Network geometry of the Drosophila brain
The recent reconstruction of the Drosophila brain provides a neural network of unprecedented size and level of details. In this work, we study the geometrical properties of this system by applying network embedding techniques to the graph of synaptic connections. Since previous analysis have revealed an inhomogeneous degree distribution, we first employ a hyperbolic embedding approach that maps the neural network onto a point cloud in the two-dimensional hyperbolic space. In general, hyperbolic embedding methods exploit the exponentially growing volume of hyperbolic space with increasing distance from the origin, allowing for an approximately uniform spatial distribution of nodes even in scale-free, small-world networks. By evaluating multiple embedding quality metrics, we find that the network structure is well captured by the resulting two-dimensional hyperbolic embedding, and in fact is more congruent with this representation than with the original neuron coordinates in three-dimensional Euclidean space. In order to examine the network geometry in a broader context, we also apply the well-known Euclidean network embedding approach Node2vec, where the dimension of the embedding space, $d$ can be set arbitrarily. In 3 dimensions, the Euclidean embedding of the network yields lower quality scores compared to the original neuron coordinates. However, as a function of the embedding dimension the scores show an improving tendency, surpassing the level of the 2d hyperbolic embedding roughly at $d=16$, and reaching a maximum around $d=64$. Since network embeddings can serve as valuable inputs for a variety of downstream machine learning tasks, our results offer new perspectives on the structure and representation of this recently revealed and biologically significant neural network.
💡 Research Summary
The adult female Drosophila melanogaster brain connectome, comprising 139,255 neurons and roughly 50 million chemical synapses, represents one of the largest publicly available neural graphs. Previous work has shown that this network exhibits a scale‑free degree distribution, a pronounced rich‑club core, and distinct functional super‑classes (e.g., “central”, “optic”). In the present study the authors investigate how well the intrinsic topology of this connectome can be captured by two fundamentally different metric‑space embeddings: a two‑dimensional hyperbolic embedding generated with the Cluster‑Level Optimised Vertex Embedding (CLOVE) algorithm, and a series of Euclidean embeddings produced by Node2vec with dimensionalities ranging from 2 to 512.
Hyperbolic space is attractive for scale‑free, small‑world graphs because its exponential volume growth allows high‑degree hubs to be placed near the origin while low‑degree nodes occupy the periphery, preserving hierarchical relationships in only two dimensions. CLOVE, a recent method designed for massive graphs, efficiently computes a 2‑D hyperbolic map (the native Poincaré disk) of the fly brain. In contrast, Node2vec learns low‑dimensional Euclidean coordinates by performing biased random walks; the authors vary the embedding dimension (d) to explore the trade‑off between representational power and computational cost.
To assess embedding quality the authors compute eight established metrics: Mapping Accuracy (MA, Spearman correlation between graph shortest‑path lengths and Euclidean/hyperbolic distances), Greedy Routing Success Rate (GR), Greedy Routing Score (GRS), Greedy Routing Efficiency (GRE), Edge Prediction Area Under the ROC Curve (EP AUC), Edge Prediction Precision (EPP), and the recall of true edges among the top 20 % and 5 % of closest node pairs (EPR20, EPR5). Because exact evaluation on a graph of this size is infeasible, they estimate each metric by random sampling of 0.1 % of node pairs.
The results (Table 1) show that the 2‑D hyperbolic embedding outperforms both the original 3‑D anatomical coordinates and the low‑dimensional Euclidean embeddings (d = 2, 3) across all metrics. Notably, GR, GRS and GRE are dramatically higher for the hyperbolic map (GR = 0.553, GRS = 0.390, GRE = 0.160) than for the physical layout (GR ≈ 0.075, GRS ≈ 0.048, GRE ≈ 0.050). This indicates that greedy navigation—an idealised model of information flow—operates far more efficiently when the network is represented in hyperbolic space.
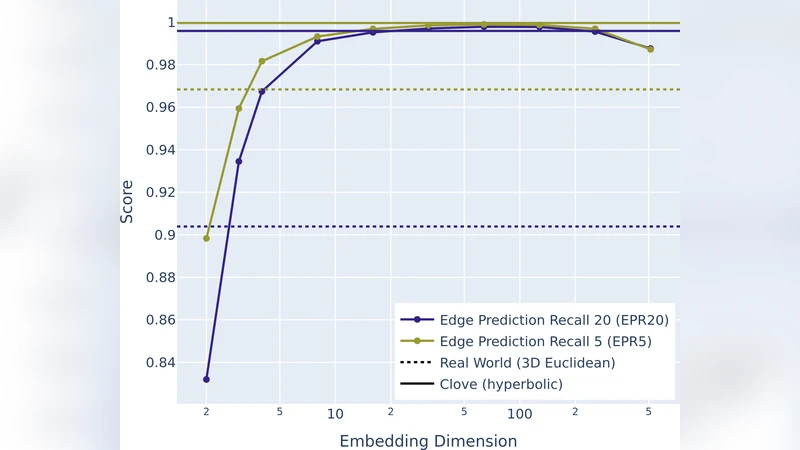
Increasing the Euclidean dimensionality improves performance substantially. At d = 4, MA reaches 0.500 and GR 0.142; at d = 16, MA = 0.653 and GR = 0.629; at d = 64 the edge‑prediction scores peak (EP AUC = 0.996, EPP = 0.995) and the recall metrics approach 1.0. Thus, around d ≈ 16 the Euclidean embedding surpasses the 2‑D hyperbolic representation, and the optimum lies near d = 64. Beyond this point (d ≥ 128) the metrics decline, reflecting the curse of dimensionality and the increasing noise in distance estimates.
Visualization (Figure 1) further supports these quantitative findings. The original 3‑D coordinates, when projected onto a 2‑D plane, retain the overall brain silhouette but suffer from edge clutter. The hyperbolic embedding spreads nodes across the disk, placing high‑degree hubs near the centre and preserving the angular separation of functional super‑classes. The 64‑D Euclidean embedding, reduced to 2‑D with UMAP, also shows clear segregation of the same super‑classes, albeit with some distortion due to dimensionality reduction.
The authors conclude that (1) hyperbolic embeddings provide a compact, low‑dimensional representation that captures both global hierarchy and local community structure, excelling especially in tasks that rely on geometric navigation; (2) high‑dimensional Euclidean embeddings can achieve comparable or superior performance for link‑prediction and similarity‑based tasks, but require careful selection of dimensionality to avoid degradation; (3) embedding quality metrics are valuable diagnostics for downstream machine‑learning pipelines, such as missing‑link prediction, node classification, or routing simulations in neural tissue. Consequently, the choice between hyperbolic and Euclidean embeddings should be guided by the specific analytical goal, computational budget, and the desired balance between interpretability (low‑dimensional hyperbolic maps) and predictive power (moderate‑dimensional Euclidean spaces).
Comments & Academic Discussion
Loading comments...
Leave a Comment