Tabular LLMs for Interpretable Few-Shot Alzheimer's Disease Prediction with Multimodal Biomedical Data
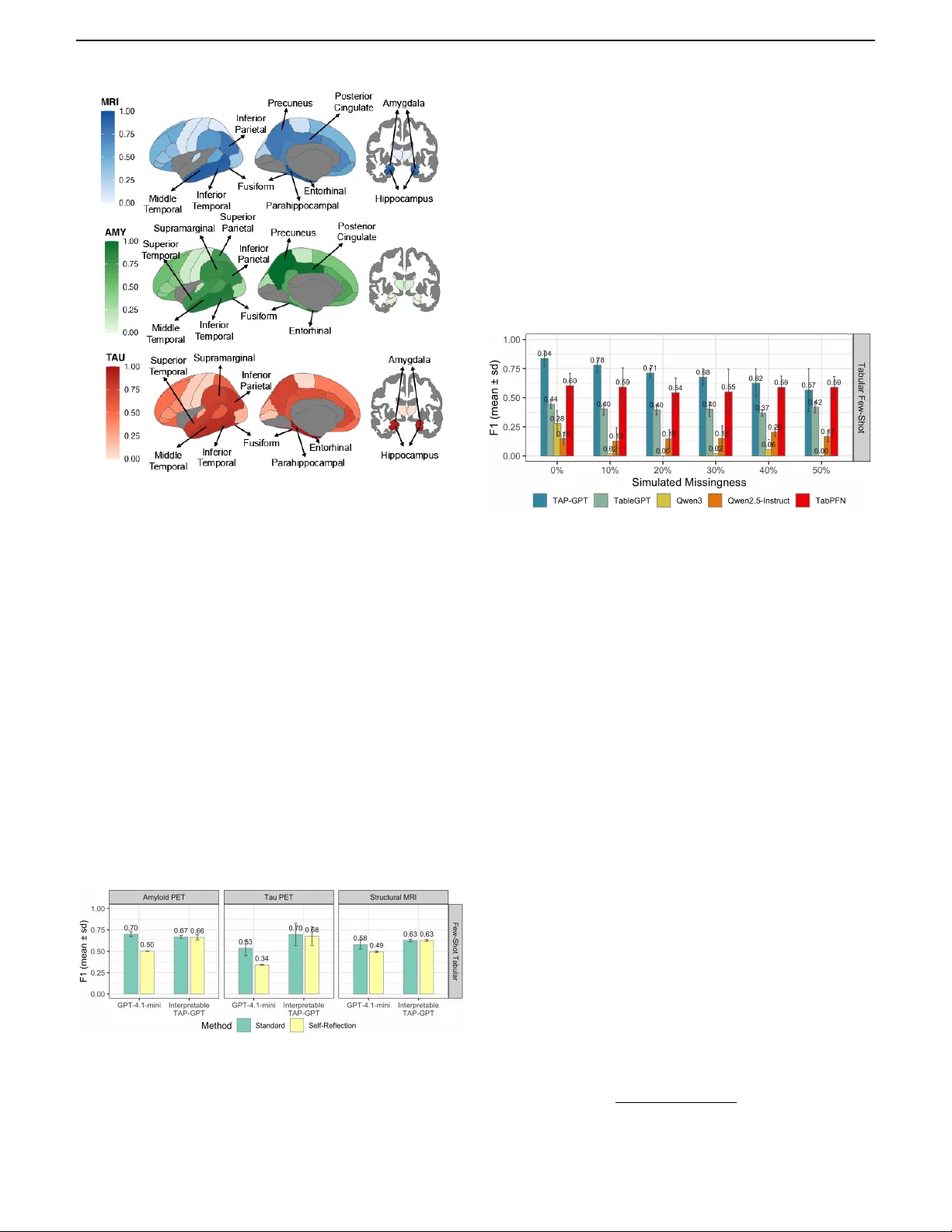
Accurate diagnosis of Alzheimer's disease (AD) requires handling tabular biomarker data, yet such data are often small and incomplete, where deep learning models frequently fail to outperform classical methods. Pretrained large language models (LLMs)…
Authors: Sophie Kearney, Shu Yang, Zixuan Wen
IEEE TRANSACTIONS AND JOURNALS TEMPLA TE 1 T ab ular LLMs f or Inter pretab le F e w-Shot Alzheimer’ s Disease Prediction with Multimodal Biomedical Data Sophie K ear ne y ∗ and Shu Y ang ∗ , Zixuan W en, Weimin L yu, Bojian Hou, Duy Duong-T ran, Tianlong Chen, Jason H. Moore , Mar ylyn D . Ritchie, Chao Chen, Li Shen † , f or the Alzheimer’ s Disease Neuroimaging Initiativ e ∗∗ Abstract — Accurate diagnosis of Alzheimer’ s disease (AD) requires handling tabular biomarker data, y et such data are often small and incomplete, where deep learning models frequently fail to outperf orm classical methods. Pretrained large language models (LLMs) offer few-shot generalization, structured reasoning, and interpretable out- puts, pr oviding a powerful paradigm shift for clinical predic- tion. W e propose T AP-GPT, Tabular Alzheimer’ s Prediction GPT , a domain-adapted tabular LLM framew ork built on T ab leGPT2 and fine-tuned for few-shot AD classification using tabular pr ompts rather than plain texts. We ev aluate T AP-GPT across four ADNI-derived datasets, including QT - P AD biomarker s and region-level structural MRI, amyloid PET , and tau PET for binary AD classification. Across mul- timodal and unimodal settings, T AP-GPT impro ves upon its backbone models and outperforms traditional machine learning baselines in the few-shot setting while remain- ing competitive with state-of-the-ar t general-purpose LLMs. W e show that feature selection mitigates degradation in high-dimensional inputs and that T AP-GPT maintains sta- ble performance under simulated and real-world missing- ness without imputation. Additionally , T AP-GPT produces structured, modality-aware reasoning aligned with estab- lished AD biology and shows greater stability under self- reflection, suppor ting its use in iterative multi-agent sys- tems. T o our knowledge, this is the first systematic applica- ∗ These authors contributed equally to this research. **Data used in the preparation of this ar ticle were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the inv estigators within the ADNI con- tributed to the design and implementation of ADNI and/or pro- vided data but did not participate in the analysis or wr iting of this repor t. A complete listing of ADNI inv estigators can be found at: http://adni.loni.usc.edu/wp- content/uploads/ how_to_apply/ADNI_Acknowledgement_List.pdf S. K ear ne y , S. Y ang, Z. W en, B. Hou, D . Duong-T ran, and L. Shen are with the University of P ennsylvania, Philadelphia, P A, USA (e-mail: sophie .kearney@pennmedicine.upenn.edu; yangsh@pennmedicine .upenn.edu; zxwen@sas.upenn.edu; houbo@pennmedicine.upenn.edu; duyanh.duong- tran@pennmedicine.upenn.edu; li.shen@pennmedicine .upenn.edu). C. Chen and W . L yu are with Stony Brook University , Stony Brook, NY , USA (e-mail: chao.chen.1@stonybrook.edu; welyu@cs.ston ybrook.edu) T . Chen is with the Univ ersity of North Carolina at Chapel Hill, Chapel Hill, NC, USA (e-mail: tianlong@cs .unc.edu). M. Ritchie is with the Medical University of South Carolina, Charleston, SC, USA (email: ritchiem@musc.edu) J. H. Moore is with Cedars-Sinai Medical Center , West Hollywood, CA, USA (e-mail: jason.moore@csmc.edu). † L. Shen is the corresponding author . tion of a tabular -specializ ed LLM to multimodal biomarker- based AD prediction, demonstrating that such pretrained models can effectively address structured clinical predic- tion tasks and la ying the foundation for tab ular LLM-driven multi-a g ent clinical decision-suppor t systems. The source code is public ly availab le on GitHub: https://github. com/sophie- kearney/TAP- GPT . Index T erms — Lar g e language models, health informat- ics, Alzheimer’ s Disease , c linical diagnosis I . I N T R O D U C T I O N The transition toward precision health in the management of complex human diseases such as Alzheimer’ s disease (AD) calls for the sophisticated inv estigation of heterogeneous, multimodal biomedical data. As a progressiv e and irrev ersible neurodegenerati ve disorder , AD represents the most common form of dementia globally and the sixth leading cause of mortality in the United States [1], [2]. Currently , curati ve treatments for AD remain limited, posing a critical challenge for both clinical neuroscience research and public health, espe- cially driven by the accelerated aging of the global population. Consequently , accurate and early detection becomes essential for proacti ve intervention. Previous studies have identified a set of AD biological markers and verified their ability to mea- sure AD pathology . The importance of these biomarkers was subsequently reflected in revised diagnostic criteria. In 2018, the National Institute on Aging (NIA) and the Alzheimer’ s Association W orkgroup’ s Research Framework proposed to use the A T[N] biomarker classification system [3], [4]. By defining AD in terms of a biological, rather than a clinical, construct, this system divides the major AD biomarkers into three categories based on the nature of their pathology: 1) ‘ A ’: the β -amyloid biomarkers (as measured by amyloid positron emission tomography (PET) imaging or cerebrospinal fluid (CSF) A β 42 ); 2) ‘T’: the pathological tau biomarkers (CSF phosphorylated T au or T au PET imaging); and 3) ‘N’: the neurodegeneration or neuronal injury biomarkers (atrophy on structural magnetic resonance imaging (MRI) scans, Fluo- rodeoxyglucose (FDG) PET scans, or CSF total T au) [4]. The multifaceted A T[N] biomarkers hav e been widely utilized in AD research and in vestigated in conjunction with other types of markers like cogniti ve measures for disease progression [5]. 2 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE From a data-science perspecti ve, these A T[N] measurements are often represented as tables: each subject corresponds to a row , each measurement (e.g., CSF tau biomarker or brain MRI-deriv ed regional feature) corresponds to a column. In the real-world setting, row/sample sizes of biomedical tables are frequently modest relative to the column/feature dimension- ality , and missingness is common during the data collection. Such tabular structure creates distinctiv e modeling challenges. Unlike language or images, tables lack a natural ordering or locality (e.g., columns are permutation inv ariant [6], [7]), and predictiv e signals may reside in cross-feature interactions that are difficult to capture with small size cohort. As a result, con- ventional machine learning methods, such as gradient-boosted and tree-ensemble approaches, have remained strong baselines for many small datasets. In particular, for AD outcome pre- diction, Random Forest (RF) and XGBoost have shown rob ust performance by modeling complex, non-linear relations among features from tabular data [8]–[10]; ev en logistic regression (LR) remains a clinical standard due to its simplicity and clarity [11]. In contrast, classic deep neural networks can be sensitiv e to limited sample sizes, feature scaling, and missing values [12], [13], which are commonly seen in AD tabular setting [14]–[16]. Despite the predictiv e performance, these con ventional machine learning and deep learning methods provide limited insight into which blood marker or brain region dri ves a given prediction, and they do not le verage prior knowledge in the biomedical domain beyond the training data. Recently , transformer-based large foundation models of fer an alternative paradigm and open up new opportunities for handling AD tabular data: rather than training a predictiv e model from scratch, one can adapt a pretrained model that has already learned broad prior knowledge ov er data dis- tributions and reasoning patterns [17], [18]. This paradigm includes both: (i) tab ular foundation models that learn in- context prediction behaviors for tables [6], [19]–[22], and (ii) large language model-based approaches that con vert tables into textual prompts through serialization or latent embeddings through table encoders [23]–[28]. Such methods can conduct few-shot prediction for very small datasets and incorporate broad prior knowledge, potentially improving performance on biomedical tables like AD A T[N] markers. T abular foun- dation models (TFMs) tailored to small-to-medium datasets hav e emerged [20]–[22]. As an example, T abPFN [6], [19] represents one of the most significant works in this class. It performs in-context learning (ICL) to predict labels from a small training table in a single forward pass without gradient updates. Although ICL was first observed in LLMs [29], later studies showed transformers can lev erage ICL to approximate classical learning algorithms such as logistic regression and neural networks [30], [31]. Pre-trained via prior-fitting on 100 million synthetic datasets spanning di verse causal structures, T abPFN achieved state-of-the-art accuracy on small, noisy tables with outliers and missing values, requiring only 50% of the data to match con ventional models [6], [19]. Notable e xten- sions hav e explored adapting T abPFN to larger or more com- plex tasks through finetuning and retrieval-augmentation [32], [33]. Howe ver , TFMs are primarily predictive engines and do not nativ ely provide natural-language reasoning that is valuable for clinical interpretability , error analysis, and down- stream agentic workflo ws. In parallel, pretrained on large text corpora, LLMs ex- hibit emergent abilities for tab ular tasks, such as tab ular data generation, table understanding, prediction, reasoning and even feature selection/engineering, etc . [23]–[28]. With prompting and finetuning, they can be adapted to tabular tasks, addressing limitations of naiv e transformer architectures for handling tabular formats. In particular , as the leading open-source ef forts of the field, T ableGPT unifies tables, language, and functional commands for table understanding and manipulation with natural language instructions [34]. It utilizes dedicated table encoders and compresses tabular information into compact latent representations to mitigate token-length constraints and capture schema-lev el context. T ableGPT2 [35] scales this idea via large-scale pretraining on 593K tables and 2.36M table-question pairs, with a schema- aware table encoder and a 7B–72B parameter QW en2.5 LLM decoder [36]. It achie ved ∼ 35–49% improvements over base LLMs on div erse table-related business intelligence tasks. The most recent variant of the family , T ableGPT -R1, released in December 2025 [37], goes beyond standard supervised fine- tuning by using a systematic reinforcement learning approach and a powerful QW en3 [38] backbone to strengthen multi- step reasoning and advanced table understanding tasks [37]. Despite these strong gains from tabular LLMs, studies show that they still lag behind humans on complex, real-world tables that require deep domain knowledge [35], [39]. Building on these advances, we introduce a domain-adapted tabular LLM frame work for A T[N]-based AD diagnosis, termed T AP-GPT : T abular Alzheimer’ s Prediction GPT , which nov elly adapts the multimodal T ableGPT2 model for few-shot classification of AD vs cognitiv ely normal (CN) individuals from A T[N] feature tables. W e formulate the task as tabu- lar fe w-shot in-context learning paradigm, where the model is prompted with a small table containing sev eral example subjects with neuroimaging, etc. , features and diagnosis labels, alongside a tar get subject whose label to be predicted. T o adapt T ableGPT2 for AD tabular prediction, we employ div erse strategies to construct dedicated training tables from clinical data, and we perform parameter-efficient finetuning [40] on these tables. T o our knowledge, this represents the first sys- tematic application of tabular LLM to AD biomarker study , uniquely combining tabular-LLM’ s table understanding and reasoning capabilities with specific medical knowledge regard- ing AD and A T[N] markers. W e ev aluated T AP-GPT against a set of general-purpose LLMs and TFM, as well classic machine learning methods, ef fectively demonstrating the po- tential of T AP-GPT for structured biomedical prediction and paving the way for future multi-agent diagnostic systems on multimodal AD data. A preliminary version of T AP-GPT has been reported in the ACM-BCB conference in 2025 [41]. W e applied T AP-GPT on the QT -P AD biomarker dataset derived from ADNI (cohort 1, GO, 2), with 15 clinical markers and 4 cov ariates as features. In this study , we present a substantial expansion of the conference proceeding by comprehensively exploring and enhancing T AP-GPT , and we make the follow- ing key contributions here: KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 3 • Generalization: V alidated model generalization by adapting T AP-GPT to three additional neuroimaging datasets: amyloid PET , tau PET , and structural MRI, with regional measures stored in tab ular format. Extended beyond the single QT -P AD dataset for fe w-shot AD prediction. • Enhanced Interpr etability: Enhanced interpretability analyses with multimodal reasoning and self-reflection prompting, laying the foundation for future multi-agent AD diagnosis workflo ws. • Featur e Selection: Dev eloped systematic feature selec- tion for high-dimensional tables via classic LASSO and LLM-guided ranking approaches, under context-length constraints. • Robustness to Missingness: Conducted robustness anal- ysis under both simulated and naturally occurring missing data deriv ed from ADNI. I I . M E T H O D S W e start this section by introducing our few-shot and in- context learning notions in the context of tabular prediction tasks follo wing previous works [6], [19], [32]–[35], as they are slightly different from the normal non-tabular scenario with LLMs. Formally , let M denote the set of modalities considered in this study , including QT -P AD clinical biomarkers, struc- tural MRI, amyloid PET , and tau PET . For each modality m ∈ M , we define a modality-specific dataset D ( m ) = { ( x i, ( m ) , y i ) } N m i =1 consisting of N m labeled subject samples. Each sample i in modality m is represented by a feature vector x i, ( m ) = ( x i, ( m ) 1 , . . . , x i, ( m ) d m ) ∈ R d m containing d m structured biomarkers or regional imaging measurements, and a corresponding clinical diagnosis label y i ∈ { 0 , 1 } , where 0 denotes cogniti vely normal (CN) and 1 denotes Alzheimer’ s disease (AD), consistent across QT -P AD and imaging datasets. For each modality m , the table columns (features) are de- scribed by a set of natural language feature names F ( m ) = { f ( m ) 1 , . . . , f ( m ) d m } , which are incorporated directly into the prompt construction. The goal of the tabular prediction task for modality m is to learn a model that, giv en a new subject’ s feature vector x ( m ) test ∈ R d m and (optionally) contextual examples in the form of a structured table, can accurately predict the corresponding diagnosis y test ∈ { 0 , 1 } . While this work focuses on binary classification, the formulation naturally generalizes to tabular regression or multiclass classification tasks. An ov erview of this framework is provided in Figure 1. In the few-shot setting, prediction for a target patient x ( m ) t is conditioned on a set of k labeled in-context examples drawn from a split-specific, disjoint in-context learning (ICL) pool. For each data split s ∈ { train , v al , test } , we define a corresponding ICL pool I ( m ) s that is used exclusi vely to construct prompts for samples in that split. Formally , for modality m ∈ M , the context set for a target sample in split s is defined as C ( m ) t = { ( x ( m ) t j , y t j ) } k j =1 , where each ( x ( m ) t j , y t j ) is sampled from the same split’ s ICL pool I ( m ) s and does not ov erlap with the target sample. The ICL pools are used Fig. 1. Overview of the T AP-GPT frame work. F or each dataset , we split the subjects into tr aining, testing, and pools f or in-conte xt e xamples, with an optional feature selection step . W e construct tab les f or finetuning and ev aluating T AP-GPT for the task of Alzheimer’ s disease prediction. solely for prompt construction and are not directly used as supervision outside their corresponding split. A structured prompt is constructed by combining the in- context examples with the unlabeled target patient: P ( m ) t = Φ C ( m ) t , x ( m ) t , where Φ( · ) denotes a deterministic formatting function that renders the examples and target patient into either a tabular representation (preserving explicit column structure) or a serialized natural language representation. T AP-GPT is an autoregressiv e language model parameter- ized by θ that defines a conditional distribution over output tokens given the prompt: p θ ( y t | P ( m ) t ) . The model is fine- tuned to maximize the likelihood of the correct diagnosis label under this conditional distribution. During inference, we restrict the output space to the binary label set { 0 , 1 } using constrained decoding. The predicted diagnosis is therefore obtained as ˆ y t = arg max y ∈{ 0 , 1 } p θ ( y | P ( m ) t ) . T o predict ˆ y t , some prior approaches rely on sim- ple serialization strategies that conv ert structured tabular data into plain-text sequences before prompting pretrained LLMs [27], [28]. In such methods, a serialization function Serialize ( x ( m ) , y , F ( m ) ) uses the feature names F ( m ) to transform each row into a natural language description (e.g., “The f 1 biomarker value is x 1 . ”), then a generic LLM directly processes the resulting text to generate a prediction: ˆ y t = LLM ( Serialize ( C ( m ) t , x ( m ) t )) . Ho wever , this approach loses explicit column alignment and structured relationships present in the original table, and can become token-inefficient as the number of features or in-context examples increases [27]. In contrast, our proposed framework, T AP-GPT , makes use of T ableGPT2’ s encoder–decoder architecture with a seman- tic table encoder , which maintains structural aw areness, and an LLM decoder . Let g θ 0 denote the pretrained T ableGPT2 model, originally trained on large-scale generic tabular data for table understanding tasks. T o adapt this model to supervised Alzheimer’ s disease 4 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE prediction, we perform task-specific fine-tuning using QLoRA. Let ∆ θ denote low-rank parameter updates applied to the de- coder weights. The adapted parameters are therefore θ = θ 0 + ∆ θ . In our framew ork, the pretrained semantic table encoder is kept fixed, preserving its learned structural representations of tabular data. Low-rank adapters are injected into the decoder , allowing the model to specialize to the binary diagnostic objectiv e while updating only a small subset of parameters. The resulting modality-specific predictor f ( m ) θ is trained to approximate the conditional distribution p ( y | x ( m ) , C ( m ) ) , by maximizing the likelihood of the correct diagnosis label giv en the structured prompt, without updating its weights during inference [31]. In the following subsections, we provide more details for the data, framew ork and experimental setup. A. Datasets Data used in the preparation of this article is obtained from the Alzheimer’ s Disease Neuroimaging Initiativ e (ADNI) database ( http://adni.loni.usc.edu ) [42], [43]. The ADNI was launched in 2003 as a public-priv ate partnership led by Principal In vestigator Michael W . W einer , MD. The primary goal of ADNI has been to test whether serial MRI, PET , other biological markers, and clinical and neuropsycho- logical assessment can be combined to measure the progres- sion of mild cogniti ve impairment (MCI) and early AD. All participants provided written informed consent, and study pro- tocols were approved by each participating site’ s Institutional Revie w Board (IRB). Up-to-date information about the ADNI is av ailable at www.adni- info.org . 1) Quantitativ e templates for the progression of AD Biomarker Data : The first dataset we focus on is a subset of the ADNI 1/Go/2 cohorts, i.e., the Quantitativ e templates for the progression of AD (QT -P AD) project dataset 1 , which was initially developed as an AD Modeling Challenge from ADNI and has been widely used for diverse AD-related research. The QT -P AD dataset contains multimodal biomarkers indicating progression of AD across the A T[N] framew ork and beyond. W e excluded the five congnitiv e score metrics because these measurements are heavily used by clinicians for diagnosis, and are therefore highly collinear to diagnosis. The resulting dataframe contained 15 clinical markers for each subject: PET measures (FDG PET and Amyloid PET), cerebrospinal fluid measures (CSF ABET A, CSF T A U, CSF PT au), structural MRI-deriv ed measures from FreeSurfer (Whole brain, hip- pocampus, entorhinal cortex, ventricles, mid-temporal lobe, fusiform gyrus), as well as one genetic marker (APOE4 status) and three demographic covariates including age, gender and years of education. W e remov ed any samples with missing values and extracted the baseline measurement of each patient which resulted in 237 CN and 96 AD patients, 333 in total. 2) Regional Summar y Imaging Data : The second dataset we focus on is a subset of participants from the ADNI-1, ADNI- GO, and ADNI-2 cohorts with av ailable amyloid PET , tau PET , and structural MRI regional summary data. Amyloid- β and T au PET images were processed by the UC Berkeley PET Analysis Group using standardized preprocessing pipelines. 1 https://www.pi4cs.org/qt- pad- challenge The extracted regional PET standardized uptake value ratios (SUVRs) using FreeSurfer-defined cortical and subcortical regions of interest (R OI). Structural MRI volumetric measures were also extracted, yielding region-le vel volumetric features aligned with PET data. As features, we used region-lev el summaries derived from FreeSurfer , including 68 cortical regions (34 per hemisphere) and four subcortical regions (thalamus, hippocampus, amyg- dala, and caudate), resulting in 74 regional summaries across all three modalities. In addition, we included three demo- graphic cov ariates (age, gender and years of education) and one genetic marker , copies of APOE4. Fig. 2. The four prompt formats used in our experiments, with tab- ular prompts shown in green and serialized prompts in blue. In each case, the model is asked to predict Alzheimer’s disease status for the same held-out patient. All v alues sho wn are synthetic with abbre viated prompts. For each modality , we selected the most recent av ailable visit to maximize the number of AD cases and improve class balance, restricting the cohort to participants with either CN or AD diagnoses. In the amyloid PET modality , this resulted in 1,031 patients (691 CN, 340 AD) and the tau PET modality , this resulted in 610 patients (484 CN, 126 AD). Structural MRI data were available for participants in both PET cohorts, so to maximize sample size, we extracted MRI features from the amyloid PET cohort, resulting in the same set of 1,031 participants. 3) Creating T abular Prompts : T o support in-conte xt learning, we designed a splitting strategy that strictly separates model training, validation, and testing data from samples used for prompt construction. For each dataset (QT -P AD, amyloid PET , tau PET , and structural MRI), we allocated approximately 40% of samples to training, 10% to validation, and 20% to testing. The remaining data were divided into three non-overlapping KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 5 ICL pools corresponding to the train, v alidation, and test splits (approximately 10% each). ICL pools are used exclusi vely to populate prompts and are ne ver included in model optimization or ev aluation. W e designed four prompt formats for the binary classifica- tion task to ev aluate performance under different contextual and representational settings (Figure 2). In each case, we construct a patient-lev el prompt and the model is asked to predict whether the patient is cognitiv ely normal or has AD. The formats vary along two dimensions: contextual exposure (zero-shot versus few-shot) and input representation (tabular versus serialized natural language). In all settings, we append the patient information to a standardized natural language instruction requesting a binary classification for the tar get sample. • Zero-Shot T abular A single unlabeled row in a tabular format (one target sample). • Zero-Shot Serialized A single unlabeled natural lan- guage description of one patient’ s information. • Few-Shot T abular For each unlabeled target patient, k labeled examples are randomly sampled from the corre- sponding ICL pool. These k rows are presented abov e the target patient in the table, with the target appearing in the final row and their diagnosis label masked. • Few-Shot Serialized Mirroring the few-shot tabular set- ting, but each patient’ s features are con verted into natural language. The k in-context examples include diagnosis labels, while the target sample is presented without a label. For serialized prompts, we conv ert each patient’ s structured features into a standardized natural language description while preserving consistent formatting across datasets. Pronouns are adjusted according to recorded sex, and measurement units are retained when applicable. For imaging modalities, we present patient-lev el cov ariates first, followed by regional imaging features formatted as structured ke y–value pairs (R OI=value) to preserve clarity across numerous measurements. In contrast, QT -P AD biomarkers, which span multiple biological domains, are assembled into a clinically interpretable narrativ e summary . W e display tabular prompts with the head() function from the pandas library . Column order is consistent between all data splits. W e embedded all formats of presenting patient data in a natural language prompt instructing the model to provide a diagnosis with instructions tailored to the prompt format. For interpretability experiments, we modified only the in- struction and expected output format while keeping patient data and in-context examples identical to the corresponding non-interpretable prompts. W e appended “Let’ s think step by step. ” and instructed the model to return a JSON object containing (i) the predicted label ( 0 =CN, 1 =AD), (ii) a free- text reasoning explanation, and (iii) a scalar confidence score. The supervised objecti ve remained binary classification, en- abling interpretable outputs without altering the training loss or architecture. B. T AP-GPT F ramew or k W e present T AP-GPT , a domain-adapted tabular large lan- guage model build on T ableGPT2-7B 2 , that integrates a static tabular encoder with an LLM decoder backbone (Qwen2.5- 7B). While T ableGPT2 was originally optimized for business- oriented tabular understanding tasks, we repurpose its de- coder for structured clinical prediction of Alzheimer’ s disease. Specifically , we finetune the decoder using labeled prompts that contain small tables of biomedical features, enabling the model to map structured patient data to diagnostic outcomes (Figure 1). For each dataset and random seed, we generate tabular in- puts for e very patient in the training, testing, and validation test splits using their corresponding ICL pools. For each patient, we con vert this structured table into four prompt formats (zero- shot tabular , zero-shot serialized, few-shot tabular , and few- shot serialized) and finetune a separate T AP-GPT model for each format to isolate the impact of structure and context. W e train T AP-GPT using supervised learning to predict a binary diagnosis label (AD=1, CN=0). W e place the diagnosis token in a fixed position within each prompt to ensure con- sistent parsing and ev aluation across seeds and formats. W e repeat all experiments across ten random seeds (36, 73, 105, 254, 314, 492, 