Tracing protein and proteome history with chronologies and networks: folding recapitulates evolution
Introduction: While the origin and evolution of proteins remain mysterious, advances in evolutionary genomics and systems biology are facilitating the historical exploration of the structure, function and organization of proteins and proteomes. Molecular chronologies are series of time events describing the history of biological systems and subsystems and the rise of biological innovations. Together with time-varying networks, these chronologies provide a window into the past. Areas covered: Here, we review molecular chronologies and networks built with modern methods of phylogeny reconstruction. We discuss how chronologies of structural domain families uncover the explosive emergence of metabolism, the late rise of translation, the co-evolution of ribosomal proteins and rRNA, and the late development of the ribosomal exit tunnel; events that coincided with a tendency to shorten folding time. Evolving networks described the early emergence of domains and a late big bang of domain combinations. Expert opinion: Two processes, folding and recruitment appear central to the evolutionary progression. The former increases protein persistence. The later fosters diversity. Chronologically, protein evolution mirrors folding by combining supersecondary structures into domains, developing translation machinery to facilitate folding speed and stability, and enhancing structural complexity by establishing long-distance interactions in novel structural and architectural designs.
💡 Research Summary
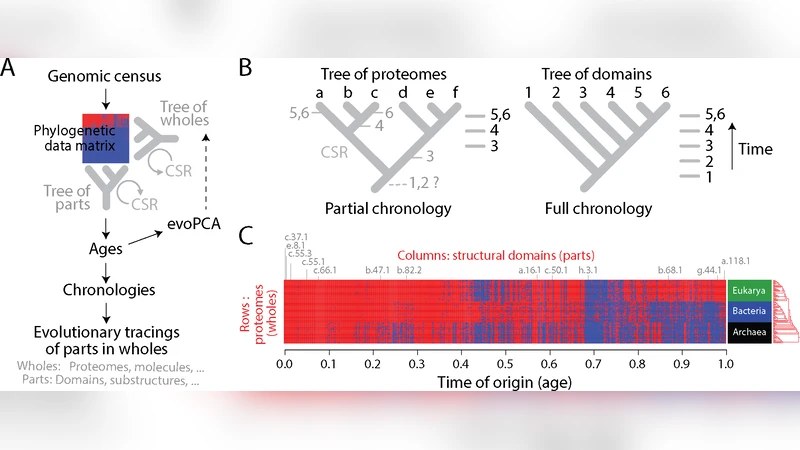
This review synthesizes recent advances in evolutionary genomics and systems biology that enable a historical reconstruction of protein and proteome evolution using molecular chronologies and time‑varying networks. The authors first describe how chronologies are built: structural domain families from databases such as SCOP and CATH are placed on calibrated phylogenetic trees using Bayesian inference, relaxed molecular clocks, and fossil constraints. This yields precise age estimates for each domain lineage.
The resulting timeline reveals four major evolutionary milestones. (1) An “explosive emergence” of metabolic domains occurs around 3.5–3.2 billion years ago (Ga), coinciding with the appearance of complex catalytic chemistries in the early Earth environment. These domains (e.g., Rossmann folds, TIM barrels) provide multifunctional enzymatic capabilities that likely powered the first proto‑metabolic networks. (2) The translation apparatus, including ribosomal proteins and ribosomal RNA, co‑evolves later, roughly 2.5 Ga, supporting the hypothesis that the ribosome and the genetic code matured after metabolism was already established. (3) A coordinated evolution of ribosomal proteins and rRNA is evident, with the ribosomal exit tunnel forming around 2.0 Ga. The tunnel’s emergence is interpreted as an adaptation that allows nascent polypeptides to begin folding while still attached to the ribosome, thereby shortening folding times and increasing protein stability. (4) A “big‑bang” of domain combinations appears after ~2 Ga, where previously isolated single‑domain proteins begin to fuse, generating multi‑domain architectures that underlie modern enzymes, signaling complexes, and structural scaffolds.
Network analyses complement the chronologies. Early in the timeline the domain interaction network is sparse, dominated by a few highly connected “core” domains. As time progresses, the network density rises sharply, reflecting the rapid recruitment of new domains onto existing scaffolds. Centrality metrics show that early core domains retain high betweenness, acting as hubs for later innovations. This pattern supports the authors’ concept of “domain recruitment” – a process that dramatically expands functional diversity by attaching novel modules to pre‑existing structures.
A central theme of the paper is the role of protein folding as an evolutionary driver. By comparing experimentally measured folding rates with the inferred ages of domain families, the authors demonstrate that early single‑domain proteins generally fold slowly and are less stable, whereas later multi‑domain proteins exhibit optimized folding pathways that achieve rapid attainment of the native state. Faster folding reduces the risk of aggregation, lengthens protein half‑life, and thus enhances evolutionary persistence. Consequently, folding speed becomes a selective pressure that shapes both sequence evolution and higher‑order architecture.
The authors propose a two‑process model for protein evolution. The first process, folding, establishes a stable structural foundation and selects for sequences that can achieve a functional conformation quickly. The second process, recruitment, builds on this foundation by fusing domains, creating long‑range interactions, and generating novel structural designs. Chronologically, the sequence is: (i) emergence of simple supersecondary structures that coalesce into domains, (ii) development of the translation machinery that accelerates folding and improves accuracy, and (iii) expansion of structural complexity through domain recruitment and network integration.
In conclusion, the paper argues that protein evolution mirrors the physical act of folding: just as folding combines elementary structural elements into a stable whole, evolution combines elementary domains into increasingly complex proteins, leverages the translation apparatus to speed up folding, and finally diversifies through domain recruitment. This integrated view reconciles phylogenetic, structural, and network data, offering a unified framework that moves beyond the traditional gradual‑mutation paradigm toward a model where folding dynamics and modular recruitment jointly drive the emergence of modern proteomes.
Comments & Academic Discussion
Loading comments...
Leave a Comment