpasses prior FSMIS baselines and achieves state- of-the-art performance. Overall, RAP demonstrates that explicit structural fitting combined with r etrieval-augmented prompting offers a simple and effective r oute to robust training-fr ee few-shot medical segmentation. Index T erms —few-shot segmentation, medical image segmen- tation, segment anything model, training-free, shape prior I . I N T R O D U C T I O N Medical image segmentation is fundamental to clinical diagnosis, treatment planning, and disease monitoring [1]. Accurate delineation of anatomical structures such as cardiac chambers, abdominal or gans, and tumors enables quantitative analysis that directly impacts patient outcomes. Deep learning methods ha ve achieved remarkable success in this domain, yet they typically require large amounts of pixel-le vel annotations that are expensi ve and time-consuming to obtain. Few-shot se gmentation has emerged as a promising paradigm to address the annotation scarcity problem, where models learn to segment nov el classes giv en only one or a few labeled support images [2], [3]. Howe ver , existing few-shot methods face critical challenges in medical imaging. First, they often require task-specific training on base classes, limiting their generalization to unseen domains and modalities [4], [5]. Second, the quality of segmentation heavily depends on the visual similarity between support and query images, which is problematic gi ven the high variability in medical scans due to different scanners, protocols, and patient populations. 1 School of Computer Science, China Uni versity of Geosciences, W uhan, China. * Equal contribution. Third, most methods f ail to exploit the strong anatomical priors inherent in medical images, where organs exhibit consistent shapes, positions, and topological structures across patients. More broadly , retriev al-oriented systems suggest that e xternal repositories can complement frozen models without retrain- ing [6], [7]. W e observe that medical images possess two distinctiv e properties that can be le veraged for training-free fe w-shot segmentation. (1) Boundary-centric characteristics : The dis- criminativ e information in medical images concentrates around object boundaries, where intensity gradients and texture pat- terns differ significantly between foreground and background regions. (2) Strong anatomical priors : Organs of the same category share similar shapes and spatial layouts, providing geometric constraints that can guide the segmentation process. Based on these observations, we propose RAP ( R etriev e, A dapt, and P rompt-Fit), a training-free framework for few- shot medical image segmentation. RAP operates in three stages: (1) Retrieve lev erages DINOv3 [8] features to iden- tify the most semantically similar support sample from a database; (2) Adapt employs oriented chamfer matching guided by DINO-based semantic gating to align the sup- port shape onto the query image, producing a preliminary mask; and (3) Pr ompt-Fit generates geometrically-aw are point prompts through V oronoi partitioning [9], which are fed into SAM2 [10] to obtain the final segmentation. Unlike existing methods that require task-specific training or fine-tuning, RAP directly lev erages pre-trained foundation models (DINOv3 and SAM2) without any gradient updates, requiring only a small database of annotated samples for retriev al. Our main contrib utions are summarized as follo ws: • W e propose RAP , a nov el training-free framew ork for few-shot medical image segmentation that synergistically combines vision foundation models with geometric shape priors. • W e introduce an oriented chamfer matching algorithm with DINO-based semantic gating that enables robust shape adaptation from support to query images, effec- tiv ely handling appearance variations across different domains. • W e design a V oronoi-based prompt generation strategy that produces geometrically-aware positiv e and nega- tiv e points, significantly improving SAM2’ s segmentation quality in medical imaging scenarios. • Extensi ve experiments on three medical imaging datasets demonstrate that RAP achiev es competiti ve performance compared to