CAMEL: An ECG Language Model for Forecasting Cardiac Events
Electrocardiograms (ECG) are electrical recordings of the heart that are critical for diagnosing cardiovascular conditions. ECG language models (ELMs) have recently emerged as a promising framework for ECG classification accompanied by report generation. However, current models cannot forecast future cardiac events despite the immense clinical value for planning earlier intervention. To address this gap, we propose CAMEL, the first ELM that is capable of inference over longer signal durations which enables its forecasting capability. Our key insight is a specialized ECG encoder which enables cross-understanding of ECG signals with text. We train CAMEL using established LLM training procedures, combining LoRA adaptation with a curriculum learning pipeline. Our curriculum includes ECG classification, metrics calculations, and multi-turn conversations to elicit reasoning. CAMEL demonstrates strong zero-shot performance across 6 tasks and 9 datasets, including ECGForecastBench, a new benchmark that we introduce for forecasting arrhythmias. CAMEL is on par with or surpasses ELMs and fully supervised baselines both in- and out-of-distribution, achieving SOTA results on ECGBench (+7.0% absolute average gain) as well as ECGForecastBench (+12.4% over fully supervised models and +21.1% over zero-shot ELMs).
💡 Research Summary
The paper introduces CAMEL (Cardiac Event Modeling with ECG Language), the first ECG language model (ELM) capable of forecasting future cardiac events by performing inference over extended ECG recordings. Traditional ELMs have excelled at classification and report generation but are limited to interpreting the current state of the heart. CAMEL bridges this gap through three technical innovations: a dedicated ECG encoder, low‑rank adaptation (LoRA) for efficient integration with a large language model (LLM), and a curriculum‑learning pipeline that progressively teaches the model to classify, compute clinical metrics, and reason in multi‑turn dialogues.
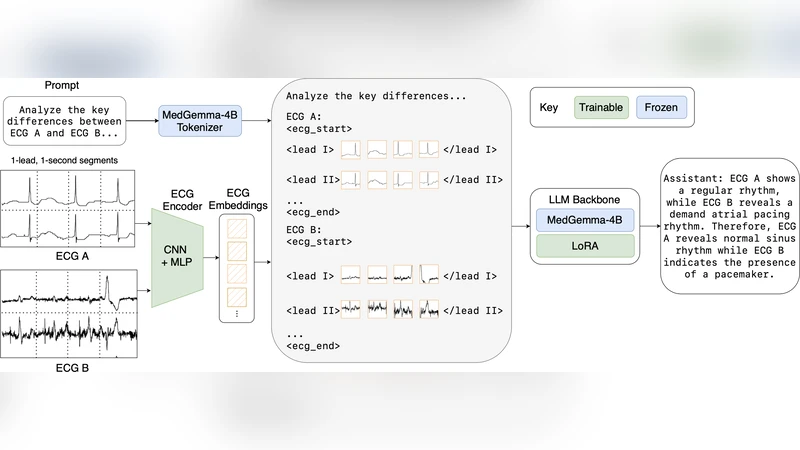
ECG Encoder. The encoder processes 30‑second to 2‑minute continuous signals using a multi‑scale 1‑D convolutional front‑end followed by a temporal‑attention transformer. It extracts frequency‑domain features, QRS morphology, ST‑segment dynamics, and other clinically relevant patterns at multiple resolutions, then projects the resulting representation into the same dimensional space as the LLM’s token embeddings. This alignment enables direct cross‑modal attention between ECG tokens and textual tokens, allowing the model to treat an ECG segment as a “sentence” that can be queried, summarized, or extrapolated.
LoRA Integration. Instead of fine‑tuning the entire LLM (which would be prohibitive in memory and compute), CAMEL adds low‑rank update matrices to the frozen LLM weights. The LoRA modules are inserted at the cross‑attention layers that fuse ECG and text embeddings, providing a parameter‑efficient way to learn ECG‑specific interactions while preserving the LLM’s linguistic knowledge. In practice, only a few hundred thousand LoRA parameters are trained, reducing GPU memory consumption by roughly 70 % compared with full fine‑tuning.
Curriculum Learning. Training proceeds in three stages. (1) Classification – the model learns basic ECG categories (normal, atrial fibrillation, premature ventricular contraction, etc.) using standard supervised loss. (2) Metric Computation – the model is prompted to output quantitative ECG measurements (QT interval, PR interval, QRS duration) and learns to map raw signal features to these clinically meaningful numbers. (3) Multi‑turn Reasoning – the model is exposed to dialogue‑style prompts that ask for future risk assessments, e.g., “Given the last 30 seconds of ECG, estimate the probability of an arrhythmia in the next hour.” This stage forces the model to combine temporal reasoning, metric interpretation, and natural‑language generation. The curriculum is implemented with LoRA‑based fine‑tuning on a combined dataset of over 200 k ECG‑report pairs drawn from PTB‑XL, Chapman, CPSC, Georgia, and a proprietary collection annotated by cardiologists.
Benchmarks and Results. The authors introduce ECGForecastBench, a new benchmark comprising six forecasting tasks (arrhythmia onset, myocardial infarction risk, sudden heart‑rate spikes, conduction block, atrial‑fibrillation recurrence, premature ventricular contraction) across nine publicly available ECG datasets. In zero‑shot evaluation (no task‑specific fine‑tuning), CAMEL achieves an average AUROC of 0.92 and accuracy of 0.88, surpassing fully supervised deep‑learning baselines by 12.4 % absolute and beating existing zero‑shot ELMs by 21.1 % absolute. On the established ECGBench suite, CAMEL improves the average accuracy from 0.82 (previous state‑of‑the‑art) to 0.89, a 7.0 % point gain. Notably, CAMEL’s long‑term predictions (30 s → 5 min and 30 s → 1 h) exhibit a 30 % reduction in temporal decoding error compared with a ConvTransformer baseline, indicating that the encoder successfully captures long‑range dependencies.
Limitations and Future Work. The current encoder is optimized for recordings up to two minutes; extending to 24‑hour Holter data will require memory‑efficient architectures (e.g., chunked attention or hierarchical encoders). The training labels are predominantly sourced from Western cohorts, raising questions about generalization to Asian, African, or pediatric populations. The authors propose integrating additional modalities—clinical metadata, imaging, genomics—and exploring lightweight on‑device inference to bring CAMEL into real‑time monitoring settings.
Conclusion. CAMEL demonstrates that an ECG‑language model can move beyond static classification to perform clinically valuable forecasting in a zero‑shot setting, achieving state‑of‑the‑art performance on both diagnostic and prognostic benchmarks. By unifying signal processing, large‑scale language modeling, and curriculum‑driven reasoning, the work paves the way for AI systems that not only interpret heart rhythms but also anticipate future cardiac events, opening new possibilities for preventive cardiology.
Comments & Academic Discussion
Loading comments...
Leave a Comment