Relating biomarkers and phenotypes using dynamical trap spaces
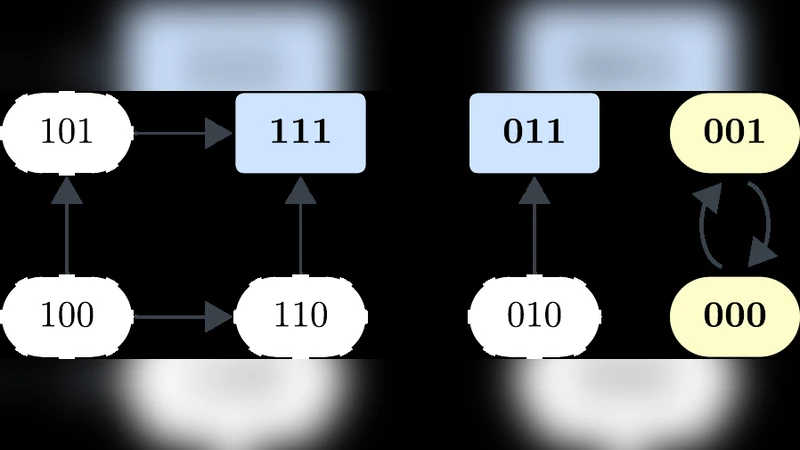
Connecting the dynamics of biomolecular networks to experimentally measurable cell phenotypes remains a central challenge in systems biology. Here we introduce a model-based definition of phenotype as a partial steady state that is committed to a certain dynamical outcome while otherwise being minimally constrained. We focus on Boolean models and define \emph{dynamical phenotypes} as complete trap spaces that maximally specify a chosen set of phenotype-determining nodes that correspond to biomarkers while keeping external inputs unconstrained. We show that dynamical phenotypes can be efficiently identified without full attractor enumeration. Using four published models, including a 70-node Boolean model of T cell differentiation, we show that dynamical phenotypes recover known cell types and activation states, and indicate the environmental conditions ensuring their existence. We also propose a method to identify informative phenotype-determining nodes based on the canalization of the Boolean functions. This method reveals biologically relevant cell state information that is complementary to the phenotypes manually defined by model creators and is validated by two attractor-based approaches. Our results demonstrate that dynamical phenotypes provide a scalable framework for linking model structure, external inputs, and phenotypic outcomes, and offer a principled tool for model-guided biomarker selection.
💡 Research Summary
The paper tackles the long‑standing problem of linking the dynamics of biomolecular networks to experimentally observable cell phenotypes. The authors propose a model‑based definition of phenotype as a partial steady state that is committed to a specific dynamical outcome while remaining otherwise unconstrained. Focusing on Boolean networks, they introduce the concept of dynamical phenotypes, which are defined as complete trap spaces that fully specify a chosen set of phenotype‑determining nodes (the biomarkers) but leave all external inputs free. This definition captures the idea that a phenotype is a set of fixed molecular markers that can arise under a variety of environmental conditions.
A trap space is a subset of the Boolean state space that, once entered, cannot be left; it therefore generalizes the notion of attractors (fixed points or cycles). The authors show that trap spaces can be identified efficiently using SAT/SMT encodings, avoiding the combinatorial explosion associated with exhaustive attractor enumeration. To select the most informative phenotype‑determining nodes, they exploit the canalization property of Boolean functions: nodes whose update functions are highly canalized are less sensitive to input variations and thus make robust biomarkers. A quantitative canalization score is computed for each node, and the top‑scoring nodes are automatically chosen as candidate biomarkers.
The methodology is applied to four published Boolean models, including a large 70‑node model of T‑cell differentiation, a metabolic network of E. coli, a human cell‑cycle regulatory network, and an oncogenic signaling model. In each case, the identified dynamical phenotypes recover known cell types or functional states (e.g., Th1, Th2, Th17, Treg in the T‑cell model) and explicitly indicate the external cytokine or growth‑factor conditions required for their existence. The authors also demonstrate that the canalization‑based biomarker selection yields markers that are consistent with two independent attractor‑based validation approaches: (i) fixed‑point enumeration and (ii) full attractor (including cycles) analysis.
Key contributions of the work are:
- Formal definition of phenotype as a minimally constrained partial steady state, bridging model structure and experimental observables.
- Efficient identification of dynamical phenotypes via trap‑space computation, which scales to large networks without full attractor enumeration.
- Systematic, data‑driven biomarker selection based on Boolean function canalization, providing a principled alternative to manual marker choice.
- Demonstration of biological relevance across multiple models, showing that dynamical phenotypes capture both known cell identities and the environmental contexts that enable them.
The authors discuss several implications. Because external inputs remain free, the framework naturally predicts the environmental cues (e.g., cytokine cocktails) that drive a cell toward a particular phenotype, offering testable hypotheses for experimental design. The scalability of trap‑space analysis suggests applicability to genome‑scale regulatory networks, while the canalization metric could be extended to multi‑valued or stochastic logical models. Future directions include integration with quantitative omics data, validation in wet‑lab experiments, and incorporation into precision‑medicine pipelines for patient‑specific biomarker discovery.
In summary, the study provides a mathematically rigorous, computationally tractable, and biologically insightful approach to connect Boolean network dynamics with measurable phenotypic outcomes, establishing dynamical trap spaces as a powerful tool for model‑guided biomarker identification and phenotype prediction.
Comments & Academic Discussion
Loading comments...
Leave a Comment