state-of-the-art methods while requiring no training. Upon acceptance, we will release our code and pre-built support databases to facilitate reproducibility and future re- search in training-free medical image segmentation. I I . R E L AT E D W O R K Few-shot Medical Image Segmentation. Few-shot seg- mentation aims to segment novel classes with limited labeled examples. Prototype-based methods [2], [11] e xtract class prototypes from support features and match them against query features. SSL-ALPNet [3] introduces self-supervised learning to improve prototype quality , while RPT [4] partitions foreground into multiple regions for comprehensive feature representation. P A TNet [5], IF A [12], and F AMNet [13] ad- dress cross-domain challenges through domain-in variant fea- tures and frequency-aw are matching. Howe ver , these methods require task-specific training and struggle to generalize across different imaging modalities. Shape Priors in Medical Segmentation. Incorporating shape priors has been a long-standing approach to improve segmentation robustness [14]. Recent methods integrate shape information through shape-aware loss functions [15] or bound- ary attention mechanisms. MA UP [16] proposes a training- free approach using multi-center prompting with V oronoi- based re gion partitioning [9]. Howe ver , e xisting methods either require training to learn shape representations or lack explicit geometric alignment between support and query images. Foundation Models f or Segmentation. SAM [17] and SAM2 [10] hav e demonstrated impressiv e zero-shot segmenta- tion capabilities by lev eraging large-scale pre-training. Sev eral works ha ve explored adapting SAM for medical imaging through prompt engineering [18], [19]. Self-supervised vision transformers like DINOv2 [20] and DINOv3 [8] provide se- mantically rich features for retriev al and correspondence. Our work combines DINOv3 for semantic guidance and SAM2 for segmentation, bridging them through geometric shape adaptation without any training. I I I . P R E L I M I N A RY A. Pr oblem F ormulation W e consider the one-shot medical image segmentation setting. Giv en a support set S = { ( I s , M s ) } containing a single image I s ∈ R H × W with its corresponding binary mask M s ∈ { 0 , 1 } H × W , and a query image I q ∈ R H × W , the goal is to predict the segmentation mask ˆ M q for the query image. In practice, we maintain a support database D = { ( I ( i ) s , M ( i ) s ) } N i =1 containing N labeled samples, from which the most suitable support is retriev ed for each query . B. Notations W e denote query and support images as I q and I s , with M s being the ground-truth mask of the support image and ˆ M q being the predicted mask for the query . The DINO feature maps are denoted as F q , F s ∈ R h × w × d , where h × w is the spatial resolution and d is the feature dimension. W e use E q ∈ R H × W to represent the edge map of the query image, D s ∈ R H × W for the signed distance field computed from M s , and M pre ∈ { 0 , 1 } H × W for the preliminary mask obtained after shape adaptation. The sets of positive and neg ativ e prompt points are denoted as P + and P − , respecti vely . C. F oundation Models DINOv3. DINOv3 [8] is a self-supervised vision trans- former that extends DINOv2 with improved feature repre- sentations. It produces semantically rich patch-level features that capture both local texture and global context. Giv en an image I , DINOv3 extracts features F = DINO ( I ) ∈ R h × w × d , where each spatial location encodes a d -dimensional descrip- tor . These features exhibit strong correspondence properties, making them suitable for cross-image matching and retriev al tasks. Segment Anything Model 2 (SAM2). SAM2 [10] is an upgraded promptable segmentation model that extends SAM with improved architecture and training. SAM2 takes an image and a set of prompts (points, boxes, or masks) as input and produces segmentation masks. Gi ven point prompts P = { ( p i , l i ) } where p i ∈ R 2 denotes the 2D coordinates and l i ∈ { 0 , 1 } indicates positive or negativ e label, SAM2 outputs ˆ M = SAM2 ( I , P ) . The quality of segmentation critically de- pends on the