Modeling Multivariate Missingness with Tree Graphs and Conjugate Odds
In this paper, we analyze a specific class of missing not at random (MNAR) assumptions called tree graphs, extending upon the work of pattern graphs. We build off previous work by introducing the idea of a conjugate odds family in which certain parametric models on the selection odds can preserve the data distribution family across all missing data patterns. Under a conjugate odds family and a tree graph assumption, we are able to model the full data distribution elegantly in the sense that for the observed data, we obtain a model that is conjugate from the complete-data, and for the missing entries, we create a simple imputation model. In addition, we investigate the problem of graph selection, sensitivity analysis, and statistical inference. Using both simulations and real data, we illustrate the applicability of our method.
💡 Research Summary
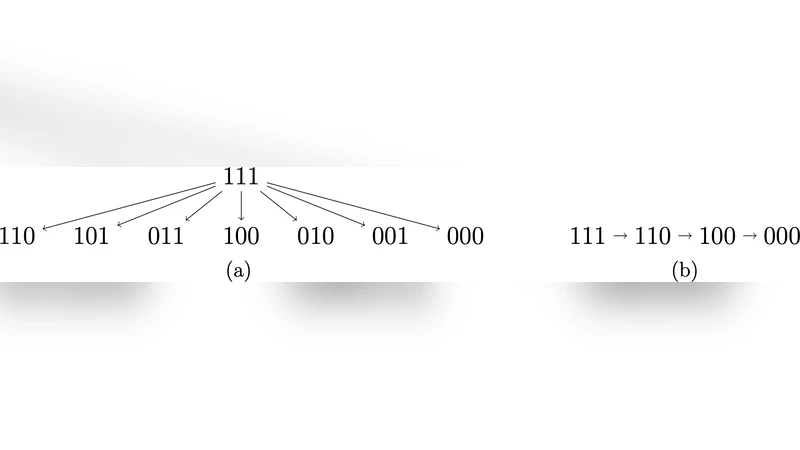
This paper tackles the challenging problem of missing‑not‑at‑random (MNAR) data in multivariate settings by introducing two complementary ideas: tree graphs, a restricted subclass of pattern graphs, and the conjugate odds family, a parametric construction that preserves the form of the full‑data distribution across missingness patterns. A pattern graph encodes relationships among missingness patterns; a tree graph further requires that every pattern be reachable from the fully observed pattern (all‑ones) by a unique directed path, making the graph an arborescence with exactly one parent per node and the minimal number of edges (2^d − 1). The authors prove that any tree graph defines a unique MNAR assumption that non‑parametrically identifies the full data distribution. Moreover, the selection odds for a pattern can be expressed as a product of conditional odds along the unique path, linking each step only to the variables observed at that stage.
The conjugate odds family builds on this structure by specifying the selection odds with a parametric form (e.g., logistic linear, beta‑ratio) that is “conjugate” to the assumed full‑data model. If the complete data follow a multivariate normal distribution, the conjugate odds ensure that the observed‑data posterior remains normal, while the missing‑data conditional distribution retains a simple closed form (e.g., normal conditional). This conjugacy yields computationally convenient Bayesian updates and enables coherent multiple imputation where the imputation model is automatically compatible with the analysis model.
Combining tree graphs with conjugate odds yields a unified probabilistic model: the observed data are modeled by the same family as the full data, and each missingness edge requires only a single parametric odds model. Consequently, the overall model is both identifiable and parsimonious, avoiding the explosion of parameters typical of general MNAR specifications.
The paper discusses three strategies for selecting an appropriate tree graph: (1) expert‑driven specification based on substantive knowledge of causal or survey design; (2) partial‑ordering constraints derived from natural variable orderings (e.g., time, monotone dropout); and (3) data‑driven search using pattern frequencies, information criteria (AIC/BIC), and cross‑validation to rank candidate trees. Sensitivity analysis is facilitated by varying the odds parameters within plausible ranges, defining “odds sensitivity intervals” that quantify how inference changes under departures from the baseline MNAR assumption.
Statistical inference proceeds via Bayesian MCMC exploiting the conjugate structure, providing posterior samples for both parameters and imputed values, as well as standard errors and credible intervals. The authors also extend the conjugate odds concept to non‑Gaussian families (binary, count) by employing appropriate exponential‑family conjugate forms.
Extensive simulations (d = 5–8, missing rates 20–50 %) demonstrate that the proposed method outperforms popular MAR‑based tools such as MissForest and MICE in terms of bias, root‑mean‑square error, and coverage. An application to an Alzheimer’s disease dataset (cognitive tests, biomarkers, clinical covariates) shows improved predictive performance and more accurate recovery of inter‑variable relationships compared with standard imputation techniques.
In summary, the work offers a theoretically grounded, practically implementable framework for MNAR data: tree graphs provide a tractable yet expressive representation of missingness mechanisms, while conjugate odds guarantee that the full‑data distribution family is preserved across patterns, enabling efficient estimation, coherent imputation, and robust sensitivity analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment