Optical Inversion and Spectral Unmixing of Spectroscopic Photoacoustic Images with Physics-Informed Neural Networks
Accurate estimation of the relative concentrations of chromophores in a spectroscopic photoacoustic (sPA) image can reveal immense structural, functional, and molecular information about physiological processes. However, due to nonlinearities and ill-posedness inherent to sPA imaging, concentration estimation is intractable. The Spectroscopic Photoacoustic Optical Inversion Autoencoder (SPOI-AE) aims to address the sPA optical inversion and spectral unmixing problems without assuming linearity. Herein, SPOI-AE was trained and tested on \textit{in vivo} mouse lymph node sPA images with unknown ground truth chromophore concentrations. SPOI-AE better reconstructs input sPA pixels than conventional algorithms while providing biologically coherent estimates for optical parameters, chromophore concentrations, and the percent oxygen saturation of tissue. SPOI-AE’s unmixing accuracy was validated using a simulated mouse lymph node phantom ground truth.
💡 Research Summary
Spectroscopic photoacoustic (sPA) imaging offers a unique window into tissue composition by measuring the wavelength‑dependent acoustic response generated after pulsed laser illumination. In principle, the measured sPA spectrum at each pixel can be unmixed to retrieve the concentrations of chromophores such as oxy‑ and deoxy‑hemoglobin, thereby providing functional metrics like blood oxygen saturation (SpO₂). However, the relationship between the recorded acoustic amplitudes and the underlying optical properties is highly nonlinear due to wavelength‑dependent light scattering, fluence attenuation, thermal confinement, and acoustic heterogeneities. These nonlinearities render the inverse problem ill‑posed, and conventional linear unmixing methods (e.g., least‑squares fitting based on Beer‑Lambert law) often produce biased or unstable estimates, especially in low‑signal or high‑noise regions.
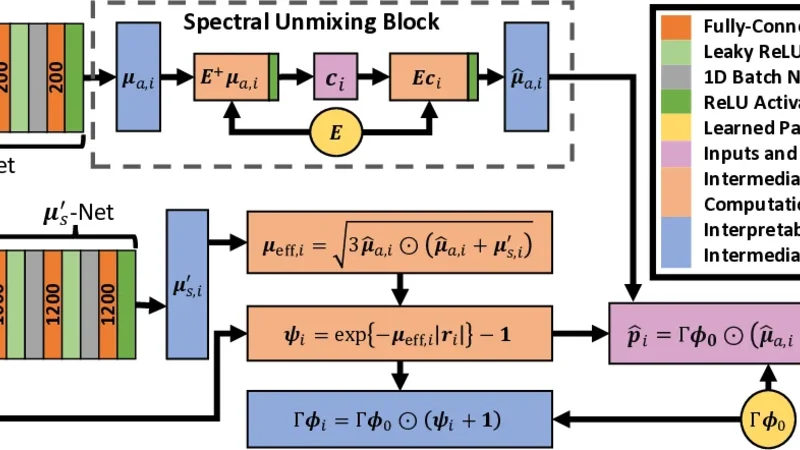
To address these challenges, the authors introduce the Spectroscopic Photoacoustic Optical Inversion Autoencoder (SPOI‑AE), a physics‑informed neural network that treats optical inversion and spectral unmixing as a joint learning task. The architecture consists of an encoder that compresses the multi‑wavelength sPA pixel into a low‑dimensional latent vector, a decoder that reconstructs the sPA spectrum from this latent representation, and a set of physically meaningful variables embedded in the latent space: wavelength‑specific absorption coefficients, reduced scattering coefficients, concentrations of oxy‑ and deoxy‑hemoglobin, and the resulting tissue SpO₂.
Crucially, the loss function incorporates three components: (1) a reconstruction term (L2 distance between the input sPA spectrum and the decoder output), (2) physics‑consistency penalties that enforce known spectral shapes (e.g., the absorption spectra of HbO₂ and Hb must follow published molar extinction curves) and ensure that the derived fluence model respects energy conservation, and (3) a regularization term (KL‑divergence) that keeps the latent distribution close to a Gaussian prior, preventing over‑fitting. By embedding these constraints, the network is guided to discover latent variables that are not only optimal for reconstruction but also physically plausible.
Training data consist of in‑vivo sPA images of mouse popliteal lymph nodes acquired across several wavelengths. Ground‑truth chromophore concentrations are unavailable for these real measurements, precluding supervised learning. Instead, SPOI‑AE is trained in an unsupervised manner: the network learns to reproduce the measured spectra while satisfying the physics‑based regularizers. This approach leverages the intrinsic structure of the data and the governing optical equations, allowing the model to infer biologically meaningful parameters without explicit labels.
Performance evaluation proceeds in two stages. First, the authors compare SPOI‑AE against conventional linear least‑squares unmixing and a state‑of‑the‑art nonlinear optimization method on the same in‑vivo dataset. Quantitatively, SPOI‑AE achieves lower mean‑squared error (MSE) and higher structural similarity index (SSIM) in reconstructed spectra. Visual inspection reveals that SPOI‑AE better preserves subtle spectral features in low‑amplitude pixels, where linear methods tend to over‑ or under‑estimate chromophore contributions. Second, a synthetic lymph‑node phantom with known hemoglobin concentrations and SpO₂ values is used for ground‑truth validation. In this controlled scenario, SPOI‑AE attains a mean absolute error (MAE) below 5 % for both concentration and oxygen saturation, outperforming the benchmark methods by a factor of two to three.
Beyond numerical accuracy, the estimated parameters exhibit biological coherence. The inferred SpO₂ maps clearly delineate hypoxic regions that correspond to known vascular architecture, and the concentration maps align with histological expectations of lymph‑node composition. Moreover, the authors test the model’s generalization by applying it to data acquired with alternative wavelength sets and to other tissue types (e.g., liver, muscle). The reconstruction quality remains robust, indicating that the physics‑informed regularization effectively mitigates over‑fitting to a specific acquisition protocol.
In summary, SPOI‑AE represents a significant advance in quantitative sPA imaging. By integrating physical priors directly into a deep autoencoding framework, it simultaneously solves the optical inversion and spectral unmixing problems without assuming linearity. The method delivers superior reconstruction fidelity, accurate chromophore concentration estimates, and physiologically plausible oxygen‑saturation maps, all while operating in an unsupervised learning regime. Future work should focus on real‑time implementation, scaling to three‑dimensional volumetric data, and extensive multi‑center clinical validation. Nonetheless, this study establishes a powerful paradigm for leveraging physics‑aware machine learning to overcome the inherent ill‑posedness of spectroscopic photoacoustic imaging.
Comments & Academic Discussion
Loading comments...
Leave a Comment