accuracy and placement of prompt points, which motiv ates our geometry-aware prompt generation strategy . I V . M E T H O D O L O G Y A. Overview Fig. 1 illustrates the overall framework of RAP . Giv en a query image I q , our method operates in three stages. (1) Retrieve : W e extract global DINO features and retriev e the most semantically similar support sample ( I s , M s ) from the database. (2) Adapt : W e employ oriented chamfer matching guided by DINO-based semantic gating to align the support mask shape onto the query image, yielding a preliminary mask M pre . (3) Prompt-Fit : W e apply V oronoi partitioning on M pre to generate geometrically-distributed positiv e points, sample negati ve points from boundary sectors, and feed these prompts along with a bounding box to SAM2 for final segmentation. B. Retrieve Stage The retriev al stage aims to find the support sample whose anatomical structure best matches the query . W e le verage DINOv3 to extract semantically meaningful global descriptors. For each image I , we compute the feature map F = DINO ( I ) and obtain the global descriptor by averaging over spatial locations: g = 1 hw X i,j F i,j (1) For support images, we optionally compute a masked descrip- tor using only foreground regions to better capture target- specific features. Query Im age Selected Support Support Pool Frequency Mixing Edge Ma p Pre-mask DINO- scoring Boundary Top 5% ➕ SAM Result Voronoi Partition Boundary Sectors Pos&Neg P oints Bounding Box Matching DINOv3 Encoder Fig. 1: Overview of the proposed RAP framew ork. Giv en a query image, we first retriev e the most similar support from a database using DINOv3 features (Retriev e). Then, we adapt the support shape to the query through oriented chamfer matching with semantic gating (Adapt). Finally , we generate V oronoi-based point prompts and feed them to SAM2 for segmentation (Prompt-Fit). Giv en the query descriptor g q and a database of support descriptors { g ( i ) s } N i =1 , we compute cosine similarities and retriev e the top- k matches: sim ( q , s ( i ) ) = g q · g ( i ) s ∥ g q ∥∥ g ( i ) s ∥ (2) In practice, we select the second-best match (rank-2) to a void potential self-matching artifacts when query and database share similar sources. C. Adapt Stag e The adapt stage transfers the support mask shape to the query image while accounting for geometric variations. This stage consists of three components: frequency-domain style adaptation, DINO-based semantic gating, and oriented cham- fer matching. 1) F r equency-domain Style Adaptation: T o reduce domain shift between support and query images caused by different scanners or imaging protocols, we employ wa velet-based style transfer . As illustrated in Fig. 2, we decompose both images using discrete wav elet transform (DWT) into four subbands: LL (low-frequenc y approximation), LH, HL, and HH (high- frequency details). The lo w-frequency subband captures global appearance such as brightness and contrast, while high- frequency subbands preserve edges and textures. W e replace the support’ s LL subband with the query’ s LL subband and apply in verse wa velet transform to obtain a style-adapted support image: ˜ I s = IDWT ( LL q , LH s , HL s , HH s ) (3) This operation transfers the query’ s global appearance to the support while preserving the support’ s edge structures, which are critical for shape matching. Query Image Support Image LL LH HL HH HH HL LH LL Replace Mixed Support Image Inverse Wavelet Tansfor m Fig. 2: Frequency-domain style adaptation via wa velet trans- form. The support’ s low-frequency (LL) subband is replaced with the query’ s LL to align global appearance while preserv- ing high-frequency edge details. 2) DINO-based Semantic Gating: T o narro w down the search space for shape matching, we compute a semantic similarity map between support and query . W e first cluster the support mask into K regions using K-Means on DINO features: { R k } K k =1 = KMeans ( F s [ M s > 0] , K ) (4) For each re gion, we compute a prototype by av eraging features within the region and measure its similarity to query features: S k ( i, j ) = F q ( i, j ) · c k ∥ F q ( i, j ) ∥∥ c k ∥ (5) where c k is the prototype of region R k . W e select the top- K ′ regions with highest average similarity and aggregate their maps to form a gating mask G : G = [ k ∈ top- K ′ ⊮ [ S k > τ s ] (6) where τ s is a threshold determined by the q -th quantile of similarity v alues. This mask identifies candidate regions in the query where the tar get structure is likely to appear . 