564, 688, 777, and 825). T o adapt the large decoder, we apply parameter-efficient finetuning using QLoRA implemented with the HuggingFace transformers, peft, and bitsandbytes libraries. QLoRA updates low-rank adaptation matrices while keeping the base model weights quantized in 4-bit precision, which reduces memory requirements without full weight retraining. Due to archi- tectural constraints of the released T ableGPT2 model, we keep the tabular encoder fixed and update only the decoder parameters during adaptation. This design allo ws us to pre- serve general-purpose tabular representations while injecting domain-specific diagnostic knowledge into the LLM. During inference, we constrain the model’ s output space to { 0,1 } using a custom logits processor to enforce valid binary predictions. For interpretability experiments, we modify the prompts to additionally request a structured reasoning explana- tion and a scalar confidence score, while maintaining the same supervised objectiv e of binary classification. This prompt-lev el modification enables interpretable outputs without altering the underlying training loss or architecture. C . Exper imental Setup W e designed a set of experiments to ev aluate T AP-GPT across four multimodal biomedical datasets. For LLM-based models, we ev aluated four prompt formats, as shown in Figure 2. Only the model type and prompt configuration varied across experiments. Performance was primarily assessed using F1 score, with balanced accuracy , precision, and recall also reported. For the QT -P AD tabular dataset, we performed a k -ablation study on the validation set to determine the optimal number of in-context examples, where k denotes the number of in-context 2 https://huggingface.co/tablegpt/TableGPT2- 7B 6 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE Fig. 3. QT -P AD mean F1 across models in zero-shot and fe w-shot ( k = 8 ) conte xts. LLMs use tabular (green) and serialized (blue) prompts with error bars for standard de viation; T abPFN and traditional ML (yello w) operate directly on structured data. examples. W e finetuned T AP-GPT models separately for each k ∈ { 2 , 4 , 6 , 8 , 10 , 12 , 16 , 20 } across three seeds under the tabular fe w-shot prompt format. W e used v alidation F1 to select the k that achieved the most stable and consistently high performance across seeds and used this value for all subsequent few-shot tabular experiments. Giv en the substantially larger feature spaces in the imaging- deriv ed datasets (MRI, amyloid PET , tau PET), we incorpo- rated feature selection prior to prompt construction to mitigate performance degradation due to prompt length. Using the training set only , we applied LASSO to select the top- p features per modality and seed. W e conducted a joint p × k ablation for p ∈ { 8 , 16 , 32 } and k ∈ { 4 , 8 , 12 } to character- ize trade-offs between feature dimensionality and in-context supervision. Optimal configurations were selected based on validation F1 and applied to final test ev aluation. T o benchmark T AP-GPT , we compared performance across all prompt formats and datasets with a diverse set of baselines. W e included unfinetuned T ableGPT2 to assess the impact of task-specific finetuning and incorporated T ableGPT -R1 for imaging analyses. T o isolate the contribution of tabular spe- cialization, we ev aluated Qwen2.5-7B-Instruct (the backbone of T ableGPT2) and Qwen3-8B as general-purpose LLMs. W e also included GPT -4.1-mini as a strong general LLM benchmark for zero- and few-shot inference. T o examine the effect of finetuning without tabular pretraining, we finetuned Qwen2.5-7B-Instruct under identical training conditions. W e included T abPFN as a non-LLM tabular foundation model, tested only on few-shot tasks. Finally , we benchmarked four traditional machine learning models on the few-shot tasks: Logistic Regression, Random Forest, XGBoost, and Support V ector Machine—on the few-shot tabular setting to provide classical baselines. For ev ery dataset, we performed hyperparameter tuning using Optuna to maximize validation F1. For finetuned T ableGPT2 and Qwen2.5 models, we tuned LoRA rank, dropout rate, learning rate, batch size, weight decay , maxi- mum training steps, and learning rate scheduler . For imaging experiments with feature selection, we treated the number of features p as a tunable parameter within the p × k ablation grid. W e tuned traditional machine learning models ov er their respectiv e standard hyperparameters (e.g., regularization strength, tree depth, kernel type, and subsampling strategies). For each model and modality , we selected the configuration achieving consistently strong validation F1 across seeds for final test ev aluation. W e conducted all experiments on a high-performance com- puting cluster using NVIDIA A100 GPUs (80 GB HBM memory) with 160 GB system RAM per node. W e allocated each training job a single GPU, 4 CPU cores, and 1 node under a SLURM scheduler . Due to the large number of experimental configurations (modalities, seeds, prompt formats, and p × k combinations), we ex ecuted several hundred GPU jobs. W e stored model checkpoints and adapters via Hugging Face Hub, with approximately 0.25 TB of model storage accumulated across 335 experiment v ariants. This large-scale compute infrastructure enabled systematic ev aluation across modalities and ablation settings while maintaining strict separation of training, validation, and test data. More details of this study , such as LLMs’ hyperparameters, various prompts, together with the source code, are all publicly available in our GitHub repository: https://github.com/sophie- kearney/ TAP- GPT . I I I . R E S U L T S A. Overall Model P erformance W e ev aluated T AP-GPT performance on the test set across four prompt formats (zero-shot tabular , zero-shot serialized, few-shot tabular , and few-shot serialized). T o contextualize performance, we ev aluated T AP-GPT against four model categories: (1) classical machine learning baselines (logistic regression, random forest, SVM, XGBoost) (2) T abPFN to represent state-of-the-art tab ular foundation model approaches; (3) vanilla LLMs without task-specific fine-tuning, including Qwen2.5-Instruct, Qwen3-Instruct, and GPT -4.1 mini as a strong external LLM from a different model family; and (4) tabular -specialized deriv ativ e T ableGPT2. Because T AP-GPT KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 7 Fig. 4. Imaging ROI mean F1 across models in the zero- and fe w-shot ( k = 4 ) conte xts. T abular and Serialized prompts are given to LLMs, with traditional ML and T abPFN models directly using the data. Model performance is e valuated across three imaging modalities, Amyloid PET , T au PET , and Structural MRI. is fine-tuned from T ableGPT (itself adapted from Qwen2.5- Instruct), this hierarchy allows us to assess the incremental contribution of tabular specialization and disease-specific fine- tuning. Interpretable T AP-GPT is the modified prompting for interpretable output from the same T AP-GPT model. 1) Model P erformance on Biomarker Data : In the zero-shot setting (Figure 3, left), serialized prompts consistently outper- formed tabular prompts. Because the tabular zero-shot format contains only a single patient row without in-context e xamples, the serialized representation provides a more natural linguistic structure for encoding clinical information. T AP-GPT achie ved strong results in both settings, with stronger results under serialization. Notably , introducing prompting for interpretable outputs with CoT reasoning produced a substantial improv e- ment over the standard T AP-GPT configuration in both tabular and serialized formats. The interpretable T AP-GPT v ariant achiev ed the highest overall zero-shot performance. T AP-GPT and Interpretable T AP-GPT have the strongest performance in the few-shot tabular context (Figure 3, right), likely due to pairing the tabular understanding of T ableGPT with task-specific knowledge acquired from finetuning. In- corporating CoT interpretable responses again boosted T AP- GPT’ s performance in this task to mean F1=0.89. As expected, T abPFN2.5 performs strongly in this context despite the very low-shot setting. Notably , T AP-GPT outperforms all traditional machine learning baselines, T abPFN, and vanilla LLMs in the few-shot tabular task. GPT -4.1-mini performs well across both formats in this context, which suggests that updating the backbone of T AP-GPT with more capable base models could further improv e prediction performance. In contrast, performance in the serialized few-shot setting is substantially weaker across most LLMs, with several models collapsing tow ard majority-class predictions. This degradation may reflect the increased prompt length and reduced structural clarity of serialized inputs when combined with multiple in-context examples. 