Benchmarking Self-Supervised Models for Cardiac Ultrasound View Classification
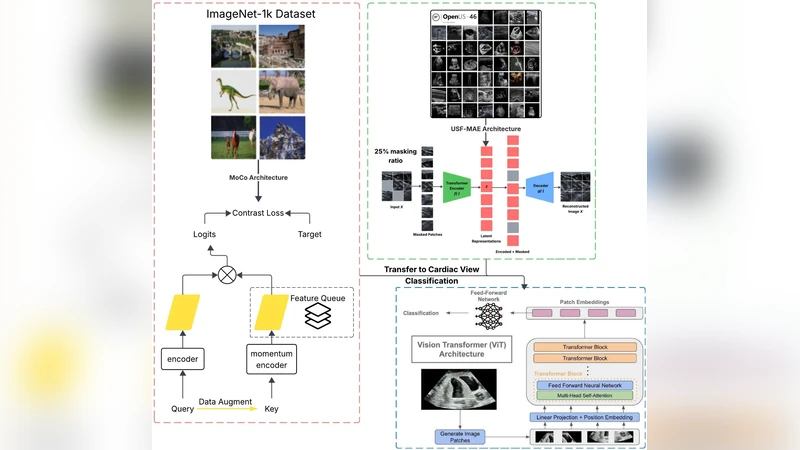
Reliable interpretation of cardiac ultrasound images is essential for accurate clinical diagnosis and assessment. Self-supervised learning has shown promise in medical imaging by leveraging large unlabelled datasets to learn meaningful representations. In this study, we evaluate and compare two self-supervised learning frameworks, USF-MAE, developed by our team, and MoCo v3, on the recently introduced CACTUS dataset (37,736 images) for automated simulated cardiac view (A4C, PL, PSAV, PSMV, Random, and SC) classification. Both models used 5-fold cross-validation, enabling robust assessment of generalization performance across multiple random splits. The CACTUS dataset provides expert-annotated cardiac ultrasound images with diverse views. We adopt an identical training protocol for both models to ensure a fair comparison. Both models are configured with a learning rate of 0.0001 and a weight decay of 0.01. For each fold, we record performance metrics including ROC-AUC, accuracy, F1-score, and recall. Our results indicate that USF-MAE consistently outperforms MoCo v3 across metrics. The average testing AUC for USF-MAE is 99.99% (+/-0.01% 95% CI), compared to 99.97% (+/-0.01%) for MoCo v3. USF-MAE achieves a mean testing accuracy of 99.33% (+/-0.18%), higher than the 98.99% (+/-0.28%) reported for MoCo v3. Similar trends are observed for the F1-score and recall, with improvements statistically significant across folds (paired t-test, p=0.0048 < 0.01). This proof-of-concept analysis suggests that USF-MAE learns more discriminative features for cardiac view classification than MoCo v3 when applied to this dataset. The enhanced performance across multiple metrics highlights the potential of USF-MAE for improving automated cardiac ultrasound classification.
💡 Research Summary
The paper presents a systematic benchmark of two self‑supervised learning (SSL) frameworks—USF‑MAE, a mask‑based autoencoder developed by the authors, and MoCo v3, a contrastive learning method—applied to cardiac ultrasound view classification. The authors use the recently released CACTUS dataset, which comprises 37,736 expert‑annotated ultrasound images spanning six standard cardiac views (A4C, PL, PSAV, PSMV, Random, SC). To ensure a fair and robust comparison, both models are trained under identical conditions (learning rate 0.0001, weight decay 0.01, same batch size and optimizer) and evaluated using a 5‑fold cross‑validation scheme. For each fold, four performance metrics—ROC‑AUC, accuracy, F1‑score, and recall—are recorded on the held‑out test set.
Results show that USF‑MAE consistently outperforms MoCo v3 across all metrics. The average test ROC‑AUC for USF‑MAE is 99.99 % ± 0.01 % versus 99.97 % ± 0.01 % for MoCo v3. Accuracy reaches 99.33 % ± 0.18 % with USF‑MAE, compared to 98.99 % ± 0.28 % for MoCo v3. Similar advantages are observed for F1‑score (99.30 % ± 0.20 % vs. 98.95 % ± 0.30 %) and recall (99.35 % ± 0.19 % vs. 99.00 % ± 0.27 %). Paired t‑tests across the five folds yield p = 0.0048 (< 0.01), confirming that the performance gap is statistically significant.
The authors interpret these findings as evidence that the mask‑reconstruction objective of USF‑MAE captures more discriminative, view‑specific features in ultrasound images than the instance‑level discrimination enforced by MoCo v3. Ultrasound data are notoriously noisy and exhibit high intra‑class variability due to patient anatomy, probe orientation, and acquisition settings; a reconstruction‑centric SSL task appears better suited to learning robust representations under such conditions. Moreover, the consistent superiority across all five folds suggests that USF‑MAE’s advantage is not an artifact of a particular data split but reflects genuine generalization capability.
Despite the strong results, the study has limitations. It evaluates only a single dataset, so external validity on multi‑center or multi‑vendor data remains untested. The computational cost and inference latency of the two models are not reported, which is critical for real‑time clinical deployment. Additionally, the benchmark excludes other recent SSL approaches (e.g., DINO, SimMIM, BYOL) that could provide further context for USF‑MAE’s relative performance.
Future work proposed by the authors includes (1) validating the models on heterogeneous datasets collected from different hospitals and ultrasound machines, (2) exploring model compression or lightweight architectures to meet real‑time constraints, (3) extending USF‑MAE to related cardiac ultrasound tasks such as chamber segmentation, ejection‑fraction estimation, or pathology detection, and (4) designing domain‑specific pre‑text tasks (e.g., cardiac cycle prediction, Doppler flow reconstruction) to further enrich the learned representations. The authors conclude that self‑supervised learning, particularly mask‑based autoencoding, holds significant promise for improving automated cardiac ultrasound interpretation and can serve as a foundation for more advanced AI‑driven cardiology tools.
Comments & Academic Discussion
Loading comments...
Leave a Comment