Oscillation Criteria in Large-Scale Gene Regulatory Networks with Intrinsic Fluctuations
Gene Regulatory Networks(GRNs) with feedback are essential components of many cellular processes and may exhibit oscillatory behavior. Analyzing such systems becomes increasingly complex as the number of components increases. Since gene regulation often involves a small number of molecules, fluctuations are inevitable. Therefore, it is important to understand how fluctuations affect the oscillatory dynamics of cellular processes, as this will allow comprehension of the mechanisms that enable cellular functions to remain even in the presence of fluctuations or, failing that, to determine the limit of fluctuations that permits various cellular functions. In this study, we investigated the conditions under which GRNs with feedback and intrinsic fluctuations exhibit oscillatory behavior. Our focus was on developing a procedure that would be both manageable and practical, even for extensive regulatory networks, that is, those comprising numerous nodes. Using the second-moment approach, we described the stochastic dynamics through a set of ordinary differential equations for the mean concentration and its second central moment. The system can attain either a stable equilibrium or oscillatory behavior, depending on its scale and, consequently, the intensity of fluctuations. To illustrate the procedure, we analyzed two relevant systems: a repressilator with three nodes and a system with five nodes, both incorporating intrinsic fluctuations. In both cases, it was observed that for very small systems, which therefore exhibit significant fluctuations, oscillatory behavior is inhibited. The procedure presented here for analyzing the stability of oscillations under fluctuations enables the determination of the critical minimum size of GRNs at which intrinsic fluctuations do not eliminate their cyclical behavior.
💡 Research Summary
This paper addresses a fundamental question in systems biology: under what conditions can gene regulatory networks (GRNs) that contain feedback loops sustain oscillatory dynamics when intrinsic molecular noise is present? Because many cellular processes rely on a relatively small number of transcription factors and mRNA molecules, stochastic fluctuations are unavoidable and can dramatically reshape the behavior predicted by deterministic models. The authors therefore set out to develop a tractable, scalable analytical framework that can be applied to large‑scale networks with many nodes, while still capturing the essential effects of intrinsic noise.
The methodological core is the second‑moment (or mean‑variance) approach. Starting from the stochastic chemical master equation that governs the probability distribution of molecular copy numbers, the authors derive ordinary differential equations (ODEs) for the first moment (the mean concentration vector μ) and the second central moment (the covariance matrix C). The resulting closed system reads
dμ/dt = f(μ) + ½ ∑ ∂²f/∂μ_j∂μ_k C_{jk},
dC/dt = J C + C Jᵀ + Q,
where f(μ) encodes the deterministic reaction rates, J is the Jacobian of f evaluated at μ, and Q is the diffusion matrix that captures the noise generated by each elementary reaction. This formulation preserves the coupling between mean dynamics and fluctuations, allowing the authors to study how noise feeds back into the average behavior of the network.
Stability analysis proceeds by locating the steady state (μ*, C*) of the ODE system and linearizing around it. The eigenvalues of the combined Jacobian determine whether the fixed point is a stable equilibrium or whether a Hopf bifurcation occurs, giving rise to a limit‑cycle oscillation. Crucially, the eigenvalues depend on the network size N (the number of genes) because N controls the average molecule number per species and therefore the magnitude of the covariance terms. The authors derive explicit criteria that relate N, the repression strength k, the degradation rate γ, and the intrinsic noise intensity D to the sign of the real part of the leading eigenvalue. In simple terms, larger networks dilute stochastic fluctuations (the coefficient of variation scales roughly as 1/√N), making it easier for the system to cross the Hopf threshold.
To illustrate the theory, two benchmark systems are examined. The first is the classic three‑node repressilator, in which each gene represses the next in a cyclic fashion. Using realistic kinetic parameters, the mean copy number per gene in this configuration stays in the tens, leading to a coefficient of variation above 30 %. The eigenvalue analysis shows that all real parts remain negative; consequently, the Hopf condition is never satisfied and the stochastic repressilator fails to oscillate. Direct Gillespie simulations confirm the absence of sustained cycles, displaying only noisy fluctuations around a steady state.
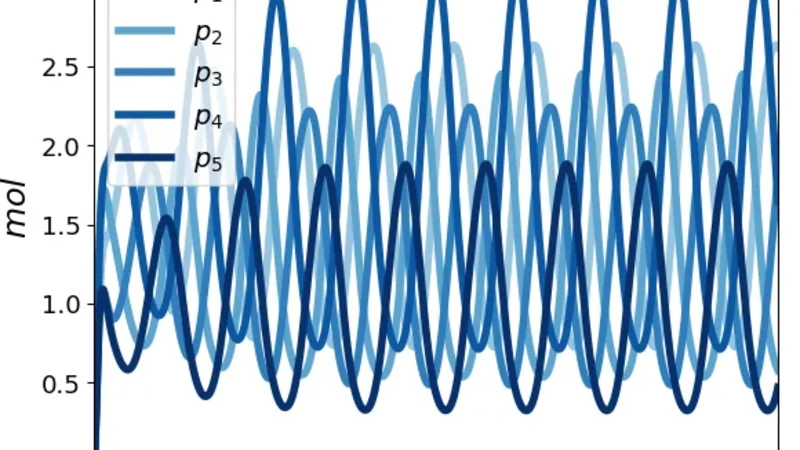
The second example expands the circuit to five nodes by adding two extra negative feedback links, thereby increasing the total number of molecular species. In this larger network the average copy number rises to 80–120 molecules per gene, reducing the coefficient of variation to below 15 %. For a range of repression strengths (k ≈ 1–2 µM⁻¹) and degradation rates (γ ≈ 0.1 min⁻¹), the linearized system exhibits a pair of complex conjugate eigenvalues whose real part crosses zero, indicating a Hopf bifurcation. Numerical integration of the mean‑variance ODEs yields a stable limit cycle with a period of roughly 0.8 h, and stochastic Gillespie runs reproduce the same rhythmic behavior, albeit with amplitude fluctuations.
A particularly useful outcome of the analysis is the concept of a “critical minimum size” N_c, defined as the smallest number of nodes for which intrinsic noise no longer suppresses oscillations. By approximating the Hopf condition, the authors obtain a simple scaling law:
N_c ≈ C · (D/k²),
where C is a constant that depends on the specific network topology and kinetic parameters, D quantifies the intrinsic noise strength, and k is the repression coefficient. This relationship provides a practical design rule: to engineer a synthetic oscillator that remains robust in the presence of molecular noise, one must either increase the number of interacting genes, strengthen the repression interactions, or reduce the intrinsic noise (for example, by using high‑copy plasmids or stabilizing mRNA).
Overall, the paper makes three major contributions. First, it delivers a mathematically rigorous yet computationally inexpensive framework for assessing stochastic stability in high‑dimensional GRNs, bridging the gap between deterministic bifurcation theory and stochastic simulation. Second, it quantifies how network scale modulates the impact of intrinsic fluctuations, confirming the intuitive notion that small circuits are more vulnerable to noise‑induced damping of oscillations. Third, it translates these insights into actionable guidelines for synthetic biology, enabling researchers to predict the minimal circuit complexity required for reliable rhythmic behavior. The authors suggest future extensions that incorporate extrinsic noise sources, time delays, and multi‑scale coupling, as well as experimental validation in living cells, which would further cement the utility of the second‑moment approach for real‑world biological design.
Comments & Academic Discussion
Loading comments...
Leave a Comment