2) Model P erformance on Imaging Data : W e next ev aluated T AP-GPT on the unimodal imaging-deriv ed datasets amyloid PET , tau PET , and structural MRI volume under a 4-shot setting with 16 selected features. W e tested the same models on these datasets as with the QT -P AD data, with the addition of T ableGPT -R1, the reasoning version of T ableGPT , built on Qwen3 backbone. The classic machine learning models can only be tested in the few-shot setting and are not subject to prompt formatting because they operate directly with the data. In this very low- shot setting of four training samples and one test sample, these methods naturally struggle to perform competitiv ely to the LLMs, which benefit from semantic knowledge and pre-training. Howe ver , T AP-GPT consistently outperforms all traditional methods and T abPFN. In the few-shot tabular setting, across each of the datasets, we observe a consistent performance progression from the base backbone to the domain-adapted model. For example, in the tau modality , Qwen2.5 started with a mean F1 of 0.43, then T ableGPT2 had a mean F1 of 0.48, and then T AP-GPT showed an F1 of 0.68. A similar stepwise trend is reflected in the amyloid modality , and although not as strong, in the MRI modality . Unlike the QT -P AD dataset, the serialized prompts (blue in Figure 4) perform similarly to the tabular prompts (green). In this setting, we implemented serialization using a structured key-v alue format appended to the minimal patient description, rather than fully narrati ve text. Because key–v alue serialization preserves much of the original structural org anization and does not substantially inflate prompt length, it remains closer to a semi-structured representation. As a result, performance between tabular and serialized formats remain similar in this unimodal imaging setting, in contrast to the larger format- driv en differences observed in QT -P AD. In contrast to the QT -P AD results, the strongest comparator in the imaging-derived setting is GPT -4.1 mini, a more ad- vanced general-purpose LLM. In the tabular few-shot setting, T AP-GPT performs similarly or better across modalities. For amyloid PET , Interpretable T AP-GPT achieves an F1 of 0.64, comparable to GPT -4.1 mini (F1 = 0.65). In the tau modality , 8 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE T AP-GPT reaches an F1 of 0.68 compared to 0.53 for GPT -4.1 mini. In the MRI modality , GPT -4.1 mini achiev es the highest tabular few-shot performance (F1 = 0.66). While T AP-GPT’ s tabular performance was lower in this modality , the serialized Interpretable T AP-GPT achiev es competitive performance (F1 = 0.65), closely matching GPT -4.1 mini. For general-purpose LLMs, the shift from zero-shot to few- shot prompting did not produce the larger gains observed in QT -P AD. Ho wever , few-shot prompting consistently improved T AP-GPT performance relative to zero-shot. A similar trend is observed for GPT -4.1 mini. These results indicate that, ev en in unimodal settings where strong prior knowledge may already exist in pretrained models, access to labeled in-context examples provides additional task-specific signal. B. Ablations Fig. 5. Number of ICL examples ( k ) ablation analysis on QT -P AD data across T ableGPT2, T abPFN, and T AP-GPT . T abPFN performance steadily improv ed with larger k, T ableGPT2 improv ed up to k = 6 and declined thereafter , and T AP-GPT peaked at k = 8 . 1) k-Ablation Analysis for Biomarker Data : T o choose an optimal number of ICL examples ( k ) in the few-shot setting, we tested a set of different k s { 2 , 4 , 6 , 8 , 10 , 12 , 16 , 20 } on T AP-GPT , T ableGPT2, and T abPFN (Figure 5). T esting a k value on T AP-GPT requires re-finetuning T ableGPT for that format of prompt across all seeds, so due to GPU memory and sample size restrictions, we restricted the upper limit of k to 20 and tested across three seeds. T ableGPT performance seems to peak around k = 6 to k = 8 but remains relativ ely steady beyond this point. As expected because it is built for much larger tables than this few-shot setting, T abPFN shows increasing performance with a higher k . T AP-GPT has the highest mean F1 at k = 8 (0.831, SD = 0.0702), with k = 6 and k = 10 achieving similar performance. W e selected k = 8 for all few-shot experiments with QT -P AD data. Fig. 6. Number of ICL examples ( k ) and number of f eatures ( p ) ablation analysis on imaging data for T AP-GPT . Mean F1 score (± standard deviation across three random seeds) is shown for each ( k , p ) combination. 2) k- and p- Ablation Analysis for Imaging Data : T o determine the appropriate values for the number of ICL examples ( k ) and selected features ( p ) for fe w-shot prompting, we conducted an ablation study for the tabular fe w-shot prompt on the T AP- GPT model for each modality . The correct choice of ( k , p ) balances prompt length and necessary informational content to optimize performance. For each modality , p denotes the num- ber of imaging-deriv ed regional features selected from the full set of 68 cortical and 4 subcortical FreeSurfer regions, while patient-lev el cov ariates (APOE4 status, age, sex, and years of education) were always included and not counted to ward p . W e finetuned T ableGPT2 on tabular few-shot prompts with each combination of p ∈ (8 , 16 , 32) and k ∈ (4 , 8 , 12) across three arbitrarily chosen seeds (36, 73, 314) for each modality dataset, as seen in Figure 6. Performance across amyloid PET and tau PET consistently peaked at lower v alues of k and p , further highlighting the limitation of tabular LLMS with large tables as text input. k = 4 shows the most stability across v alues of p for amyloid and tau modalities and pairs best with moderate feature counts of p = 16 . In contrast, the MRI volumetric modality showed weaker signal o verall with some inconsistent and v ariable gains at larger p or k v alues. T o maintain consistency across modalities, we set p = 16 and k = 4 for all imaging modality prompts. In addition, p = 16 provides a feature dimensionality similar to the QT -P AD biomarker dataset, enabling more consistent comparison of T AP-GPT performance across all datasets ev aluated in this study . Fig. 7. T AP-GPT P erformance using all 72 ROIs compared to selecting the top 16 features f or each modality . 3) Impact of Feature-Dimensionality on T abular LLMs : Fig- ure 7 examines the impact of feature dimensionality on few- shot tabular performance across amyloid PET , tau PET , and MRI, comparing T AP-GPT to two related baselines, Qwen3- 8B and T ableGPT -R1. For each modality , we ev aluate perfor- mance using either all 72 av ailable regional features ( p = 72 ; light green) or a reduced subset of the top 16 features selected via LASSO ( p = 16 ; dark green). This experiment isolates the effect of feature selection in a long-context setting and assesses whether ne wer foundation models better tolerate high- dimensional tabular inputs. Across all three modalities, T AP-GPT shows consistent improv ement in performance when feature selection is applied. In contrast, using all 72 features leads to performance degrada- tion, particularly for tau PET and MRI, supporting the use of feature selection for high-dimensional tables for tab ular LLMs. While Qwen3 largely maintains comparable performance be- tween p = 72 and p = 16 , its overall performance remains KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 9 lower than T AP-GPT . T ableGPT -R1 exhibits sensitivity to feature dimensionality , with near-zero performance in the p = 72 setting across modalities, indicating dif ficulty handling long tabular contexts without feature reduction. These results suggest improv ements in base model archi- tecture may not be sufficient to address long-context chal- lenges with high-dimensional clinical tables. Feature selection enables effecti ve performance for tabular LLMs. Fig. 8. Example inter pretab le outputs from T AP-GPT across all datasets. For each modality , the model correctly predicts the patient’s diagnosis and returns a structured JSON containing the prediction, confidence score, and reasoning. C . Inter pretability 1) Interpretable Prompting Rev eals Multimodal Reasoning : In addition to improving predictiv e performance, interpretable prompting provides direct insight into ho w T AP-GPT syn- thesizes multimodal clinical features (Figure 8). Across all four datasets, T AP-GPT produces structured JSON outputs containing the binary prediction, a probability estimate, and a stepwise reasoning explanation. By qualitatively inspecting the outputs, we noticed modality-specific feature prioritization. In the QT -P AD dataset, T AP-GPT often references features spanning multiple biological domains when explaining its predictions, including genetic risk factors, patterns in amyloid and tau biomarkers, and measures of neurodegeneration. This cross-domain reasoning suggests that the model integrates multimodal inputs to form a more comprehensiv e patient profile. For the imaging modalities, even when restricted to the selected subset of 16 features, T AP-GPT emphasizes regions consistent with established Alzheimer’ s disease biology . In the structural MRI example shown in Figure 8, the model correctly interprets higher volumetric values as preserved structure rather than increased atrophy and cites normal hip- pocampal and amygdala v olumes to support the classifica- tion of cognitively normal. In the tau PET and amyloid PET examples, the model references APOE4 status alongside modality-specific regional uptake patterns, combining genetic risk with imaging evidence in its reasoning. These examples illustrate how finetuning enables T AP-GPT to lev erage both domain-specific training signals and the underlying semantic knowledge encoded in the base LLM to generate clinically interpretable explanations. The interpretability framew ork also provides flexibility in output structure. Because interpretability is induced via prompt design rather than architectural modification, we can request alternativ e structured outputs (e.g., predicted AD risk prob- ability , ranked important features, or feature-level justifica- tions) without retraining the model. This adaptability enables downstream integration into clinical decision-support pipelines where explanation format may vary by use case. At the same time, our qualitati ve analysis rev eals reasoning inconsistencies. In some cases, the model misinterprets feature directionality (e.g. treating elev ated FDG values as indicativ e of AD when reduced metabolism is typically associated with disease) or incorrectly references values from other ro ws as belonging to the target patient. These errors underscore the need to validate generated explanations and caution against interpreting reasoning strings as causal evidence. Exposing the model’ s intermediate reasoning provides feedback that can guide targeted refinement of prompting strategy or additional finetuning. The ability to surface and inspect model reasoning provides additional diagnostic signal unav ailable in con ven- tional tabular classifiers. 