3) Oriented Chamfer Matching: W e compute the query edge map E q by fusing multi-scale Laplacian of Gaussian (LoG) responses with Sobel gradients: E q = w LoG · max σ | LoG σ ( I q ) | + w grad · ∥∇ I q ∥ (7) where σ ∈ { 1 , 2 , 4 , 8 } are the LoG scales. From the support mask M s , we extract boundary points { ( x i , y i ) } along with their normal directions { θ i } by comput- ing tangent vectors along the contour . These points form the shape template to be matched against the query . T o enable orientation-aware matching, we partition the query edge pixels into K θ angular bins based on their gradient directions. For each bin k , we compute a distance transform DT k on the corresponding edge pixels: DT k ( x, y ) = min ( x ′ ,y ′ ) ∈ E ( k ) q ∥ ( x, y ) − ( x ′ , y ′ ) ∥ (8) where E ( k ) q denotes edge pixels in angular bin k . W e search for the optimal transformation ( t x , t y , s, r ) (trans- lation, scale, rotation) that minimizes the oriented chamfer distance: C ( t x , t y , s, r ) = 1 |B | X ( x i ,y i ,θ i ) ∈B DT k ( θ i + r ) ( T ( x i , y i )) (9) where B is the support boundary template, T ( · ) applies the transformation, and k ( θ ) maps angle to bin inde x. The search is constrained within the gating mask G to improve efficienc y and robustness. The optimal transformation is applied to the support mask to obtain the preliminary mask: M pre = T ∗ ( M s ) ∩ G (10) D. Pr ompt-F it Stag e The prompt-fit stage conv erts the preliminary mask into effecti ve SAM2 prompts through geometry-aware sampling. 1) V or onoi-based P ositive P oints: Inspired by recent works on region-based prompting [4], [16], we apply Farthest Point Sampling (FPS) on M pre to obtain N v seed points that are maximally spread: V = FPS ( M pre , N v ) (11) These seeds induce a V oronoi partition of the mask region, where each cell V i contains pixels closest to seed v i . For each V oronoi cell, we select its centroid as a positi ve prompt point: P + = { centroid ( V i ) | i = 1 , . . . , N v } (12) This strategy ensures that positi ve points are geometrically distributed across the entire target region, providing compre- hensiv e coverage for SAM2. 2) Sector-based Ne gative P oints: W e partition the boundary of M pre into N s angular sectors centered at the mask centroid. W ithin each sector , we sample negati ve points from the exterior region based on DINO dissimilarity scores: P − = N s [ j =1 arg min p ∈ Sector j \ M pre S DINO ( p ) (13) where S DINO ( p ) is the semantic similarity at point p . This ensures negati ve points are distrib uted around the object boundary with lo w similarity to the foreground. 3) SAM2 Infer ence: W e combine the positive points P + , negati ve points P − , and a bounding box B box deriv ed from M pre as the complete prompt set: ˆ M q = SAM2 ( I q , P + , P − , B box ) (14) The bounding box provides a spatial prior that constrains SAM2’ s attention to the region of interest, while the point prompts of fer fine-grained foreground/background guidance. V . E X P E R I M E N T S A. Datasets W e e valuate our method on three widely-used medical image segmentation benchmarks spanning dif ferent modalities and anatomical structures. Abd-MRI [21] consists of 20 cases of abdominal MRI scans from the ISBI 2019 Combined Healthy Abdominal Organ Segmentation (CHA OS) challenge, with annotations for four abdominal organs: li ver , left kidney (LK), right kidney (RK), and spleen. Abd-CT [22] contains 20 cases of abdominal CT scans from the MICCAI 2015 Multi-Atlas Labeling Beyond the Cranial V ault (BTCV) chal- lenge, with the same four organs annotated. Card-MRI [23] comprises 45 cases of cardiac MRI scans from the MICCAI 2019 Multi-Sequence Cardiac MRI Segmentation Challenge, with three cardiac structures annotated: left ventricle blood pool (L V -BP), left ventricle myocardium (L V -MYO), and right ventricle (R V). B. Evaluation Metrics Follo wing prior works [4], [16], we adopt the Dice Sim- ilarity Coefficient (DSC) as the primary ev aluation metric: DSC = 2 | P ∩ G | | P | + | G | × 100% , where P and G denote the predicted and ground-truth masks, respectiv ely . All experiments are conducted under the 1-way 1-shot setting to simulate the scarcity of labeled data in medical scenarios. C. Implementation Details W e employ the pre-trained DINOv3-V iT -L/16 [8] as the feature extractor and SAM2-V iT -H [10] as the segmentation backbone. All images are resized to 512 × 512 for DINO feature extraction. For oriented chamfer matching, we use K θ = 8 angular bins, scales s ∈ { 0 . 6 , 0 . 7 , . . . , 1 . 4 } , and rotations r ∈ {− 20 ◦ , − 10 ◦ , 0 ◦ , 10 ◦ , 20 ◦ } . The DINO gating T ABLE I: Quantitativ e comparison (Dice %) on Abd-MRI and Abd-CT datasets. Method Abd-MRI Abd-CT Liver LK RK Spleen Mean Liver LK RK Spleen Mean P ANet 39.24 26.47 37.35 26.79 32.46 40.29 30.61 26.66 30.21 31.94 SSL-ALPNet 70.74 55.49 67.43 58.39 63.01 71.38 34.48 32.32 51.67 47.46 RPT 49.22 42.45 47.14 48.84 46.91 65.87 40.07 35.97 51.22 48.28 P A TNet 57.01 50.23 53.01 51.63 52.97 75.94 46.62 42.68 63.94 57.29 IF A 50.22 35.99 34.00 42.21 40.61 46.62 25.13 26.56 24.85 30.79 F AMNet 73.01 57.28 74.68 58.21 65.79 73.57 57.79 61.89 65.78 64.75 MA UP 78.16 58.23 72.34 59.65 67.09 78.25 59.41 71.80 60.38 67.46 RAP (Ours) 79.84 60.28 72.06 64.46 69.16 80.16 60.62 69.54 67.93 69.56 threshold is set to the 90th percentile of similarity values. For V oronoi-based prompt generation, we sample N v = 6 positiv e points via FPS and N s = 8 negati ve points from boundary sectors. During retriev al, we select the rank-2 match instead of rank-1 to a void potential self-matching when query and support slices originate from the same patient volume in leav e- one-out ev aluation. All experiments are conducted on a single NVIDIA R TX 3090 GPU with 24GB memory using PyT orch. D. Compar ed Methods W e compare RAP with se veral state-of-the-art fe w-shot medical image segmentation methods. The training-based methods include P ANet [2], SSL-ALPNet [3], RPT [4], P A T - Net [5], IF A [12], and F AMNet [13], all of which require train- ing on source medical domains. For training-free comparison, we include MA UP [16], which leverages DINOv2 and SAM without gradient updates. V I . R E S U LT S A N D D I S C U S S I O N A. Comparison with State-of-the-art T able I presents the quantitative comparison on Abd-MRI and Abd-CT datasets. Our RAP achieves the best average Dice scores of 69.16% on Abd-MRI and 69.56% on Abd-CT , outperforming all compared methods. Notably , RAP surpasses the recent MA UP [16] by 2.07% and 2.10% on the tw o datasets, respecti vely , v alidating the ef fectiveness of our shape- guided approach. T able II shows the results on the Card-MRI dataset. RAP achiev es an av erage Dice of 75.27%, improving upon MA UP by 2.14%. The improv ement is particularly significant on the challenging L V -MYO structure (+1.27%) and R V (+3.58%), where our oriented chamfer matching effecti vely captures the thin myocardium boundaries. B. Ablation Study W e conduct ablation studies on the R V structure of Card- MRI to validate the contribution of each component in RAP . As shown in T able III, each module progressiv ely improves the segmentation performance. Effect of OCM. Adding Oriented Chamfer Matching im- prov es Dice from 78.29% to 80.17% (+1.88%), demonstrating the importance of orientation-a ware shape alignment. Effect of SG. DINO-based Semantic Gating further im- prov es to 81.02% (+0.85%) by constraining the search space and pre venting false matches. T ABLE II: Quantitative comparison (Dice %) on Card-MRI dataset. Method L V -BP L V -MY O R V Mean P ANet 51.42 25.75 25.75 36.66 SSL-ALPNet 83.47 22.73 66.21 57.47 RPT 60.84 42.28 57.30 53.47 P A TNet 65.35 50.63 68.34 61.44 IF A 50.43 31.32 30.74 37.50 F AMNet 86.64 51.82 76.26 71.58 MA UP 88.36 52.74 78.29 73.13 RAP (Ours) 89.92 54.01 81.87 75.27 T ABLE III: Ablation study on Card-MRI (R V structure). OCM: Oriented Chamfer Matching, SG: Semantic Gating, VP: V