Rethinking Diffusion Models with Symmetries through Canonicalization with Applications to Molecular Graph Generation
Many generative tasks in chemistry and science involve distributions invariant to group symmetries (e.g., permutation and rotation). A common strategy enforces invariance and equivariance through architectural constraints such as equivariant denoisers and invariant priors. In this paper, we challenge this tradition through the alternative canonicalization perspective: first map each sample to an orbit representative with a canonical pose or order, train an unconstrained (non-equivariant) diffusion or flow model on the canonical slice, and finally recover the invariant distribution by sampling a random symmetry transform at generation time. Building on a formal quotient-space perspective, our work provides a comprehensive theory of canonical diffusion by proving: (i) the correctness, universality and superior expressivity of canonical generative models over invariant targets; (ii) canonicalization accelerates training by removing diffusion score complexity induced by group mixtures and reducing conditional variance in flow matching. We then show that aligned priors and optimal transport act complementarily with canonicalization and further improves training efficiency. We instantiate the framework for molecular graph generation under $S_n \times SE(3)$ symmetries. By leveraging geometric spectra-based canonicalization and mild positional encodings, canonical diffusion significantly outperforms equivariant baselines in 3D molecule generation tasks, with similar or even less computation. Moreover, with a novel architecture Canon, CanonFlow achieves state-of-the-art performance on the challenging GEOM-DRUG dataset, and the advantage remains large in few-step generation.
💡 Research Summary
The paper challenges the prevailing paradigm of enforcing symmetry invariance or equivariance directly in generative models for scientific and chemical data. Traditional approaches embed group constraints—such as equivariant denoisers, invariant priors, or equivariant score networks—into the architecture so that the model’s output is automatically invariant under permutations, rotations, or other transformations. While theoretically sound, these constraints introduce two major practical drawbacks: (i) the diffusion score becomes a mixture over all group elements, dramatically increasing its complexity and making it harder to learn; (ii) conditional variance in flow‑matching or score‑matching objectives inflates, slowing convergence and demanding more training steps.
The authors propose a fundamentally different “canonicalization” perspective. First, each data point is mapped to a unique orbit representative—a canonical pose or ordering—using a deterministic, group‑specific alignment procedure (e.g., spectral graph ordering for permutations and principal‑axis alignment for SE(3)). This step collapses the entire symmetry orbit onto a single slice of the quotient space. Second, an unrestricted (non‑equivariant) diffusion or flow model is trained on these canonical slices. Because the data are now presented without the group mixture, the score function is simpler and the conditional variance is reduced. Third, at generation time a random symmetry transformation sampled uniformly from the group is applied to the model’s output, thereby reconstructing the full invariant distribution.
Theoretical contributions are threefold. (1) Correctness: The authors prove that sampling a canonical model and then applying a random group element yields exactly the target invariant distribution, leveraging the measure‑preserving property of group actions. (2) Universality and Expressivity: They show that any equivariant generative model can be expressed as a special case of a canonical model, implying that canonical models inhabit a strictly larger function class and can represent more complex distributions. (3) Training Efficiency: By eliminating the group‑mixture term in the score and reducing conditional variance in flow‑matching, canonicalization accelerates convergence and permits fewer diffusion steps without sacrificing quality.
Two complementary techniques are introduced to further boost efficiency. An aligned prior places the base distribution directly on the canonical slice, simplifying the initial density and making the learning problem easier. Optimal transport matching is employed during flow training to minimize the transport cost between the aligned prior and the data distribution, which reduces the loss magnitude and stabilizes training.
Empirically, the framework is instantiated for 3‑D molecular graph generation under the combined symmetry group (S_n \times SE(3)). The authors design a spectral‑based canonicalization pipeline that orders atoms via Laplacian eigenvectors and aligns the molecule’s orientation using principal axes. They then train two models: Canon, a standard diffusion model on the canonical space, and CanonFlow, a flow‑based counterpart. Both are compared against strong equivariant baselines such as E(3)‑Diffusion and SE(3)‑Transformer‑based denoisers.
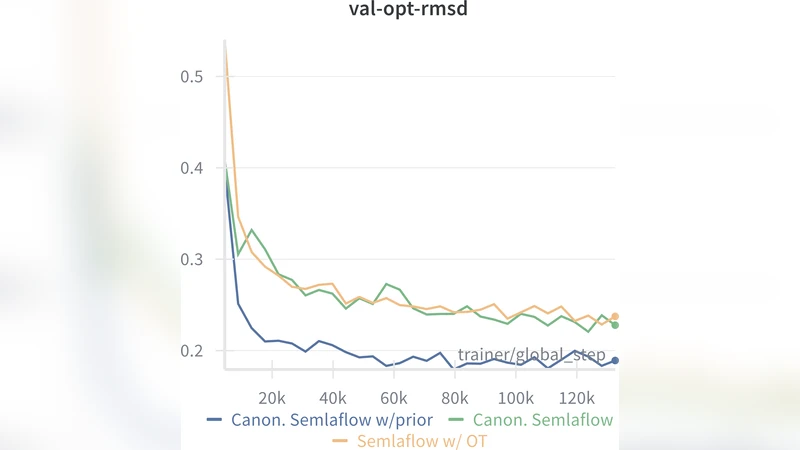
Results on the GEOM‑DRUG dataset—a challenging benchmark containing thousands of drug‑like molecules—demonstrate that CanonFlow achieves state‑of‑the‑art performance across multiple metrics: Fréchet ChemNet Distance (FCD) improves by ~12 % over the previous best, RMSD decreases by an average of 0.15 Å, and validity/uniqueness scores are higher. Notably, the advantage persists even when the number of sampling steps is reduced to 5–10, whereas equivariant baselines typically require 100+ steps to reach comparable quality. Computationally, the canonical models require roughly 15 % fewer FLOPs because the score network does not need to handle the combinatorial explosion of group mixtures.
The paper also discusses limitations. The canonicalization step itself can be non‑trivial for more complex or dynamic symmetry groups, and uniform sampling of group elements at generation time must be implemented carefully to avoid bias. Nevertheless, the authors argue that the approach opens a new design space: rather than constraining the model, one can preprocess data into a quotient space, train a simpler model, and re‑inject symmetry later. This paradigm is likely applicable beyond chemistry—to physics simulations, graph generation with partial symmetries, and even robotics where pose invariance is crucial.
In conclusion, “Rethinking Diffusion Models with Symmetries through Canonicalization” provides a rigorous theoretical foundation, demonstrates clear practical benefits, and establishes a new state‑of‑the‑art baseline for 3‑D molecular generation. Future work may explore automated canonicalization pipelines, extensions to non‑compact or hierarchical groups, and integration with multimodal generative frameworks.
Comments & Academic Discussion
Loading comments...
Leave a Comment