2) LLM-Derived Feature Ranking Across Imaging Modalities : W e prompted GPT -4.1-mini to rank each of the R OIs found in the imaging datasets according to their importance in diag- nosing AD for each modality and mapped these normalized rankings onto anatomical space in Figure 9. W e used GPT -4.1- mini because T AP-GPT is finetuned on a subset of features, and GPT -4.1-mini performs strongly on this task. In the MRI modality , GPT -4.1-mini prioritized sev eral me- dial temporal and subcortical structures, including the hip- pocampus and amygdala, alongside posterior cortical areas such as the precuneus and posterior cingulate. Many of these regions are established in neurodegeneration and were also selected by LASSO, demonstrating overlap between LLM- deriv ed and data-driven feature selection. In the amyloid modality , the model emphasized posterior cortical regions but placed less weight on frontal areas than both prior literature and the LASSO-based selection, suggesting a different pattern of importance across cortical lobes. For tau PET , GPT -4.1- mini concentrated importance in medial temporal and inferior temporal regions, including the entorhinal cortex and parahip- pocampal areas, aligning with the known spatial progression of tau pathology in AD. Despite these anatomically plausible patterns, GPT -4.1-mini consistently selected a partially distinct subset of regions com- pared to LASSO across modalities. These differences indicate that the LLM may rely on broader clinical and biological knowledge rather than purely on statistical sparsity in the train- ing data, supporting the use of LLM-deri ved rankings as an alternativ e or complementary feature selection strategy . GPT - 4.1-mini could act as a helper agent to create an optimized 10 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE Fig. 9. Regional feature importance as rank ed by GPT -4.1-mini across modalities. Cor tical and subcor tical regions are colored according to their nor malized impor tance scores (0–1), derived from the LLM- generated ranking of input features. Labeled regions reflect the top 10 ROIs chosen f or each modality . set of features prior to prompt creation, operating in a similar knowledge space as tabular LLM predictor agents. 3) Self-Reflection : T o ev aluate stability under iterati ve rea- soning, we introduced a self-reflection step where both T AP- GPT and GPT -4.1-mini were prompted to re view and op- tionally revise their own predictions across three arbitrarily selected seeds (688, 825, and 492). Figure 10 shows that GPT -4.1-mini shows considerable performance loss under self- reflection across modalities. In contrast, T AP-GPT maintains similar performance between standard and self-reflection con- ditions across all three imaging modalities. Although both models achieve comparable performance in the standard few-shot tabular setting, T AP-GPT demonstrates significantly greater prediction stability when subjected to iterativ e prompt- ing. Fig. 10. Effect of self-reflection on GPT -4.1-mini and T AP-GPT across modalities. We compare standard prompting to a self-reflection setting in which each model re views and potentially re vises its initial prediction. Consistency under self-reflection is important for multi- agent system (MAS) deployment, where models engage in multiple rounds of critique, discussion, and voting. A candi- date model for MAS must tolerate iterative reasoning without destabilizing its predictions. Although this represents a sim- ple single round of self-reflection, T AP-GPT’ s stability here indicates reduced susceptibility to drift during debate. D . Missingness W e ev aluated the robustness of T AP-GPT to missing data in two contexts on QT -P AD biomarker dataset: (i) controlled simulated missingness applied uniformly across the input table, and (ii) real-world missingness presented in held-out clinical samples. Fig. 11. P erformance of multiple models on simulated missingness in the QT -P AD dataset at inference . T AP-GPT retains high performance until high missingness at 50%. 1) P erformance Under Simulated Missingness : In the sim- ulated missingness setting, we randomly masked 10%, 20%, 30%, 40%, and 50% of feature values across the entire input table provided to each model under a missing-completely- at-random assumption. As shown in Figure 11, T AP-GPT maintained a clear performance advantage over baseline tab- ular LLMs and classical models as missingness increased. Performance declined gradually with increasing masking, but T AP-GPT remained competitiv e through 40% missingness. At the most extreme setting (50% missingness), T abPFN achiev ed performance comparable to T AP-GPT , while other baselines degraded substantially . These results suggest that T AP-GPT is robust to moderate lev els of uniformly distributed missing data and that tool-calling T abPFN may be appropriate in extreme cases of missingness. 2) P erformance Under Naturally Occurr ing Missingness : T o assess robustness under realistic clinical conditions, we next ev aluated T AP-GPT on naturally occurring missingness. Patients from QT -P AD with missing v alues were excluded from both training and prior ev aluation of T AP-GPT and therefore pro vide an independent cohort for this analysis. From this subset, 541 patients with naturally occurring missing data were identified, and 20% were reserved exclusiv ely to serve as an ICL pool, leaving 433 target patients. For each target patient in the ev aluation set, tabular prompts were constructed by randomly sampling k = 8 examples from this pool, which has an av erage missingness of 0.247. Missingness is defined for one patient as number of missing features number of total features . The ICL examples reflect the natural, heterogeneous missingness present in QT - P AD and were not stratified by missingness lev el. All 10 T AP- GPT models, each trained on a different random seed from KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 11 Fig. 12. P erformance across real-world target missingness levels . est samples were grouped b y the propor tion of missing features in the target patient only , while the in-context learning (ICL) pool retained its natural missingness distr ib ution across all patients with missing data (mean missingness = 0.247). the fully observed cohort, were ev aluated on the same set of target patients to characterize robustness across independently trained models. Figure 12 stratifies results by target-le vel missingness, iso- lating the effect of missing data in the patient being ev aluated while preserving a deployment-like prompting scenario. T AP- GPT exhibits stable performance across missingness bins, with limited degradation and relativ ely low variability across seeds. T abPFN maintains similar performance to T AP-GPT across increasing missingness. I V . D I S C U S S I O N In this work, we present T AP-GPT , a domain-adapted tabular-LLM framew ork for fe w-shot AD prediction from mul- timodal biomedical tables, and a comprehensi ve analysis of its ability . In biomedical settings, where datasets are low-sample, partially observed, and multimodal, con ventional deep learning methods often fail to outperform tree-based models. Our work introduces a new paradigm for structured clinical prediction by adapting tabular -specialized large language models to mul- timodal Alzheimer’ s disease biomarker and imaging-derived data. T o our knowledge, this is the first systematic applica- tion of tabular LLMs to multimodal biomedical biomarkers and imaging-derived modalities, showing that structured LLM architectures can bridge prior knowledge, in-context learning, and domain adaptation to address long-standing challenges in clinical tabular modeling. Across experiments spanning one multimodal biomark er dataset (QT -P AD), and three unimodal imaging-deriv ed datasets, T AP-GPT shows strong and consistent performance in low-shot settings. On the QT -P AD dataset, T AP-GPT achiev es particularly strong results in the fe w-shot tabu- lar setting, outperforming vanilla LLMs, traditional machine learning models, and T abPFN. On the imaging datasets, T AP- GPT shows competitive performance across prompt settings, although in some configurations, strong general-purpose LLMs achiev e similar results. GPT -4.1-mini and Qwen3 are both stronger models than the backbone of T AP-GPT , Qwen2.5, but we see that in most cases T AP-GPT keeps up with or surpasses these models. It is possible that moving towards a multimodal analysis of the imaging data w ould further impro ve T AP-GPT’ s performance on these datasets, similar to what we observed in the multimodal dataset QT -P AD, which benefits from multiple biological domains. Our results also highlight the importance of the underly- ing LLM backbone. Even without task-specific finetuning, GPT -4.1-mini achiev ed strong performance on the QT -P AD data, and ev en outperformed T AP-GPT in some imaging experiments. This underscores the growing tabular reason- ing capacity of high-capacity general-purpose LLMs. W ithin our framew ork, howe ver , we observe consistent gains from Qwen2.5 to T ableGPT2 through tab ular specialization, and from T ableGPT2 to T AP-GPT with task-specific fine-tuning, indicating that each stage of specialization contributes mean- ingfully to performance. Although T ableGPT -R1, built on Qwen3, did not perform strongly in our fe w-shot imaging experiments (and we similarly did not observe strong perfor- mance from Qwen3 in this setting) these findings reinforce that backbone selection remains critical. Notably , we did not enable extended “thinking” or reasoning modes for these models, as our objectiv e was constrained to binary classification. At the same time, the fixed architecture of T ableGPT2 limits our ability to replace its decoder with newer , more capable LLMs. Future work could explore reconstructing a similar tabular - LLM architecture using more modern backbones for clinical prediction tasks. An important finding is that T AP-GPT remains competitiv e with T abPFN in the low-shot settings common in biomedical research. T abPFN is specifically designed for few-shot tabular prediction and sets a strong baseline for small structured datasets. In the QT -P AD dataset with 4–10 examples (Figure 5), T AP-GPT performs comparably while also providing inter- pretable outputs. In the 4-shot imaging experiments, T AP-GPT often outperforms T abPFN, depending on the prompt format. Unlike T abPFN, which relies solely on tabular structure, T AP- GPT combines a tabular encoder with a language model decoder , allowing it to use both structural patterns and prior semantic knowledge. This may offer an advantage in clinical settings, where labeled data are scarce and feature names carry meaningful domain information. Another practical advantage of LLM-based prediction mod- els is their flexibility in handling missing data. Unlike tradi- tional machine learning models or tabular foundation models that typically require explicit imputation, T AP-GPT can repre- sent missingness directly within the prompt without artificially filling v alues. In our experiments, T AP-GPT maintained stable performance under both simulated and real-world missingness, remaining competitive up to moderate lev els of missing data and showing limited degradation across target-le vel missing- ness bins. These results suggest that LLM-based models can tolerate incomplete clinical tables without heavy preprocess- ing, which is especially important in real-world biomedical settings where data are often sparse or irregular . Beyond performance, sev eral properties of T AP-GPT posi- tion it as a candidate agent for clinical multi-agent systems. First, its robustness to both real-world and simulated missing- ness in the multimodal setting suggests it can operate under the incomplete and heterogeneous data conditions typical to 12 IEEE TRANSACTIONS AND JOURNALS TEMPLA TE clinical workflows, where different agents may receiv e partial views of a patient’ s record. Second, the model produces interpretable outputs that explicitly reference multimodal ev- idence, demonstrating that it performs cross-modal reasoning rather than relying on isolated signals. Importantly , these explanations can be generated in flexible natural language or constrained structured formats, making them suitable for both clinician-facing interpretation and machine-readable commu- nication between agents. Third, while GPT -4.1-mini achiev ed comparable or stronger performance in some settings, T AP- GPT exhibited greater stability under self-reflection, indicat- ing more consistent reasoning across iterativ e ev aluations. This stability is particularly relev ant for multi-agent chains of communication, where models repeatedly critique, re vise, and exchange intermediate conclusions. Finally , the strongest ov erall performance emerged in the multimodal dataset, while unimodal results remained competiti ve, suggesting a natural architecture in which modality-specific T AP-GPT agents com- municate to simulate collaborativ e clinical decision-making. T ogether, these properties support framing T AP-GPT not merely as a predictor , but as a reasoning module suited for multimodal, communicativ e diagnostic systems. By operating on shared evidence rather than only probability scores, such a system would more closely resemble multidisciplinary clinical decision-making in the real world and provide a pra ctical foun- dation for collaborati ve, multimodal LLM-based diagnostic support. V . C O N C L U S I O N In this work, we introduced T AP-GPT , a tabular large language model framework for few-shot Alzheimer’ s disease prediction on multimodal biomarker and imaging-deri ved data, and demonstrated that tabular -specialized LLMs can effec- tiv ely reason over structured clinical data without serialization. Across multimodal and unimodal datasets, T AP-GPT con- sistently improv ed upon its backbone models, outperformed classical machine learning baselines, and remained competitiv e with dedicated tabular foundation models in low-shot settings, while additionally providing structured, clinically meaningful reasoning. W e sho wed that T AP-GPT maintained robustness under missingness, generated structured and interpretable mul- timodal reasoning, and sho wed stability under self-reflection. T AP-GPT highlights a new direction for integrating multi- modal biomarkers through structured LLM reasoning, laying the groundwork for future multi-agent diagnostic systems in which modality-specific expert models collaborate to produce more reliable and transparent clinical decisions. A C K N OW L E D G M E N T S This work was supported in part by the NIH grants U01 A G066833, P30 A G073105, U01 A G068057, U19 A G074879, U01 AG088658, R01 LM013463 and R01 A G071470; and the NSF grant SCH 2500343. Data collection and sharing for this project were funded by the Alzheimer’ s Disease Neuroimaging Initiati ve (ADNI) (National Institutes of Health Grant U01 A G024904) and DOD ADNI (Department of Defense award number W81XWH-12- 2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineer- ing, and through generous contributions from the following: AbbV ie, Alzheimer’ s Association; Alzheimer’ s Drug Discov- ery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Ei- sai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F . Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Dev el- opment, LLC.; Johnson & Johnson Pharmaceutical Research & Dev elopment LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neu- rotrack T echnologies; Nov artis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; T akeda Pharmaceu- tical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Priv ate sector contributions are facilitated by the Foundation for the National Institutes of Health (www .fnih.org). The grantee or ganization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’ s Therapeutic Research Institute at the Univ ersity of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California. R E F E R E N C E S [1] M. Prince, R. Bryce, E. Albanese, A. Wimo, W . Ribeiro, and C. P . Ferri, “The global prev alence of dementia: A systematic revie w and metaanalysis, ” Alzheimer’s & Dementia , vol. 9, no. 1, pp. 63–75.e2, 2013. [2] E. Nichols et al. , “Estimation of the global prev alence of dementia in 2019 and forecasted prev alence in 2050: an analysis for the global burden of disease study 2019, ” The Lancet Public Health , vol. 7, no. 2, pp. e105–e125, 2022. [3] C. R. Jack, D. A. Bennett et al. , “Nia-aa research framework: T oward a biological definition of alzheimer’ s disease, ” Alzheimer’ s and Dementia , vol. 14, no. 4, pp. 535–562, 2018. [4] C. R. Jack Jr , D. A. Bennett, K. Blennow , M. C. Carrillo, H. H. Feldman, G. B. Frisoni, H. Hampel, W . J. Jagust, K. A. Johnson, D. S. Knopman et al. , “ A/t/n: An unbiased descriptiv e classification scheme for alzheimer disease biomarkers, ” Neur ology , vol. 87, no. 5, pp. 539– 547, 2016. [5] L. Mosconi, M. Brys et al. , “Early detection of alzheimer’ s disease using neuroimaging, ” Experimental Ger ontology , vol. 42, no. 1, pp. 129–138, 2007. [Online]. A vailable: https://www .sciencedirect.com/ science/article/pii/S0531556506001355 [6] N. Hollmann, S. M ¨ uller , K. Eggensperger , and F . Hutter , “T abpfn: A transformer that solves small tabular classification problems in a second, ” arXiv preprint , 2022. [7] S. Dash, S. Bagchi, N. Mihindukulasooriya, and A. Gliozzo, “Permutation in variant strategy using transformer encoders for table understanding, ” in F indings of the Association for Computational Linguistics: NAA CL 2022 , M. Carpuat, M.-C. de Marneffe, and I. V . Meza Ruiz, Eds. Seattle, United States: Association for Computational Linguistics, Jul. 2022, pp. 788–800. [Online]. A vailable: https://aclanthology .org/2022.findings- naacl.59/ [8] Y .