oronoi-based Prompts. OCM SG VP Mean Dice (%) 78.29 ✓ 80.17 ✓ ✓ 81.02 ✓ ✓ ✓ 81.87 Effect of VP . V oronoi-based Prompts achieve 81.87% (+0.85%) through comprehensive geometric cov erage of the target region. C. Qualitative Analysis Fig. 3 presents qualitativ e comparisons on the Card-MRI dataset for three cardiac structures (L V -BP , L V -MY O, and R V). Our RAP produces more accurate boundaries compared to baseline methods, especially for the thin myocardium (L V - MY O) where precise boundary delineation is critical. The V oronoi-based prompts effecti vely guide SAM2 to segment complete structures without missing thin regions or including spurious areas. D. Discussion W e clarify that “training-free” refers to requiring no gradient-based optimization or weight updates during infer- ence. While RAP utilizes a pre-built support database for retriev al, this database only stores raw images and masks without learned representations, distinguishing our approach from methods that require task-specific training on source domains. RAP of fers sev eral key adv antages. First, it enables imme- diate deployment to new domains without retraining. Second, the shape-guided approach is robust to appearance variations across dif ferent scanners and protocols. Third, the oriented chamfer matching effecti vely handles gradient direction infor- mation for subtle boundaries. Suppor t LV-BP LV-MYO RV Ground Truth RPT FAMNet MAUP RAP Fig. 3: Qualitative results on Card-MRI for three cardiac structures. Our method produces more accurate boundaries, especially for the thin myocardium (L V -MYO). Regarding the support database, our experiments use leav e- one-out ev aluation where each test case retriev es from remain- ing samples. The retriev al mechanism is robust to database variations: since DINOv3 features capture semantic similarity rather than exact pixel matching, morphologically compatible supports can be retrieved ev en from small databases (20-45 cases in our benchmarks). Howe ver , performance may degrade for rare anatomical structures or pathological cases that deviate significantly from normal anatomy in the database. V I I . C O N C L U S I O N W e presented RAP , a training-free frame work for few- shot medical image segmentation that synergistically com- bines vision foundation models with geometric shape pri- ors. Our method introduces oriented chamfer matching with DINO-based semantic gating for rob ust shape adaptation, and V oronoi-based prompt generation for geometry-aware SAM2 guidance. Experiments on three datasets demonstrate that RAP achiev es competiti ve performance compared to training-based methods while requiring no task-specific optimization. Future work includes extending RAP to 3D volumetric segmentation by le veraging inter-slice consistency . A C K N O W L E D G M E N T S R E F E R E N C E S [1] R. Azad, E. K. Aghdam, A. Rauland, Y . Jia, A. H. A vval, A. Bozor gpour, S. Karimijafarbigloo, J. P . Cohen, E. Adeli, and D. Merhof, “Medical image segmentation review: The success of u-net, ” IEEE T ransactions on P attern Analysis and Machine Intelligence , vol. 46, no. 12, pp. 10 076– 10 095, 2024. [2] K. W ang, J. H. Liew , Y . Zou, D. Zhou, and J. Feng, “Panet: Few-shot image semantic segmentation with prototype alignment, ” in proceedings of the IEEE/CVF international conference on computer vision , 2019, pp. 9197–9206. [3] C. Ouyang, C. Biffi, C. Chen, T . Kart, H. Qiu, and D. Rueckert, “Self- supervised learning for few-shot medical image segmentation, ” IEEE T ransactions on Medical Imaging , vol. 41, no. 7, pp. 1837–1848, 2022. [4] Y . Zhu, S. W ang, T . Xin, and H. Zhang, “Few-shot medical image segmentation via a region-enhanced prototypical transformer, ” in In- ternational Conference on Medical Image Computing and Computer- Assisted Intervention . Springer , 2023, pp. 271–280. [5] S. Lei, X. Zhang, J. He, F . Chen, B. Du, and C.-T . Lu, “Cross-domain few-shot semantic segmentation, ” in European Confer ence on Computer V ision , 2022, pp. 73–90. [6] M. Li, K. Li, Y . Liu, J. Chen, Z. Zheng, Y . W u, and X. Chen, “Hi- erarchical scheduling for multi-vector image retrie val, ” arXiv pr eprint arXiv:2510.08976 , 2025. [7] Z. Zheng, Z. Mao, S. Tian, M. Li, J. Chen, X. Sun, Z. Zhang, X. Liu, D. Cao, H. Mei et al. , “Heisd: Hybrid speculativ e decoding for embodied vision-language-action models with kinematic awareness, ” arXiv pr eprint arXiv:2603.17573 , 2026. [8] O. Sim ´ eoni, H. V . V o, M. Seitzer, F . Baldassarre, M. Oquab, C. Jose, V . Khalidov , M. Szafraniec, S. Y i, M. Ramamonjisoa, F . Massa, D. Haziza, L. W ehrstedt, J. W ang, T . Darcet, T . Moutakanni, L. Sentana, C. Roberts, A. V edaldi, J. T olan, J. Brandt, C. Couprie, J. Mairal, H. J ´ egou, P . Labatut, and P . Bojanowski, “Dinov3, ” 2025. [Online]. A vailable: https://arxiv .org/abs/2508.10104 [9] F . Aurenhammer , “V oronoi diagrams: a surve y of a fundamental geo- metric data structure, ” A CM Computing Surve ys (CSUR) , vol. 23, no. 3, pp. 345–405, 1991. [10] N. Ravi, V . Gabeur, Y .-T . Hu, R. Hu, C. Ryali, T . Ma, H. Khedr, R. R ¨ adle, C. Rolland, L. Gustafson, E. Mintun, J. Pan, K. V . Alwala, N. Carion, C.-Y . Wu, R. Girshick, P . Doll ´ ar , and C. Feichtenhofer, “Sam 2: Segment anything in images and videos, ” 2024. [Online]. A vailable: https://arxiv .org/abs/2408.00714 [11] J. Snell, K. Swersky , and R. Zemel, “Prototypical networks for fe w-shot learning, ” Advances in neural information processing systems , vol. 30, 2017. [12] J. Nie, Y . Xing, G. Zhang, P . Y an, A. Xiao, Y .-P . T an, A. C. Kot, and S. Lu, “Cross-domain few-shot segmentation via iterative support-query correspondence mining, ” IEEE/CVF Conference on Computer V ision and P attern Recognition , pp. 3380–3390, 2024. [13] Y . Bo, Y . Zhu, L. Li, and H. Zhang, “F amnet: Frequency-a ware matching network for cross-domain few-shot medical image segmentation, ” in AAAI Confer ence on Artificial Intelligence , 2025. [14] H. Kerv adec, J. Bouchtiba, C. Desrosiers, E. Granger, J. Dolz, and I. B. A yed, “Boundary loss for highly unbalanced segmentation, ” in International conference on medical imaging with deep learning , 2019, pp. 285–296. [15] J. Ma, J. Chen, M. Ng, R. Huang, Y . Li, C. Li, X. Y ang, and A. L. Martel, “Loss odyssey in medical image segmentation, ” Medical Image Analysis , vol. 71, p. 102035, 2021. [16] Y . Zhu and H. Zhang, “ MA UP: Training-free Multi-center Adaptiv e Uncertainty-aware Prompting for Cross-domain Few-shot Medical Im- age Segmentation , ” in proceedings of Medical Imag e Computing and Computer Assisted Intervention – MICCAI 2025 , vol. LNCS 15966. Springer Nature Switzerland, September 2025. [17] A. Kirillov , E. Mintun, N. Ravi, H. Mao, C. Rolland, L. Gustafson, T . Xiao, S. Whitehead, A. C. Berg, W .-Y . Lo et al. , “Segment anything, ” in Proceedings of the IEEE/CVF international conference on computer vision , 2023, pp. 4015–4026. [18] K. Huang, T . Zhou, H. Fu, Y . Zhang, Y . Zhou, C. Gong, and D. Liang, “Learnable prompting sam-induced knowledge distillation for semi- supervised medical image segmentation, ” IEEE T ransactions on Medical Imaging , pp. 1–1, 2025. [19] R. W ang, Z. Y ang, and Y . Song, “Osam-fundus: A training-free, one- shot segmentation framew ork for optic disc and cup in fundus images, ” Biomedical Signal Processing and Contr ol , vol. 100, p. 107069, 2025. [20] M. Oquab, T . Darcet, T . Moutakanni, H. V o, M. Szafraniec, V . Khalidov , P . Fernandez, D. Haziza, F . Massa, A. El-Nouby et al. , “Dinov2: Learning robust visual features without supervision, ” arXiv preprint arXiv:2304.07193 , 2023. [21] A. E. Kavur , N. S. Gezer, M. Barıs ¸, S. Aslan, P .-H. Conze, V . Groza, D. D. Pham, S. Chatterjee, P . Ernst, S. ¨ Ozkan et al. , “Chaos challenge- combined (ct-mr) healthy abdominal organ segmentation, ” Medical Image Analysis , vol. 69, p. 101950, 2021. [22] B. Landman, Z. Xu, J. Igelsias, M. Styner , T . Langerak, and A. Klein, “Miccai multi-atlas labeling beyond the cranial vault–workshop and challenge, ” in Proceedings of the MICCAI—W orkshop Challenge , vol. 5, 2015, p. 12. [23] X. Zhuang, “Multiv ariate mixture model for myocardial segmentation combining multi-source images, ” IEEE tr ansactions on pattern analysis and machine intelligence , vol. 41, no. 12, pp. 2933–2946, 2018.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment