Omni-iEEG: A Large-Scale, Comprehensive iEEG Dataset and Benchmark for Epilepsy Research
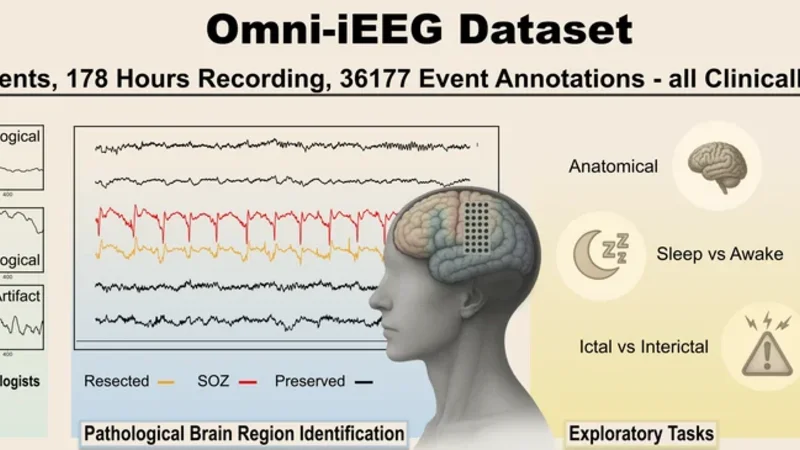
Epilepsy affects over 50 million people worldwide, and one-third of patients suffer drug-resistant seizures where surgery offers the best chance of seizure freedom. Accurate localization of the epileptogenic zone (EZ) relies on intracranial EEG (iEEG). Clinical workflows, however, remain constrained by labor-intensive manual review. At the same time, existing data-driven approaches are typically developed on single-center datasets that are inconsistent in format and metadata, lack standardized benchmarks, and rarely release pathological event annotations, creating barriers to reproducibility, cross-center validation, and clinical relevance. With extensive efforts to reconcile heterogeneous iEEG formats, metadata, and recordings across publicly available sources, we present $\textbf{Omni-iEEG}$, a large-scale, pre-surgical iEEG resource comprising $\textbf{302 patients}$ and $\textbf{178 hours}$ of high-resolution recordings. The dataset includes harmonized clinical metadata such as seizure onset zones, resections, and surgical outcomes, all validated by board-certified epileptologists. In addition, Omni-iEEG provides over 36K expert-validated annotations of pathological events, enabling robust biomarker studies. Omni-iEEG serves as a bridge between machine learning and epilepsy research. It defines clinically meaningful tasks with unified evaluation metrics grounded in clinical priors, enabling systematic evaluation of models in clinically relevant settings. Beyond benchmarking, we demonstrate the potential of end-to-end modeling on long iEEG segments and highlight the transferability of representations pretrained on non-neurophysiological domains. Together, these contributions establish Omni-iEEG as a foundation for reproducible, generalizable, and clinically translatable epilepsy research. The project page with dataset and code links is available at omni-ieeg.github.io/omni-ieeg.
💡 Research Summary
Omni‑iEEG introduces a large‑scale, harmonized intracranial EEG (iEEG) resource that directly addresses the reproducibility, standardization, and clinical relevance challenges that have hampered data‑driven epilepsy research. By aggregating publicly available recordings from multiple centers, the authors have assembled data from 302 presurgical patients, amounting to 178 hours of high‑resolution (generally ≥1 kHz) iEEG. The dataset is accompanied by a richly curated set of clinical metadata—seizure onset zones (SOZ), resection maps, and postoperative outcomes (Engel grades)—all validated by board‑certified epileptologists. In addition, more than 36 000 pathological events (spikes, high‑frequency oscillations, rhythmic discharges, etc.) have been expert‑annotated with precise temporal boundaries and electrode‑specific locations, providing a rare depth of labeling for biomarker discovery.
To transform this resource into a usable benchmark, the authors define four clinically meaningful tasks: (1) SOZ prediction from raw iEEG, (2) detection of pathological events, (3) outcome prediction (Engel classification) using combined iEEG and metadata, and (4) electrode selection optimization for maximal diagnostic yield with minimal invasiveness. Each task is paired with evaluation metrics that reflect clinical priorities—sensitivity, specificity, F1‑score, AUROC—rather than simplistic accuracy, thereby ensuring that algorithmic improvements translate into potential therapeutic benefit.
Technical experiments demonstrate the feasibility of end‑to‑end modeling on long continuous recordings using transformer‑based architectures, and they reveal that representations pretrained on non‑neurophysiological domains (speech, video) can be successfully transferred to iEEG, yielding measurable gains in both SOZ localization and event detection. This finding suggests that large‑scale, domain‑agnostic sequence models may serve as a universal foundation for neurophysiological signal analysis.
The paper also discusses limitations. Heterogeneity in recording hardware and electrode implantation strategies across sites prevents perfect spatial standardization; some patients have relatively short recordings, limiting long‑term trend analyses; and expert annotation, while high‑quality, remains subject to inter‑rater variability, calling for systematic reliability assessments. Moreover, the current cohort is adult‑centric, leaving pediatric generalization an open question.
In summary, Omni‑iEEG constitutes the first comprehensive, publicly released iEEG dataset that couples extensive raw recordings with validated clinical metadata and dense event annotations, all wrapped in a clear benchmark framework. By providing standardized data formats, unified evaluation protocols, and baseline models, the work lays a solid foundation for reproducible, generalizable, and clinically translatable machine‑learning research in epilepsy. Future directions include multimodal integration (imaging, genetics, electronic health records), real‑time decision support systems, and the development of transferable neural representations that can be fine‑tuned across diverse neurophysiological tasks.
Comments & Academic Discussion
Loading comments...
Leave a Comment