R2GenCSR: Mining Contextual and Residual Information for LLMs-based Radiology Report Generation
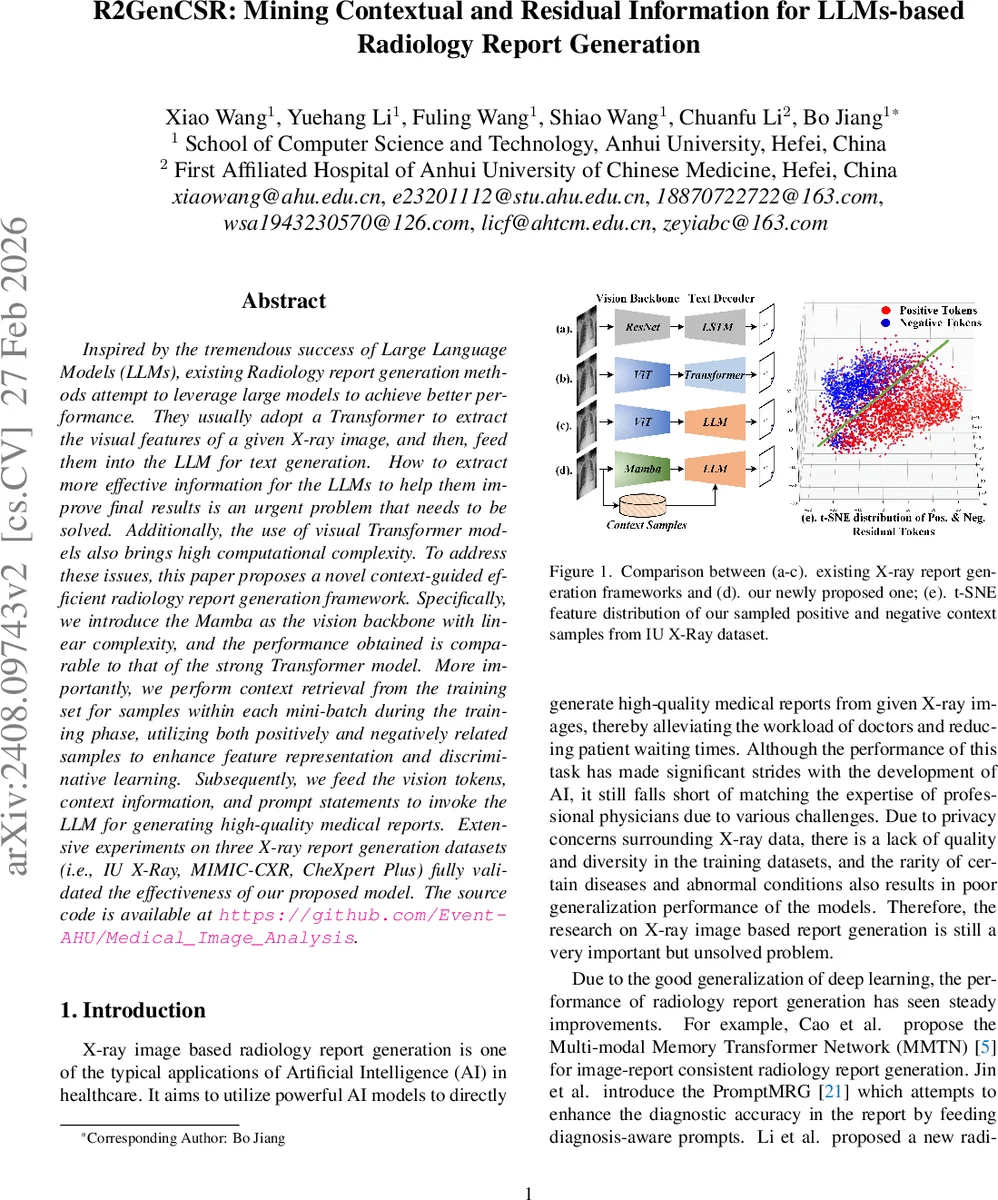
Inspired by the tremendous success of Large Language Models (LLMs), existing Radiology report generation methods attempt to leverage large models to achieve better performance. They usually adopt a Transformer to extract the visual features of a given X-ray image, and then, feed them into the LLM for text generation. How to extract more effective information for the LLMs to help them improve final results is an urgent problem that needs to be solved. Additionally, the use of visual Transformer models also brings high computational complexity. To address these issues, this paper proposes a novel context-guided efficient radiology report generation framework. Specifically, we introduce the Mamba as the vision backbone with linear complexity, and the performance obtained is comparable to that of the strong Transformer model. More importantly, we perform context retrieval from the training set for samples within each mini-batch during the training phase, utilizing both positively and negatively related samples to enhance feature representation and discriminative learning. Subsequently, we feed the vision tokens, context information, and prompt statements to invoke the LLM for generating high-quality medical reports. Extensive experiments on three X-ray report generation datasets (i.e., IU X-Ray, MIMIC-CXR, CheXpert Plus) fully validated the effectiveness of our proposed model. The source code is available at https://github.com/Event-AHU/Medical_Image_Analysis.
💡 Research Summary
The paper introduces R2GenCSR, a novel framework for generating radiology reports from chest X‑ray images that leverages large language models (LLMs) while addressing two major bottlenecks of existing approaches: (1) the high computational cost of transformer‑based visual encoders and (2) the limited contextual information supplied to LLMs.
Vision Backbone – Mamba
Instead of the conventional Vision Transformer (ViT), the authors adopt the Mamba architecture, a state‑space model (SSM) that processes sequences with linear O(N) complexity. An X‑ray image is first split into patches, each treated as a token. Mamba updates a hidden state hₜ recursively, effectively accumulating information from all previous patches and providing an implicit global receptive field without quadratic self‑attention. This design dramatically reduces GPU memory consumption (≈30 % less) and speeds up inference (≈1.8× faster) while preserving or even improving visual representation quality for medical images, where pathological patterns can be spatially dispersed.
Contextual Retrieval and Residual Tokens
During training, for every image in a mini‑batch the method retrieves a set of context samples from the training pool: one or more positive samples (images with similar disease patterns) and one or more negative samples (normal images). Retrieval is performed in the visual token space using cosine similarity or a learned embedding distance. Both the query image and the retrieved samples are passed through the same Mamba backbone, yielding visual token sequences. The global token of the query (analogous to a
Comments & Academic Discussion
Loading comments...
Leave a Comment