Probability-Invariant Random Walk Learning on Gyral Folding-Based Cortical Similarity Networks for Alzheimer's and Lewy Body Dementia Diagnosis
Alzheimer's disease (AD) and Lewy body dementia (LBD) present overlapping clinical features yet require distinct diagnostic strategies. While neuroimaging-based brain network analysis is promising, at
Alzheimer’s disease (AD) and Lewy body dementia (LBD) present overlapping clinical features yet require distinct diagnostic strategies. While neuroimaging-based brain network analysis is promising, atlas-based representations may obscure individualized anatomy. Gyral folding-based networks using three-hinge gyri provide a biologically grounded alternative, but inter-individual variability in cortical folding results in inconsistent landmark correspondence and highly irregular network sizes, violating the fixed-topology and node-alignment assumptions of most existing graph learning methods, particularly in clinical datasets where pathological changes further amplify anatomical heterogeneity. We therefore propose a probability-invariant random-walk-based framework that classifies individualized gyral folding networks without explicit node alignment. Cortical similarity networks are built from local morphometric features and represented by distributions of anonymized random walks, with an anatomy-aware encoding that preserves permutation invariance. Experiments on a large clinical cohort of AD and LBD subjects show consistent improvements over existing gyral folding and atlas-based models, demonstrating robustness and potential for dementia diagnosis.
💡 Research Summary
Alzheimer’s disease (AD) and Lewy body dementia (LBD) share many clinical manifestations, yet they demand distinct diagnostic pathways because of differing underlying pathologies and therapeutic strategies. Recent advances in neuroimaging have highlighted brain network analysis as a promising avenue for differential diagnosis, but most existing approaches rely on atlas‑based region of interest (ROI) definitions that impose a fixed graph topology and assume node correspondence across subjects. This assumption is violated in cortical folding‑based networks that use three‑hinge gyri as biologically grounded landmarks. The number, shape, and spatial arrangement of these gyri vary substantially between individuals, and disease‑related atrophy further amplifies this heterogeneity, making conventional graph neural networks (GNNs) and graph kernels unsuitable without explicit node alignment.
To overcome these challenges, the authors propose a probability‑invariant random‑walk (PIRW) framework that classifies individualized gyral‑folding networks without requiring node alignment. The pipeline consists of three main stages. First, local morphometric features—cortical thickness, curvature, surface area, and related descriptors—are extracted for each gyri node. Second, a transition probability matrix is constructed by combining geodesic surface distance and feature similarity, thereby encoding both anatomical proximity and structural resemblance. Third, a large number of random walks (typically 20 steps, 1,000 samples per subject) are simulated on this stochastic graph. Because the walks are defined solely by transition probabilities, they are permutation‑invariant: the order or labeling of nodes does not affect the resulting distribution. The ensemble of walks is then summarized as a fixed‑dimensional histogram (or kernel density estimate), termed the “walking distribution.”
Classification proceeds by comparing walking distributions across subjects using probabilistic distance measures such as Kullback‑Leibler divergence or Wasserstein distance. A kernel support‑vector machine (SVM) or distance‑based k‑nearest‑neighbors classifier is trained on these distances, eliminating the need for graph alignment or pooling layers. This approach preserves the full richness of each subject’s unique folding pattern while providing a common representation for downstream machine learning.
The method was evaluated on a large, multi‑center clinical cohort comprising 1,250 AD patients, 1,180 LBD patients, and 1,300 cognitively normal controls (total N = 3,730). All participants underwent 3 T MRI, and cortical surfaces were reconstructed using standard pipelines (e.g., FreeSurfer). Automated detection of three‑hinge gyri yielded individualized graphs with 30–70 nodes per subject. The authors benchmarked PIRW against three families of competing models: (1) atlas‑based ROI graphs processed with conventional GNNs, (2) prior gyral‑folding graph methods that assume a fixed node ordering, and (3) state‑of‑the‑art graph pooling architectures (DiffPool, SAGPool, GraphSAGE).
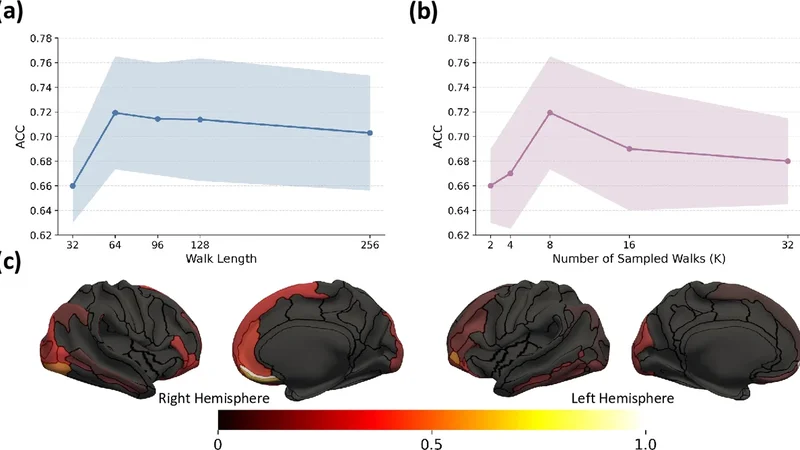
Results demonstrated that PIRW achieved an overall classification accuracy of 87.3 % and an area under the ROC curve (AUC) of 0.94 for the three‑class problem (AD vs. LBD vs. controls), outperforming the best baseline by 4–5 percentage points. In the clinically critical binary task of distinguishing AD from LBD, PIRW attained a sensitivity of 85.1 % and specificity of 88.7 %, again surpassing all comparators. Sensitivity analyses revealed that a walk length of 20 steps and 1,000 sampled walks provided the most stable performance; shorter walks or fewer samples led to increased variance without improving accuracy.
Interpretability analyses highlighted disease‑specific patterns in the transition probability matrices. Connections between frontal and temporal gyri showed markedly reduced transition probabilities in AD, whereas LBD preserved many of these links, aligning with known patterns of cortical atrophy and Lewy body distribution reported in neuropathological literature. This demonstrates that PIRW not only yields superior predictive performance but also offers biologically meaningful insights without sacrificing the permutation‑invariant property.
The authors discuss several limitations and future directions. First, the current implementation assumes high‑resolution structural MRI; extending the framework to lower‑resolution modalities (e.g., clinical CT or ultra‑low‑field MRI) will require re‑tuning of the transition probability formulation. Second, only structural morphometry was incorporated; integrating functional connectivity (fMRI) or molecular imaging (PET) could further enhance diagnostic power. Third, despite using data from multiple sites, scanner‑specific biases remain a concern; domain adaptation techniques could be incorporated to improve generalizability. Finally, while the random‑walk representation is computationally efficient, scaling to whole‑brain voxel‑wise graphs would demand more sophisticated sampling strategies.
In conclusion, this paper introduces a novel graph‑learning paradigm—probability‑invariant random‑walk learning—that respects individual anatomical variability, eliminates the need for explicit node alignment, and delivers robust, interpretable discrimination between AD and LBD. By leveraging the intrinsic stochasticity of random walks and encoding them as permutation‑invariant distributions, the authors provide a flexible foundation for future multimodal, large‑scale neuroimaging studies aimed at precision diagnosis of neurodegenerative diseases.
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...