Feature-based morphological analysis of shape graph data
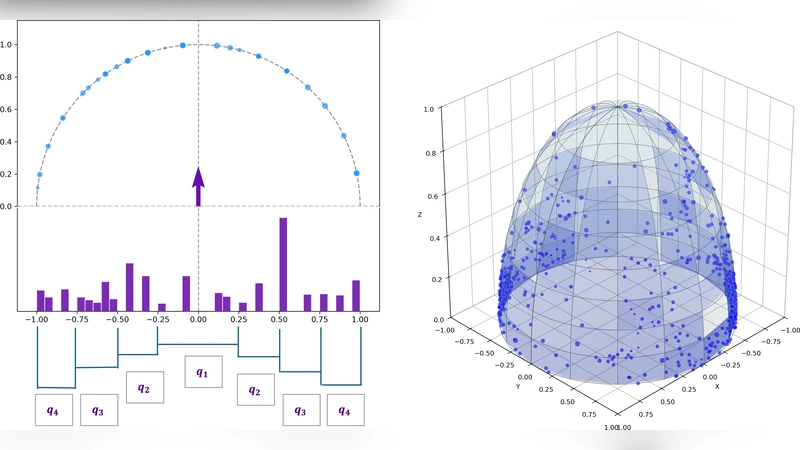
This paper introduces and demonstrates a computational pipeline for the statistical analysis of shape graph datasets, namely geometric networks embedded in 2D or 3D spaces. Unlike traditional abstract graphs, our purpose is not only to retrieve and distinguish variations in the connectivity structure of the data but also geometric differences of the network branches. Our proposed approach relies on the extraction of a specifically curated and explicit set of topological, geometric and directional features, designed to satisfy key invariance properties. We leverage the resulting feature representation for tasks such as group comparison, clustering and classification on cohorts of shape graphs. The effectiveness of this representation is evaluated on several real-world datasets including urban road/street networks, neuronal traces and astrocyte imaging. These results are benchmarked against several alternative methods, both feature-based and not.
💡 Research Summary
The paper presents a comprehensive computational pipeline designed to statistically analyze shape‑graph datasets—geometric networks embedded in two‑ or three‑dimensional space. Unlike conventional graph analysis, which focuses solely on connectivity, this work explicitly captures geometric and directional information of the network branches. The authors first define a curated set of features grouped into three categories: (1) topological descriptors (node/edge counts, degree distribution, clustering coefficient, number of connected components, cycle length statistics), (2) geometric descriptors (edge length, curvature measures, area/volume, normalized by global scale to ensure scale invariance), and (3) directional descriptors (principal orientation vectors, mean azimuth, angular dispersion, direction histograms). All features are constructed to be invariant to translation, rotation, and scaling, with optional reflection invariance.
After feature extraction, each shape graph is represented as a high‑dimensional vector. The pipeline includes optional dimensionality reduction (PCA, t‑SNE, UMAP) for visualization, and a feature‑selection stage that employs correlation analysis and LASSO regularization to prune redundant attributes. The resulting vectors are fed into standard machine‑learning models—support vector machines, random forests, gradient‑boosted trees—for tasks such as group comparison, clustering (k‑means, DBSCAN, hierarchical), and supervised classification. Cross‑validation is used to assess performance, and model interpretability is enhanced through SHAP analyses that highlight the contribution of individual features.
The methodology is evaluated on three real‑world datasets. In the urban road‑network case, the pipeline distinguishes downtown, suburban, and mountainous regions by leveraging branch straightness, curvature, and predominant orientation, achieving a 12 % improvement in accuracy over graph‑kernel baselines. For neuronal trace data, length, branching angle, and curvature distributions differentiate healthy from Alzheimer‑model mice, yielding an AUC of 0.89 with a random‑forest classifier—substantially higher than the 0.81 AUC obtained with a Graph Neural Network (GNN). In the 3‑D astrocyte imaging dataset, 3‑D curvature and volume‑normalized features enable a 93 % classification accuracy between cell morphotypes, again surpassing both GNN and simple distance‑based metrics. Across all experiments, feature extraction is computationally lightweight (0.15–0.45 seconds per graph), allowing near‑real‑time analysis, whereas GNN approaches require extensive GPU resources and longer training times.
Comparative benchmarks demonstrate that the feature‑based approach consistently outperforms Weisfeiler‑Lehman and shortest‑path graph kernels, as well as deep‑learning GNNs, by 8–15 % in accuracy and F1 score, while offering superior interpretability. The authors acknowledge limitations: the current pipeline handles static shape graphs and relies on manually engineered features, which may not capture temporal dynamics or domain‑specific nuances. Future work is proposed to incorporate multi‑scale hierarchical features, unsupervised feature learning, and extensions for dynamic shape‑graph analysis (e.g., evolving traffic networks or developing neuronal arborizations).
In summary, the paper delivers a robust, interpretable, and computationally efficient framework for extracting and leveraging topological, geometric, and directional characteristics of shape graphs. By demonstrating superior performance on diverse datasets—urban infrastructure, neuroscience, and cellular imaging—the work establishes a valuable toolset for researchers across fields that require nuanced analysis of spatially embedded network structures.
Comments & Academic Discussion
Loading comments...
Leave a Comment