Piecewise integrability of the discrete Hasimoto map for analytic prediction and design of helical peptides
The representation of protein backbone geometry through the discrete nonlinear Schrödinger equation provides a theoretical connection between biological structure and integrable systems. Although the global application of this framework is constrained by chiral degeneracies and non-local interactions we propose that helical peptides can be effectively modeled as piecewise integrable systems in which the discrete Hasimoto map remains applicable within specific geometric boundaries. We delineate these boundaries through an analytic characterization of the mapping between biochemical dihedral angles and Frenet frame parameters for a dataset of 50 helical peptide chains. We demonstrate that the transformation is information-preserving globally but ill-conditioned within the helical basin characterized by a median Jacobian condition number of 31 which suggests that the loss of chiral information arises primarily from local coordinate compression rather than topological singularities. We define a local integrability error $E[n]$ derived from the discrete dispersion relation to show that deviations from integrability are driven predominantly by torsion non-uniformity while curvature remains structurally rigid. This metric identifies integrable islands where the analytic dispersion relation predicts backbone coordinates with sub-angstrom accuracy yielding a median root-mean-square deviation of 0.77,Å and enables a segmentation strategy that isolates structural defects. We further indicate that the inverse design of peptide backbones is feasible within a quantitatively defined integrability zone where the design constraint reduces essentially to the control of torsion uniformity. These findings advance the Hasimoto formalism from a qualitative descriptor toward a precise quantitative framework for analyzing and designing local protein geometry within the limits of piecewise integrability.
💡 Research Summary
The paper investigates how the discrete Hasimoto map—an isomorphism that translates the geometric description of a space curve into the language of the discrete nonlinear Schrödinger equation (DNLS)—can be employed to predict and design the backbone geometry of helical peptides. While previous work has shown that the global application of this map to whole proteins is hampered by chiral degeneracies (left‑handed and right‑handed helices map to the same complex wavefunction) and by long‑range, non‑local interactions, the authors propose that a piecewise‑integrable approach can circumvent these limitations for locally regular structures such as helices.
Data set and mapping
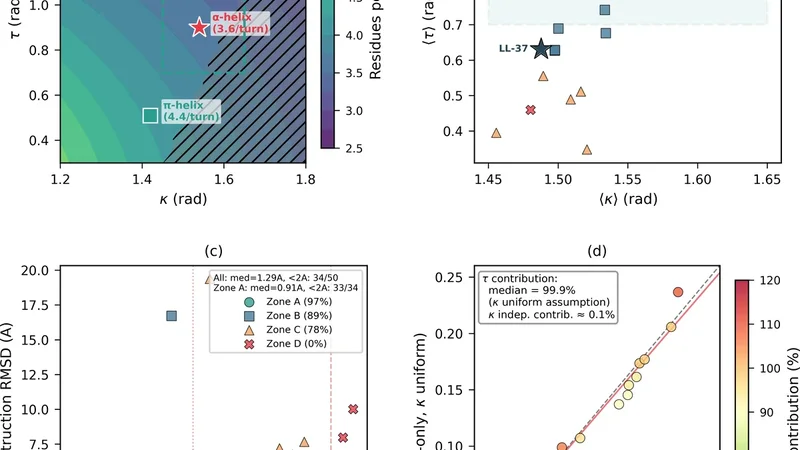
Fifty short helical peptide chains (including α‑helices, 3₁₀‑helices, and π‑helices) were extracted from the Protein Data Bank. For each residue the backbone dihedral angles φ and ψ were measured and then converted into discrete Frenet frame parameters—curvature κₙ and torsion τₙ—using the relations
\
Comments & Academic Discussion
Loading comments...
Leave a Comment