-W . Bao, Z.-J. W ang, Y .-F . Shea, P . K.-C. Chiu, J. S. Kwan et al. , “Combined quantitati ve amyloid- β pet and structural mri features improve alzheimer’s disease classification in random forest model-a multicenter study , ” Academic Radiology , vol. 31, no. 12, pp. 5154–5163, 2024. [9] F . Yi, H. Y ang, D. Chen, Y . Qin, H. Han et al. , “Xgboost-shap- based interpretable diagnostic framework for alzheimer’ s disease, ” BMC medical informatics and decision making , vol. 23, no. 1, p. 137, 2023. KEARNEY & Y ANG et al. : T ABULAR LLMS FOR INTERPRET ABLE FEW -SHOT ALZHEIMER’S DISEASE PREDICTION WITH MUL TIMODAL BIOMEDICAL D A T A 13 [10] A. Y . Guo, J. P . Laporte, K. Singh, J. Bae, K. Bergeron et al. , “Ma- chine learning diagnosis of mild cognitiv e impairment using advanced diffusion mri and csf biomarkers, ” Alzheimer’ s & Dementia: Diagnosis, Assessment & Disease Monitoring , vol. 17, no. 3, p. e70182, 2025. [11] Q. Li, L. Cui, Y . Guan, Y . Li, F . Xie, and Q. Guo, “Prediction model and nomogram for amyloid positivity using clinical and mri features in individuals with subjective cognitive decline, ” Human Brain Mapping , vol. 46, no. 8, p. e70238, 2025. [12] L. Grinsztajn, E. Oyallon, and G. V aroquaux, “Why do tree-based models still outperform deep learning on typical tabular data?” Advances in neural information pr ocessing systems , vol. 35, pp. 507–520, 2022. [13] D. McElfresh, S. Khandagale, J. V alverde, V . Prasad C, G. Ramakrish- nan, M. Goldblum, and C. White, “When do neural nets outperform boosted trees on tabular data?” Advances in Neural Information Pro- cessing Systems , vol. 36, pp. 76 336–76 369, 2024. [14] L. Shen and P . M. Thompson, “Brain imaging genomics: Integrated analysis and machine learning, ” Pr oceedings of the IEEE , v ol. 108, no. 1, pp. 125–162, 2020. [15] J. Bao, B. N. Lee, J. W en, M. Kim, S. Mu, S. Y ang, C. Davatzik os, Q. Long, M. D. Ritchie, and L. Shen, “Employing informatics strategies in alzheimer disease research: A revie w from genetics, multiomics, and biomarkers to clinical outcomes, ” Annual Review of Biomedical Data Science , 2024. [16] E. Lin, C.-H. Lin, and H.-Y . Lane, “Deep learning with neuroimaging and genomics in alzheimer’ s disease, ” International journal of molecular sciences , vol. 22, no. 15, p. 7911, 2021. [17] X. Fang, W . Xu, F . A. T an, Z. Hu, J. Zhang, Y . Qi, S. H. Sengamedu, and C. Faloutsos, “Large language models (LLMs) on tabular data: Prediction, generation, and understanding - a survey , ” Tr ansactions on Machine Learning Researc h , 2024. [18] G. Badaro, M. Saeed, and P . Papotti, “Transformers for tabular data representation: A survey of models and applications, ” T ransactions of the Association for Computational Linguistics , vol. 11, pp. 227–249, 2023. [19] N. Hollmann, S. M ¨ uller , L. Purucker , A. Krishnakumar, M. K ¨ orfer , S. B. Hoo, R. T . Schirrmeister, and F . Hutter , “ Accurate predictions on small data with a tabular foundation model, ” Nature , vol. 637, no. 8045, pp. 319–326, 2025. [20] X. Zhang, D. C. Maddix, J. Y in, N. Erickson, A. F . Ansari, B. Han, S. Zhang, L. Akoglu, C. F aloutsos, M. W . Mahoney et al. , “Mitra: Mixed synthetic priors for enhancing tabular foundation models, ” in The Thirty- ninth Annual Conference on Neural Information Pr ocessing Systems . [21] J. Ma, V . Thomas, R. Hosseinzadeh, A. Labach, J. C. Cresswell, K. Golestan, G. Y u, A. L. Caterini, and M. V olkovs, “T abdpt: Scaling tabular foundation models on real data, ” in The Thirty-ninth Annual Confer ence on Neural Information Processing Systems . [22] V . Balazadeh, H. Kamkari, V . Thomas, J. Ma, B. Li, J. C. Cresswell, and R. Krishnan, “Causalpfn: Amortized causal effect estimation via in-context learning, ” in The Thirty-ninth Annual Conference on Neural Information Pr ocessing Systems . [23] T . Zhang, S. W ang, S. Y an, L. Jian, and Q. Liu, “Generativ e table pre-training empowers models for tabular prediction, ” in Proceedings of the 2023 Conference on Empirical Methods in Natural Language Pr ocessing , 2023, pp. 14 836–14 854. [24] Z. W ang, H. Zhang, C.-L. Li, J. M. Eisenschlos, V . Perot, Z. W ang, L. Miculicich, Y . Fujii, J. Shang, C.-Y . Lee et al. , “Chain-of-table: Evolving tables in the reasoning chain for table understanding, ” in The T welfth International Conference on Learning Repr esentations , 2024. [25] S. Han, J. Y oon, S. O. Arik, and T . Pfister, “Large language models can automatically engineer features for few-shot tabular learning, ” in F orty-first International Conference on Machine Learning , 2024. [26] T . Li, S. Shetty , A. Kamath, A. Jaiswal, X. Jiang, Y . Ding, and Y . Kim, “Cancergpt for few shot drug pair synergy prediction using large pretrained language models, ” NPJ Digital Medicine , vol. 7, no. 1, p. 40, 2024. [27] S. Hegselmann, A. Buendia, H. Lang, M. Agrawal, X. Jiang, and D. Sontag, “T abllm: Few-shot classification of tabular data with large language models, ” in International Conference on Artificial Intelligence and Statistics . PMLR, 2023, pp. 5549–5581. [28] J. Lee, S. Y ang, J. Y . Baik, X. Liu, Z. T an, D. Li, Z. W en, B. Hou, D. Duong-Tran, T . Chen et al. , “Knowledge-dri ven feature selection and engineering for genotype data with large language models, ” AMIA Summits on T ranslational Science Pr oceedings , vol. 2025, p. 250, 2025. [29] T . Brown, B. Mann, N. Ryder , M. Subbiah, J. D. Kaplan, P . Dhariwal, A. Neelakantan, P . Shyam, G. Sastry , A. Askell et al. , “Language mod- els are few-shot learners, ” Advances in neural information processing systems , vol. 33, pp. 1877–1901, 2020. [30] H. Zhou, A. Bradley , E. Littwin, N. Razin, O. Saremi, J. Susskind, S. Bengio, and P . Nakkiran, “What algorithms can transformers learn? a study in length generalization, ” arXiv preprint , 2023. [31] S. M ¨ uller , N. Hollmann, S. P . Arango, J. Grabocka, and F . Hutter , “T rans- formers can do bayesian inference, ” arXiv preprint , 2021. [32] V . Thomas, J. Ma, R. Hosseinzadeh, K. Golestan, G. Y u, M. V olkovs, and A. L. Caterini, “Retriev al & fine-tuning for in-context tabular models, ” Advances in Neural Information Processing Systems , vol. 37, pp. 108 439–108 467, 2024. [33] J. W u and M. Hou, “ An efficient retriev al-based method for tabular prediction with llm, ” in Pr oceedings of the 31st International Conference on Computational Linguistics , 2025, pp. 9917–9925. [34] L. Zha, J. Zhou, L. Li, R. W ang, Q. Huang, S. Y ang, J. Y uan, C. Su, X. Li, A. Su et al. , “T ablegpt: T owards unifying tables, nature language and commands into one gpt, ” arXiv pr eprint arXiv:2307.08674 , 2023. [35] A. Su, A. W ang, C. Y e, C. Zhou, G. Zhang, G. Chen, G. Zhu, H. W ang, H. Xu, H. Chen et al. , “T ablegpt2: A large multimodal model with tabular data integration, ” arXiv preprint , 2024. [36] B. Hui, J. Y ang, Z. Cui, J. Y ang, D. Liu, L. Zhang, T . Liu, J. Zhang, B. Y u, K. Lu et al. , “Qwen2. 5-coder technical report, ” arXiv pr eprint arXiv:2409.12186 , 2024. [37] S. Y ang, Q. Huang, J. Y uan, L. Zha, K. T ang, Y . Y ang, N. W ang, Y . W ei, L. Li, W . Y e et al. , “T ablegpt-r1: Advancing tabular reasoning through reinforcement learning, ” arXiv preprint , 2025. [38] A. Y ang, A. Li, B. Y ang, B. Zhang, B. Hui, B. Zheng, B. Y u, C. Gao, C. Huang, C. Lv , C. Zheng, D. Liu, F . Zhou, F . Huang, F . Hu, H. Ge, H. W ei, H. Lin, J. T ang, J. Y ang, J. T u, J. Zhang, J. Y ang, J. Y ang, J. Zhou, J. Zhou, J. Lin, K. Dang, K. Bao, K. Y ang, L. Y u, L. Deng, M. Li, M. Xue, M. Li, P . Zhang, P . W ang, Q. Zhu, R. Men, R. Gao, S. Liu, S. Luo, T . Li, T . T ang, W . Y in, X. Ren, X. W ang, X. Zhang, X. Ren, Y . Fan, Y . Su, Y . Zhang, Y . Zhang, Y . W an, Y . Liu, Z. W ang, Z. Cui, Z. Zhang, Z. Zhou, and Z. Qiu, “Qwen3 technical report, ” 2025. [Online]. A vailable: https://arxiv .org/abs/2505.09388 [39] X. Wu, J. Y ang, L. Chai, G. Zhang, J. Liu, X. Du, D. Liang, D. Shu, X. Cheng, T . Sun et al. , “T ablebench: A comprehensiv e and complex benchmark for table question answering, ” in Proceedings of the AAAI Confer ence on Artificial Intelligence , vol. 39, no. 24, 2025, pp. 25 497– 25 506. [40] T . Dettmers, A. Pagnoni, A. Holtzman, and L. Zettlemoyer , “Qlora: Efficient finetuning of quantized llms, ” Advances in neural information pr ocessing systems , vol. 36, pp. 10 088–10 115, 2023. [41] S. Kearney , S. Y ang, Z. W en, B. Hou, D. Duong-Tran, T . Chen, J. Moore, M. Ritchie, and L. Shen, Enabling F ew-Shot Alzheimer’s Disease Diagnosis on Biomarker Data with T abular LLMs . Ne w Y ork, NY , USA: Association for Computing Machinery , 2025. [Online]. A vailable: https://doi.org/10.1145/3765612.3767229 [42] M. W . W einer, D. P . V eitch, P . S. Aisen, L. A. Beckett, N. J. Cairns, R. C. Green, D. Harvey , C. R. Jack, W . Jagust, E. Liu et al. , “The alzheimer’ s disease neuroimaging initiativ e: a revie w of papers published since its inception, ” Alzheimer’ s & Dementia , vol. 9, no. 5, pp. e111–e194, 2013. [43] M. W . W einer , k. P . V eitch, P . S. Aisen, L. A. Beckett, N. J. Cairns, R. C. Green, D. Harvey , C. R. Jack Jr , W . Jagust, J. C. Morris et al. , “Re- cent publications from the alzheimer’s disease neuroimaging initiativ e: Revie wing progress toward improved ad clinical trials, ” Alzheimer’s & Dementia , vol. 13, no. 4, pp. e1–e85, 2017.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment