When Stability Fails: Hidden Failure Modes Of LLMS in Data-Constrained Scientific Decision-Making
Large language models (LLMs) are increasingly used as decision-support tools in data-constrained scientific workflows, where correctness and validity are critical. However, evaluation practices often emphasize stability or reproducibility across repe…
Authors: Nazia Riasat
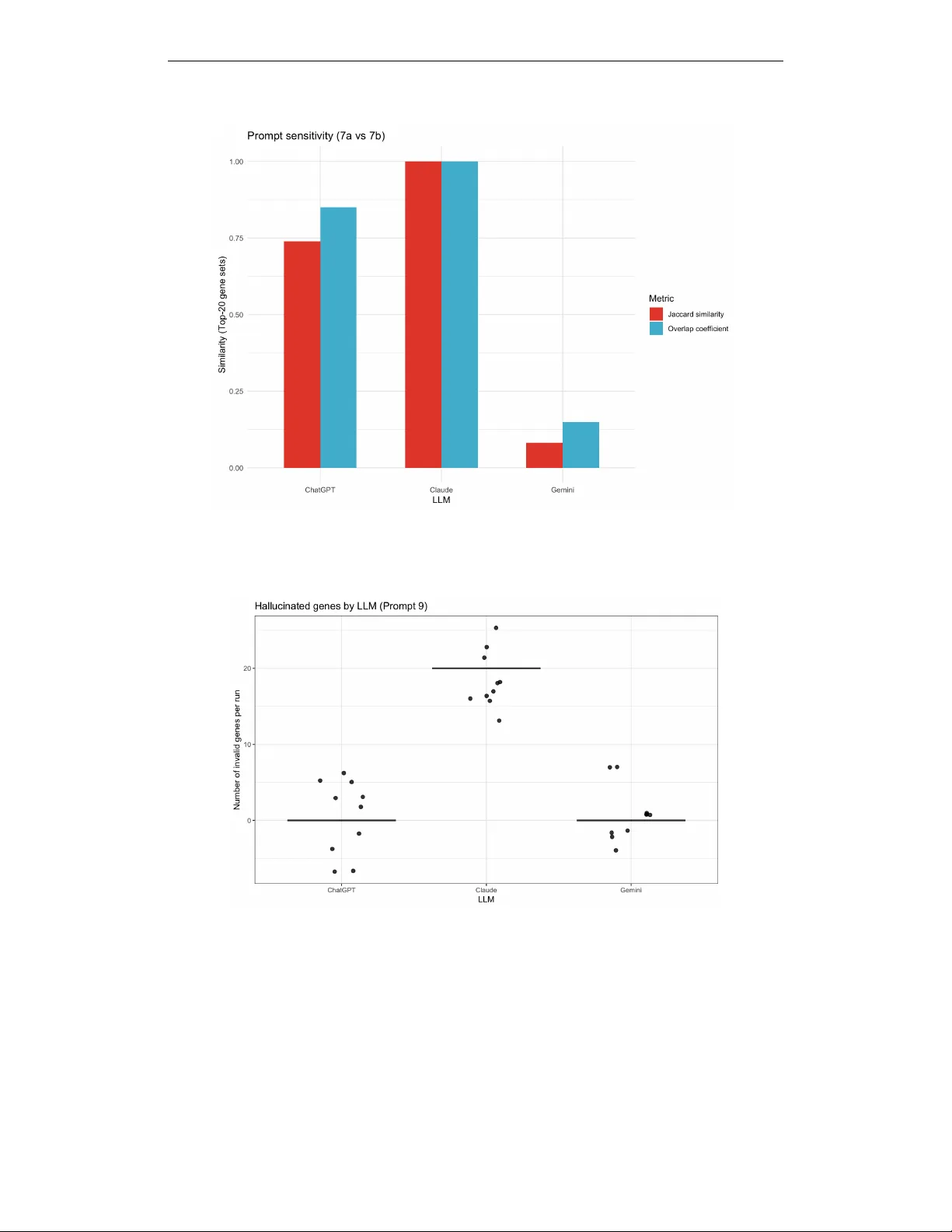
I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 W H E N S T A B I L I T Y F A I L S : H I D D E N F A I L U R E M O D E S O F L L M S I N D A T A - C O N S T R A I N E D S C I E N T I FI C D E C I S I O N - M A K I N G Nazia Riasat North Dakota State Uni versity nazia.riasat@ndsu.edu A B S T R AC T Large language models (LLMs) are increasingly used as decision-support tools in data-constrained scientific workflo ws, where correctness and validity are critical. Howe ver , evaluation practices often emphasize stability or reproducibility across repeated runs. While these properties are desirable, stability alone does not guar- antee agreement with statistical ground truth when such references are av ailable. W e introduce a controlled beha vioral ev aluation framew ork that explicitly sep- arates four dimensions of LLM decision-making: stability , correctness, prompt sensitivity , and output validity under fixed statistical inputs. W e ev aluate multi- ple LLMs using a statistical gene prioritization task derived from differential ex- pression analysis across prompt regimes in volving strict and relaxed significance thresholds, borderline ranking scenarios, and minor wording variations. Our ex- periments sho w that LLMs can exhibit near -perfect run-to-run stability while sys- tematically div erging from statistical ground truth, over -selecting under relaxed thresholds, responding sharply to minor prompt wording changes, or producing syntactically plausible gene identifiers absent from the input table. Although sta- bility reflects robustness across repeated runs, it does not guarantee agreement with statistical ground truth in structured scientific decision tasks. These findings highlight the importance of explicit ground-truth validation and output validity checks when deploying LLMs in automated or semi-automated scientific work- flows. 1 I N T RO D U C T I O N Large language models (LLMs) are increasingly e xplored as decision-support tools in scientific and biomedical workflo ws, including tasks such as data interpretation, hypothesis generation, and candi- date gene prioritization (Singhal et al., 2023; OpenAI, 2023). In bioinformatics settings, LLMs are often applied to summarize or rank outputs from statistical analyses, where their fluent reasoning and apparent consistency across repeated runs can encourage trust among users. Ho wev er , ev alu- ation practices frequently emphasize stability or reproducibility across repeated runs as indicators of reliability . While stability is generally desirable, it does not necessarily imply correctness in structured scientific decision tasks. In many machine learning settings, stability is vie wed as an indicator of robustness to perturbations in data or model inputs. Howe ver , stability alone cannot substitute for explicit validation when reliable statistical references exist. When the underlying sta- tistical reference is kno wn, stable model outputs may still div erge systematically from statistically valid conclusions. In this work, we inv estigate this phenomenon through a controlled e valuation of LLM behavior in a statistical gene prioritization task deri ved from dif ferential expression anal- ysis. In differential expression studies, statistical methods identify genes whose expression differs significantly between experimental conditions. T ools such as DESeq2 estimate gene-level effect sizes and statistical significance while controlling the false discovery rate (FDR) (Lov e et al., 2014; Benjamini & Hochberg, 1995). Here we treat the set of genes identified by a deterministic DESeq2 analysis as a fixed statistical reference that enables controlled comparison of LLM outputs under identical inputs. Using a fixed differential e xpression table as input, we query multiple LLMs across prompt regimes that vary statistical thresholds, borderline selection scenarios, and prompt word- ing. W e e valuated each configuration over repeated runs to measure stability , agreement with the 1 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 statistical reference, prompt sensitivity , and output v alidity . This design isolates model behavior from data variability and enables systematic identification of failure modes that may remain hidden in more open-ended LLM ev aluations. Importantly , our analysis does not challenge the value of stability itself, but demonstrates that stability alone can be insufficient for reliable decision-making when ground-truth statistical references are av ailable. Our results rev eal sev eral characteristic be- haviors. First, models often produce highly stable outputs across repeated runs while exhibiting low agreement with the statistical reference. Second, small wording changes in prompts can alter gene prioritization outcomes, reflecting shifts in how models interpret statistical objectiv es. Third, relaxed statistical thresholds encourage over -selection relati ve to the reference procedure. Finally , models may produce syntactically plausible but in valid gene identifiers that do not appear in the input table. Collecti vely , these results indicate that stability should be interpreted as a complemen- tary diagnostic rather than a proxy for correctness in data-constrained scientific w orkflows. Figure 1 summarizes the e xperimental setup and primary failure modes observed in our e valuation. W e there- fore introduce a controlled behavioral e valuation frame work that separates four dimensions of LLM decision-making: stability , correctness relati ve to a statistical reference, prompt sensiti vity , and out- put validity . 2 E X P E R I M E N T A L S E T U P A N D E V A L UA T I O N W e study LLM behavior in a controlled statistical gene prioritization task deriv ed from differential expression (DE) analysis. In this setting, genes are ranked according to significance estimates pro- duced by the DESeq2 statistical model. A single fixed differential expression table is provided as input to all models, and no external biological knowledge or contextual information is permitted. Differential e xpression statistics are computed using standard normalization, dispersion estimation, and multiple-testing correction procedures (Ji et al., 2023; Benjamini & Hochberg, 1995). W e ev al- uate three large language models: ChatGPT (GPT -5.2), Google Gemini 3, and Claude Opus 4.5, across se veral prompt regimes reflecting common analytical scenarios. These include strict statis- tical thresholding ( FDR ≤ 0 . 05 ), relaxed thresholding ( 0 . 05 < FDR ≤ 0 . 10 ), borderline T op-20 selection, and ranked T op-20 prioritization. T o assess prompt sensiti vity , we construct two semanti- cally equiv alent prompts that dif fer only in wording (P7a vs. P7b). Although both prompts encode the same decision criteria, minor dif ferences in emphasis (e.g., statistical significance v ersus ranking instructions) may introduce subtle shifts in task interpretation. Each configuration is ev aluated over ten repeated runs with identical inputs. Model beha vior is assessed along four complementary ev al- uation dimensions. First, we measure run-to-run stability of model outputs using the Jaccard index (Jaccard, 1901), which quantifies similarity between gene sets produced across repeated executions. Because predicted gene sets may v ary in size across prompt regimes, we additionally report the ov er- lap coefficient (Szymkiewicz–Simpson coefficient) (V ijaymeena & Kavitha, 2016), a containment- based similarity measure that captures cases where one set is largely included within another . T o ev aluate correctness, we compare model outputs against a deterministic statistical reference deri ved from DESeq2 differential expression analysis (Lov e et al., 2014). Prompt sensiti vity is quantified by measuring div ergence between outputs generated from semantically equiv alent prompts (Zhu et al., 2023). Finally , we examine output validity by identifying gene identifiers produced by the model that do not appear in the input table, reflecting hallucination-related failure modes previously doc- umented in language models (Li et al., 2024; Kaddour et al., 2023). In this study , the DESeq2 out- put serves as a deterministic statistical reference that enables controlled comparisons across model outputs. While borderline FDR regimes may introduce ambiguity in biological interpretation, the reference functions here as a consistent e valuation baseline rather than a claim of absolute biological ground truth. Code and e xperimental outputs are pro vided in the supplementary repository described in Appendix H4. 3 R E S U L T S Failure Modes in LLM-Based Gene Prioritization: W e organize the results by failure mode rather than by individual prompt so that stability , agreement with the DESeq2-deri ved statistical reference, prompt sensitivity , and output v alidity can be compared directly under identical inputs. 2 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 (A) Experimental setup Same DE table → 3 LLMs (ChatGPT , Gemini, Claude), 10 r uns Regimes: FDR ≤ 0.05, 0.05 < FDR ≤ 0.10, border line T op-20, wording (7a vs 7b), ranked T op-20 (B) Stability = correctness Identical runs (Jaccard ≈ 1) can still disagree with ground truth. (D) Hallucinated identifiers Outputs may include gene IDs ab- sent from the input table. (C) Prompt sensitivity Minor wording changes yield different T op-20 sets. (E) Failure modes Over-selection, prompt sensitivity , hallucination. Figure 1: Failure modes in LLM-based statistical gene prioritization under fixed input data. Despite high internal stability , models may disagree with the DESeq2-derived statistical reference (B), ex- hibit prompt sensiti vity (C), or produce in valid gene identifiers (D), Panel (E) summarizes observed behaviors. ChatGPT Claude Gemini 0.00 0.25 0.50 0.75 1.00 0.50 0.75 1.00 1.25 1.50 Within−LLM stability (Jaccard) Agreement with ground truth Stability does not imply correctness Figure 2: Stability does not imply correctness. Within-LLM stability (pairwise Jaccard similarity across repeated runs; x-axis) versus agreement with the DESeq2-deriv ed statistical reference (Jac- card against ground truth; y-axis). Each point represents the mean stability and agreement values aggregated across 10 repeated runs per configuration. Run-lev el variability was minimal due to near - perfect within-model stability . 3 . 1 S TA B I L I T Y D O E S N O T I M P LY C O R R E C T N E S S A C R O S S L L M S Across all e valuated prompt regimes, the three large language models exhibit near-perfect run-to-run stability . In threshold-based tasks (e.g., FDR ≤ 0 . 05 and 0 . 05 < FDR ≤ 0 . 10 ), repeated queries under fixed inputs typically produce identical outputs. Howe ver , while this apparent determinism reflects high internal stability , it does not imply correctness. When compared against the DESeq2- deriv ed statistical reference, stable outputs may still show low or e ven zero agreement with the statistical reference. A Jaccard value of zero indicates that the predicted gene set shares no identifiers with the reference set; this may arise either from returning an empty set or from selecting genes entirely outside the ground-truth set. As shown in Figures 2 and 3, models achie ve maximal within- LLM stability while e xhibiting substantial variation in agreement with the reference. Under relaxed thresholds ( 0 . 05 < FDR ≤ 0 . 10 ), corresponding to the borderline discov ery regime described in Section 2, some models consistently return lar ge gene sets that only partially ov erlap with DESeq2- significant genes, while others return incorrect or empty selections. These results demonstrate that run-to-run consistency alone is insuf ficient as evidence of correct statistical reasoning. 3 . 2 P R O M P T R E G I M E A N D W O R D I N G S E N S I T I V I T Y U N D E R I D E N T I C A L I N P U T S In addition to the Jaccard index, we report the overlap coefficient to capture containment relation- ships between gene sets of dif ferent sizes. T o assess sensiti vity to prompt formulation, we ev aluated model outputs across multiple prompt regimes while holding the underlying differential expression 3 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 table fixed. Figure 3 summarizes mean Jaccard similarity of T op-20 gene sets across threshold- based selection and borderline prioritization regimes, while Figure 4 isolates sensitivity to wording variants. Despite identical inputs, models exhibit marked dif ferences across prompt regimes, re- flecting systematic shifts rather than stochastic variability . W e further isolate the effect of wording by comparing two semantically similar prompts (Prompt P7a vs. P7b) that differ only in whether statistical significance or effect size is emphasized. Although Prompt P7a and Prompt P7b are se- mantically aligned and operate on the same borderline gene set, they differ slightly in emphasis, so the observed dif ferences may reflect both linguistic sensitivity and subtle shifts in instructed prioriti- zation criteria. These prompts operate on the same borderline gene set and request a ranked T op-20 list; ne vertheless, small wording changes can lead to substantial dif ferences in selected gene sets for some models, with overlap ranging from near -complete agreement to minimal intersection. Notably , this prompt sensitivity persists ev en when each prompt individually produces highly stable outputs across repeated runs. This decoupling between internal stability and cross-prompt consistency sug- gests that prompt phrasing may function as an implicit decision v ariable rather than a purely neutral instruction. 0.00 0.25 0.50 0.75 1.00 P1 (FDR<=0.05) P5 (FDR<=0.10) P6 (Borderline T op−20) P7 (7a vs 7b) Prompt Mean Jaccard similarity llm ChatGPT Claude Gemini LLM behavior across prompt regimes Figure 3: LLM behavior across prompt regimes. Mean Jaccard similarity to DESeq2 reference of T op-20 gene sets across threshold-based selection, borderline prioritization, and prompt wording variants. 3 . 3 O V E R - S E L E C T I O N U N D E R R E L A X E D T H R E S H O L D S Relaxing the significance threshold from FDR ≤ 0 . 05 (P1) to 0.05 < FDR ≤ 0 . 10 (P5) system- atically shifts LLM behavior from selective filtering to ward permissiv e inclusion. As summarized in T able 1, agreement with the DESeq2-deriv ed reference under P5 remains limited despite larger returned sets: ChatGPT attains Jaccard = 0 . 47 , Gemini drops to 0 . 28 , and Claude collapses to 0 . 00 . The apparent gains in recall therefore reflect over -selection or degenerate inclusion rather than im- prov ed discrimination. This pattern extends to ranking under uncertainty . In the borderline T op-20 setting (P6), Gemini matches the deterministic reference (Jaccard = 1 . 00 ), while ChatGPT di verges substantially (Jaccard = 0 . 14 ) and Claude ag ain returns no true signal. Notably , all models maintain perfect within-LLM stability across repeated runs in this setting (pairwise Jaccard = 1 . 00 ; T able 1), reinforcing that internal reproducibility does not imply ef fectiv e prioritization. Overall, relaxed statistical criteria may function as a failure trigger for LLM-based gene prioritization, promoting broad inclusion or collapse rather than principled sensitivity–specificity trade-offs. Apparent im- prov ements in sensitivity under relaxed thresholds should therefore not be interpreted as e vidence of more reliable selection. Overlap coef ficient values follow the same qualitati ve pattern as the Jaccard results across prompt regimes. In particular , the ordering of models is unchanged, indicating that the observed discrepancies are not merely a consequence of unequal set sizes b ut reflect substantiv e differences in the selected gene sets rather than artif acts of set size. 3 . 4 H A L L U C I NAT E D G E N E I D E N T I FI E R S W e next e valuate output validity by checking whether returned gene identifiers appear in the pro- vided dif ferential expression table (P9). T able 1 shows that ChatGPT and Gemini consistently re- strict outputs to valid identifiers (0 in valid genes/run), whereas Claude systematically produces in- valid gene IDs (20 in valid genes/run). The frequenc y of hallucinated identifiers across repeated runs is illustrated in Figure 5. Importantly , this failure mode is not explained by stochastic sampling vari- 4 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 T able 1: Summary of LLM behavior across prompt regimes, illustrating the separation between stability , agreement with ground truth, prompt sensitivity , and output validity . Prompt T ask T ype Metric ChatGPT Gemini Claude Interpretation P1 (FDR ≤ 0.05) Thresholded DE Jaccard vs truth 1.00 1.00 0.00 Claude fails to recov er DE genes P5 (FDR ≤ 0.10) Relax ed threshold Jaccard vs truth 0.47 0.28 0.00 Over-selection / collapse P6 (Borderline) Ranking uncertainty Jaccard vs truth 0.14 1.00 0.00 Gemini recovers truth P6 (stability) W ithin-LLM Pairwise Jaccard 1.00 1.00 1.00 Perfect internal stability P7a vs P7b Prompt sensitivity Jaccard 0.74 0.08 1.00 High wording sensitivity P9 (ranking) V alidity check In valid gene IDs per run 0 0 20 Hallucinated identifiers ability: in valid identifiers recur across repeated runs and take the form of syntactically well-formed, gene-like tokens absent from the input table, indicating a systematic violation of the input-domain constraint rather than occasional formatting noise, consistent with prior observations of systematic hallucination in constrained generation tasks (Li et al., 2024). This finding highlights output validity as a distinct e valuation dimension. Even when model outputs are highly stable across repeated runs, they may still violate basic input-domain constraints. In other words, internal consistency does not guarantee that generated entities are supported by the underlying data. The recurrence of such iden- tifiers further distinguishes stability from validity: outputs can remain internally consistent while nonetheless introducing entities that are absent from the input table. For data-constrained scien- tific workflo ws, these hallucinated identifiers represent a qualitativ ely different risk. They suggest that e valuation based solely on stability or overlap with a statistical reference may be insufficient, and that explicit validity checks are necessary to ensure that model outputs remain grounded in the provided data. 4 D I S C U S S I O N A N D C O N C L U S I O N Across prompt regimes, we find that stability , reproducibility , and internal consistency alone is an insufficient proxy for correctness in data-constrained scientific workflo ws.While stability is gener- ally viewed as a desirable property reflecting robustness across repeated runs, our results show that stability alone can be insufficient as evidence of correctness when reliable statistical references are av ailable. Models frequently exhibit near -perfect run-to-run stability while showing poor agreement with the DESeq2-deri ved statistical reference, indicating that deterministic beha vior may reflect internal consistency rather than reliable statistical reasoning. Small changes in prompt wording induce large shifts in prioritization outcomes ev en under identical inputs, while relaxed statistical thresholds systematically promote ov er-selection rather than principled sensiti vity–specificity trade- offs, consistent with documented prompt sensitivity in LLMs (Zhu et al., 2023; Gendron et al., 2023). Critically , some models generate syntactically plausible gene identifiers absent from the in- put data despite high internal stability , violating basic input-domain constraints. This failure mode is systematic and aligns with prior evidence of hallucination in constrained generation tasks (Li et al., 2024), reinforcing broader concerns about the reliability of foundation models in high-stakes decision-making contexts (Bommasani et al., 2022; Zhu et al., 2023). Ov erall, our results disen- tangle four behavioral dimensions, stability , correctness, sensitivity , and validity that are often con- flated in informal LLM e valuations. For data-constrained scientific tasks, internal consistenc y alone is an insufficient proxy for correctness, motiv ating explicit ground-truth-based e valuation and input- domain validity checks. Practically , these findings suggest that LLM-assisted scientific pipelines require explicit ground-truth validation and input-domain checks, rather than reliance on output sta- bility or internal consistency alone. Full prompt specifications and additional figures are provided in Appendices A and C. While the e valuation focuses on a single differential-e xpression dataset and statistical paradigm, the framework provides a controlled setting for diagnosing LLM behavior in data-constrained scientific workflows. Extending the analysis to additional datasets and statistical paradigms is an important direction for future work. 5 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 R E F E R E N C E S Y oa v Benjamini and Y osef Hochberg. Controlling the false disco very rate: A practical and po werful approach to multiple testing. Journal of the Royal Statistical Society: Series B (Methodological) , 57(1):289–300, 1995. doi: https://doi.org/10.1111/j.2517- 6161.1995.tb02031.x. Rishi Bommasani, Dre w A. Hudson, Ehsan Adeli, et al. On the opportunities and risks of foundation models, 2022. Gaël Gendron, Qiming Bao, et al. Large language models are not strong abstract reasoners, 2023. T om Hope, Doug Do wney , et al. A computational inflection for scientific discov ery . Communica- tions of the A CM , 2022. doi: 10.1145/3576896. Paul Jaccard. Étude comparativ e de la distribution florale dans une portion des alpes et des jura. Bulletin de la Société V audoise des Sciences Naturelles , 37:547–579, 1901. doi: https://doi.org/ 10.5169/seals- 266450. Ziwei Ji et al. Survey of hallucination in natural language generation. ACM Computing Surveys , 2023. doi: https://doi.org/10.1145/3571730. Jean Kaddour , Joshua Harris, et al. Challenges and applications of lar ge language models, 2023. Y ifan Li et al. Hallucinations in lar ge language models: A survey , 2024. Michael I. Love, W olfgang Huber, and Simon Anders. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biolo gy , 15(12):550, 2014. doi: https: //doi.org/10.1186/s13059- 014- 0550- 8. OpenAI. Gpt-4 technical report. T echnical report, OpenAI, 2023. Marco T ulio Ribeiro, T ongshuang W u, et al. Beyond accuracy: Behavioral testing of nlp models. In Pr oceedings of the 58th Annual Meeting of the Association for Computational Linguistics , 2020. URL https://aclanthology.org/2020.acl- main.442/ . Karan Singhal et al. Large language models encode clinical kno wledge. Nature , 2023. doi: https: //doi.org/10.1038/s41586- 023- 06455- 0. M.K. V ijaymeena and K. Ka vitha. A surve y on similarity measures in text mining. Machine Learn- ing and Applications: An International Journal , 2016. doi: https://doi.org/10.5121/mlaij.2016. 3103. Shichao Zhu, Y ong Chen, Zhe Gan, Jian W ang, W einan Zhang, et al. Promptbench: T owards ev aluating the robustness of large language models on adv ersarial prompts, 2023. 6 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 A P RO M P T S P E C I FI C A T I O N S This appendix provides the full set of prompts used in our experiments. All prompts operate on the same fixed differ ential expr ession table produced using the DESeq2 statistical model, where genes are ranked based on statistical significance (false discovery rate, FDR) and ef fect size estimates. No external biological kno wledge, pathway information, or gene annotations were permitted in any prompt (Singhal et al., 2023). For presentation clarity , prompt text is sho wn in fix ed-width verbatim format and line-broken to fit within the page margins without altering semantic content. Prompt groups correspond directly to the e valuation regimes (P1–P9) introduced in Section 2 and e valuated in Section 3. F or clarity , the appendix presents the full prompt text used in each regime while maintaining the same notation used throughout the main paper . Differential expression statistics were computed using the DESeq2 model, which serves as the deterministic statistical reference used for ev aluation in the main analysis (Love et al., 2014). A . 1 C O M M O N I N P U T F O R M AT All prompts were provided with a table of differential expression results containing the following columns: • gene • log2FoldChange • lfcSE • stat • pvalue • padj • baseMean These columns correspond to the standard output fields of the DESeq2 differential expression frame- work. Unless otherwise stated, models were instructed to use only the supplied table and to base all decisions solely on the reported statistics without inferring biological function or pathway relev ance. A . 2 G R O U P 1 : S T R I C T S I G N I FI C A N C E F I LT E R I N G Prompt P1 (FDR ≤ 0.05). You are given a table of differential expression results. Task: 1. Identify all genes that are statistically significant at FDR <= 0.05. 2. Return ONLY a JSON object with the following keys: - "n_significant": integer - "significant_genes": array of gene names (strings), sorted alphabetically Rules: - Use ONLY the provided table. - Use the "padj" column for FDR. - Do NOT add or remove genes. - Do NOT include explanations. A . 3 G R O U P 2 : R E L A X E D / B O R D E R L I N E S I G N I FI C A N C E Prompt P5 (0.05 < FDR ≤ 0.10). You are given a table of differential expression results. Task: 1. Identify all genes with adjusted p-values at 0.05 < FDR <= 0.10. 2. Return ONLY a JSON object with the following keys: - "n_significant": integer - "significant_genes": array of gene names (strings), sorted alphabetically Rules: 7 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 - Use ONLY the provided table. - Use the "padj" column for FDR. - Do NOT add or remove genes. - Do NOT include explanations. A . 4 G R O U P 3 : R A N K E D P R I O R I T I Z A T I O N O F B O R D E R L I N E G E N E S These prompts operate on a subset of 127 borderline genes with adjusted p-v alues between 0.05 and 0.15. The explanatory “rule” field is included for interpretability but is not used in any quantitativ e ev aluation. Prompts P7a and P7b are wording variants of the borderline ranking task (P6) designed to ev aluate prompt sensitivity while keeping the underlying task objecti ve identical (see Section 3.3). Prompt P6 (Balanced ranking). You are given DE results for 127 borderline genes with adjusted p-values at 0.05 < FDR <= 0.15. Task: Select the top 20 genes that you would prioritize as "most likely true positives" for follow-up, using both effect size (|log2FoldChange|) and statistical evidence (padj). Return ONLY valid JSON: { "top20_genes": [...], "rule": "one sentence describing how you prioritized" } Rules: - Use ONLY the table. - No outside biology. Prompt P7a (Balanced e vidence wording). You are given DE results for 127 borderline genes with adjusted p-values at 0.05 < FDR <= 0.15. Task: Select the top 20 genes that you would prioritize as "most likely true positives" for follow-up, using both effect size (|log2FoldChange|) and statistical evidence (padj). Return ONLY valid JSON: { "top20_genes": [...], "rule": "one sentence describing how you prioritized" } Rules: - Use ONLY the table. - No outside biology or prior knowledge. - Output exactly 20 unique genes. - Do not include any text outside the JSON. Prompt P7b (Effect-size dominant w ording). You are given DE results for 127 borderline genes with adjusted p-values at 0.05 < FDR <= 0.15. Task: Select the TOP 20 genes you would prioritize as "most likely true positives". Prioritize PRIMARILY by effect size (larger |log2FoldChange| is better), and use padj ONLY to break ties or near-ties. Return ONLY valid JSON with exactly these keys: { "top20_genes": [...], "rule": "one sentence describing how you prioritized" } Rules: - Use ONLY the table (no outside biology or prior knowledge). - Output exactly 20 unique genes. - Do not include any text outside the JSON. 8 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 While P7a and P7b differ only slightly in wording emphasis, they implicitly prioritize different objectiv es (effect size vs statistical significance), illustrating how small prompt v ariations can shift task interpretation. A . 5 G R O U P 4 : R A N K E D O U T P U T W I T H E X P L I C I T O R D E R I N G Prompt P9 (Explicit ranking). You are given DE results for 127 borderline genes with adjusted p-values at 0.05 < FDR <= 0.15. Task: Select the TOP 20 genes you would prioritize as “most likely true positives” for follow-up, using BOTH: (1) statistical evidence (smaller padj is better), and (2) effect size (larger |log2FoldChange| is better). Return ONLY valid JSON: { "ranked_top20": [ {"rank": 1, "gene": "GENEID"}, ... {"rank": 20, "gene": "GENEID"} ], "rule": "one sentence describing how you prioritized" } Rules: - Use ONLY the table (no outside biology). - Ranks must be 1..20 with no gaps. - Genes must be unique. - Do not include any text outside the JSON. A . 6 G R O U P 5 : P R O M P T S E N S I T I V I T Y V A R I A N T S T o assess prompt sensiti vity , semantically similar prompts were constructed that dif fered only in em- phasis (e.g., statistical significance versus ef fect size, or ranking versus selection) while preserving the same underlying task objectiv e. These variants were otherwise identical in input data, output for- mat, and constraints. The prompt variants correspond to the sensiti vity analyses reported in Figure 3 (regime comparison) and Figure 4 (prompt wording sensiti vity). A . 7 N O T E S O N R E P E A T E D R U N S Each prompt was executed multiple times per model under identical conditions to assess run-to- run stability . Any dif ferences across repeated outputs using a fixed prompt therefore reflect model stochasticity or internal decision v ariability (e.g., sampling effects) rather than changes in input data or task specification. B G RO U N D - T R U T H C O N S T R U C T I O N F R O M D I FF E R E N T I A L E X P R E S S I O N Ground-truth gene sets were constructed deterministically from the same differential expression analysis used as input to the LLMs. Dif ferential expression was performed using DESeq2 with standard normalization, dispersion estimation, and multiple-testing correction. For threshold-based tasks, genes with adjusted p-v alues (Benjamini–Hochberg FDR) below the specified cutoff were treated as true positi ves. For relaxed or borderline regimes, genes with adjusted p-values within the designated interval were retained as candidate sets for ranking or prioritization tasks but were not treated as ground-truth positives. For ranking-based e valuations, genes were ordered by adjusted p- value in ascending order, with ties broken deterministically . This ordering defines a fixed reference ranking against which model-produced ranked lists were compared. Importantly , the ground-truth construction is entirely data-driv en and does not incorporate external biological kno wledge, pathw ay annotations, or functional priors, ensuring alignment with the information av ailable to the models. This construction ensures that any disagreement reflects model behavior rather than ambiguity in the reference construction. 9 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 Figure 4: Prompt sensitivity under minor wording changes (Prompt 7a vs. 7b). Jaccard similarity and ov erlap coefficient between T op-20 gene sets produced under semantically similar prompts dif- fering only in emphasis. Both metrics exhibit the same qualitativ e pattern, indicating that prompt wording can alter selected gene sets e ven when accounting for dif ferences in set containment. Figure 5: Evaluates validity in ranked gene lists. ChatGPT and Gemini consistently return identifiers present in the input table. In contrast, Claude frequently outputs ranked lists composed largely of identifiers absent from the input table under this prompt configuration. C A D D I T I O N A L P R O M P T S E N S I T I V I T Y A N A L Y S E S W e conducted additional analyses to verify that the observed failure modes persist across repeated runs and prompt regimes. For each prompt configuration, models were queried repeatedly under identical conditions to assess run-to-run stability . Stability was quantified using pairwise Jaccard similarity and ov erlap coefficient between gene sets returned across repeated runs. Both metrics showed consistent qualitativ e patterns, indicating that the observed stability and sensitivity ef fects are not driven solely by differences in set size. These analyses confirm that the observed stability and 10 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 T able 2: Overlap coefficient between LLM-selected gene sets and the DESeq2 reference across prompt regimes. V alues follow the same qualitativ e trends observed with Jaccard similarity but additionally provide a containment-based measure that remains stable when returned gene-set sizes differ . Prompt T ask T ype Metric ChatGPT Gemini Claude Interpretation P1 (FDR ≤ 0.05) Thresholded DE Overlap coef ficient - - 0.00 Fails to recov er DE genes P5 (FDR ≤ 0.10) Relaxed threshold Overlap coef ficient 0.74 1.00 0.00 Claude returns no overlapping genes P6 (Borderline) Ranking uncertainty Overlap coef ficient 0.25 1.00 0.00 Gemini recovers truth P6 (stability) W ithin-LLM Pairwise ov erlap coefficient 1.00 1.00 1.00 Perfect internal stability P7 (7a vs 7b) Prompt sensiti vity Overlap coef ficient 0.85 0.15 1.00 High wording sensitivity Note: “–” indicates metrics that are not defined for that evaluation setting (N A). prompt sensitivity ef fects persist across regimes and repeated runs. W e further examined sensitivity across semantically similar prompt v ariants that dif fer only in emphasis (e.g., effect size versus statistical significance, selection versus ranking). Consistent with the main results, small wording changes frequently produced large shifts in selected gene sets, even when internal run-to-run stability within models remained high. T aken together , these supplementary analyses reinforce that the observed behaviors reflect system- atic properties of model decision-making rather than artifacts of stochastic sampling or isolated prompt formulations. D E M P I R I C A L C O M PA R I S O N O F OV E R L A P C O E FFI C I E N T A N D J AC C A R D T o complement the Jaccard-based analyses reported in the main paper, we also computed the overlap coefficient between LLM-selected gene sets and the DESeq2-deriv ed reference sets across prompt regimes. Unlike Jaccard similarity , the overlap coef ficient measures set containment and is particu- larly informativ e when the sizes of compared gene sets differ . T able 2 reports overlap coef ficient values for each prompt re gime and model. The results follo w the same qualitativ e trends observed with Jaccard similarity in the main analysis. In particular , regimes exhibiting strong agreement or strong div ergence under Jaccard sho w the same patterns under the ov erlap coef ficient. This consistency indicates that the observed discrepancies are not driven solely by differences in gene-set size b ut reflect substantive dif ferences in the selected gene sets. E M E T R I C D E FI N I T I O N S Jaccard Similarity . For two gene sets A and B , Jaccard similarity is defined as J ( A, B ) = | A ∩ B | | A ∪ B | . This metric is used to quantify both run-to-run stability and overlap between model-selected and reference gene sets (Jaccard, 1901). Overlap Coefficient. F or two gene sets A and B , the overlap coef ficient is defined as O ( A, B ) = | A ∩ B | min( | A | , | B | ) . Unlike Jaccard similarity , the ov erlap coefficient measures containment between sets and is particu- larly informativ e when comparing sets of unequal size. Agreement with Ground T ruth. Agreement with ground truth is computed as the Jaccard simi- larity between the set of genes selected by a model and the DESeq2-deriv ed reference set under the corresponding e valuation regime. This metric is e valuated under the same prompt-specific regime used to construct the reference. Run-to-Run Stability . Run-to-run stability is measured as the mean pairwise Jaccard similarity across repeated outputs of the same prompt and model under identical input data. 11 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 Prompt Sensitivity . Prompt sensitivity is assessed by comparing gene sets produced by semanti- cally similar prompts using Jaccard similarity and ov erlap coefficient, isolating the ef fect of prompt wording from input data v ariation. F R E L A T E D W O R K A N D C O N T E X T Prior work has documented hallucination and inv alid output generation in large language models (Gendron et al., 2023) across a range of tasks, including clinical reasoning (Singhal et al., 2023), general natural language generation, and large-scale model ev aluation (Ribeiro et al., 2020). These findings motiv ate careful e valuation of LLM outputs in statistically constrained scientific workflows, where correctness and input validity are critical (Hope et al., 2022). Our work complements this literature by focusing on controlled statistical decision tasks with fully specified ground truth. G R E P R O D U C I B I L I T Y N OT E All experiments were conducted using a fixed differential expression table deriv ed from a single DESeq2 analysis. For each prompt configuration, models were queried repeatedly using identical inputs to assess stability , correctness, prompt sensiti vity , and output validity , enabling measure- ment of within-prompt stability across repeated runs. Outputs were parsed automatically to extract selected gene sets, which were then compared against a deterministic DESeq2-deri ved reference using Jaccard similarity . No external biological kno wledge, tool use, or retriev al was permitted. All prompt templates are provided v erbatim in Appendix A. H M O D E L S , D A TA , A N D C O M P U TA T I O N A L E N V I R O N M E N T H . 1 M O D E L S W e ev aluate three widely used large language models: ChatGPT (GPT -5.2), Google Gemini 3, and Claude Opus 4.5. All models were accessed through their publicly av ailable API endpoints using deterministic decoding settings (temperature = 0). Experiments were conducted using default API configurations, and token limits were not manually constrained beyond the platform defaults. No fine-tuning, system-lev el customization, retriev al augmentation, or external tools were used. Each model was queried repeatedly under identical inputs to measure run-to-run stability and sensitivity to prompt formulation. H . 2 D A TA All experiments use a single fixed differential expression table derived from RN A-seq data from the GEO dataset (GSE239514), comprising non–small cell lung cancer (NSCLC) tumor sam- ples and tumor-draining lymph node (TDLN) samples. Raw HTSeq count matrices provided by the original study were processed using a deterministic DESeq2 pipeline (Lov e et al., 2014), with Benjamini–Hochberg false discovery rate (FDR) control (Benjamini & Hochberg, 1995). In the differential expression reference table, 0 genes satisfied FDR ≤ 0 . 05 , 35 genes fell within 0 . 05 < FDR ≤ 0 . 10 , and 127 genes within 0 . 05 < FDR ≤ 0 . 15 , defining the candidate sets used in the strict, relaxed, and borderline prompt regimes. These thresholds between 0.05 and 0.10 are often treated as borderline discoveries, reflecting weaker statistical support while still capturing potentially biologically relev ant signals. The resulting differential expression table contains gene identifiers, log2 fold changes, test statistics, and FDR-adjusted p -values, and it serves as the sole input provided to all models. No additional biological annotations, pathway databases, metadata beyond group labels, or external knowledge sources were supplied. This controlled setup isolates model behavior from data variability and enables direct comparison against a fixed, statistically grounded reference. H . 3 C O M P U TA T I O N A L E N V I RO N M E N T All experiments were conducted using scripted prompt ex ecution with automated output parsing. Analysis code was implemented in R, and e valuation metrics were computed directly from raw 12 I Can’t Belie ve It’ s Not Better W orkshop @ ICLR 2026 model outputs without post-processing beyond JSON parsing and set extraction. The full compu- tational en vironment, including package dependencies and v ersion information, is specified in a configuration file provided with the accompanying code release. Exact prompts, hyperparameters, and ev aluation scripts are included in the supplementary materials. T ogether, these controls ensure that observed differences reflect model behavior rather than data variability , ev aluation artifacts, or hidden system augmentation. H . 4 C O D E A V A I L A B I L I T Y All prompts, ra w LLM outputs, e valuation scripts, and the DESeq2 reference tables used in this study are publicly av ailable in the project repository at: https://github.com/NaziaRiasat/ llm- prompt- sensitivity . I U S E O F L A R G E L A N G U AG E M O D E L S Large language models were used as experimental subjects in this study and were queried via their public interfaces under controlled prompting conditions, as described in the manuscript. LLMs were also used in a limited capacity to assist with code debugging and language polishing during manuscript preparation. All experimental design decisions, data analysis, result interpretation, and conclusions were made by the author . J E T H I C S S T A T E M E N T This study ev aluates the behavior of large language models in a controlled computational setting using publicly a vailable data. No human subjects, personal data, or sensiti ve information were in volv ed. The findings are intended to inform responsible use of LLMs in scientific workflows by highlighting limitations related to reliability and reproducibility . K R E P R O D U C I B I L I T Y S T A T E M E N T All experiments were conducted using fix ed input data, deterministic reference analyses, and scripted prompt execution. Exact prompts, ev aluation metrics, and analysis code are provided in the supplementary materials. Each prompt was executed repeatedly under identical conditions to assess model stability and prompt sensitivity . 13
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment