Hidden Markov Individual-level Models of Infectious Disease Transmission
Individual-level epidemic models are increasingly being used to help understand the transmission dynamics of various infectious diseases. However, fitting such models to individual-level epidemic data is challenging, as we often only know when an individual’s disease status was detected (e.g., when they showed symptoms) and not when they were infected or removed. We propose an autoregressive coupled hidden Markov model to infer unknown infection and removal times, as well as other model parameters, from a single observed detection time for each detected individual. Unlike more traditional data augmentation methods used in epidemic modelling, we do not assume that this detection time corresponds to infection or removal or that infected individuals must at some point be detected. Bayesian coupled hidden Markov models have been used previously for individual-level epidemic data. However, these approaches assumed each individual was continuously tested and that the tests were independent. In practice, individuals are often only tested until their first positive test, and even if they are continuously tested, only the initial detection times may be reported. In addition, multiple tests on the same individual may not be independent. We accommodate these scenarios by assuming that the probability of detecting the disease can depend on past observations, which allows us to fit a much wider range of practical applications. We illustrate the flexibility of our approach by fitting two examples: an experiment on the spread of tomato spot wilt virus in pepper plants and an outbreak of norovirus among nurses in a hospital.
💡 Research Summary
This paper tackles a central challenge in individual‑level epidemic modelling: the latent nature of infection and removal times when only a single detection time (e.g., the first positive test or symptom onset) is recorded for each case. Traditional Bayesian data‑augmentation approaches typically assume that the observed detection coincides with infection or removal, or that every infected individual will eventually be detected. Both assumptions are rarely satisfied in practice, where testing may stop after the first positive result, and many infections remain undetected.
To overcome these limitations, the authors propose an autoregressive coupled hidden Markov model (AR‑coupled HMM). The model consists of two hidden state sequences—infection (I) and removal (R)—each evolving as a first‑order Markov chain. Transmission between individuals is governed by a network‑based infection rate β, while removal follows an individual‑specific rate γ. The novel observation component is a detection probability function ψ_t that depends on the entire past detection history of the individual. By allowing ψ_t to be a function of previous positive tests and the elapsed time since the last test, the model captures (i) the fact that detection does not necessarily align with infection or removal, (ii) the common practice of stopping testing after the first positive, and (iii) potential dependence among repeated tests on the same person.
For inference, the authors adopt a fully Bayesian framework. Priors are placed on β, γ, and the parameters governing ψ (typically a logistic autoregressive regression). Posterior sampling is performed with a hybrid Markov chain Monte Carlo (MCMC) scheme: the hidden infection and removal trajectories are sampled using a forward‑backward algorithm, while the autoregressive coefficients of ψ are updated via Metropolis‑Hastings steps. The conditional posterior distributions retain tractable forms because ψ is modeled with a logistic link, which preserves conjugacy‑like properties and improves mixing.
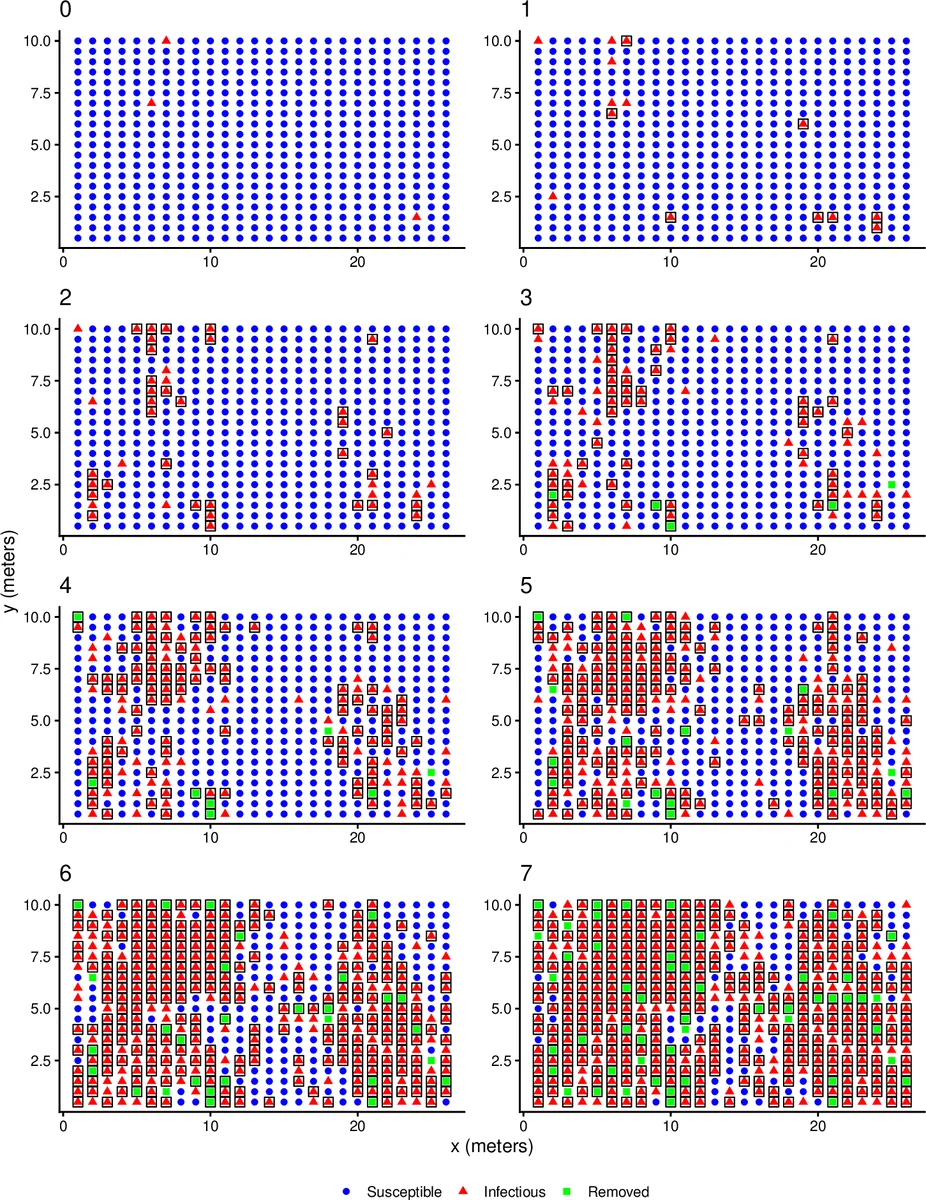
The methodology is validated through simulation studies that vary network topology, detection probabilities, and the degree of autocorrelation in ψ. Across all scenarios the AR‑coupled HMM accurately recovers the true infection times, removal times, and transmission parameters, demonstrating robustness to misspecification of detection processes.
Two real‑world applications illustrate the practical value of the approach.
- Tomato spot wilt virus experiment – In a controlled greenhouse study, pepper plants were inoculated with a virus, but only the first positive PCR result for each plant was recorded. Applying the AR‑coupled HMM, the authors reconstructed the latent infection chronology and estimated the plant‑to‑plant transmission rate. The inferred infection order matched the known experimental inoculation sequence, and the estimated removal (plant death) rate aligned with observed mortality patterns.
- Norovirus outbreak among hospital nurses – During a hospital outbreak, nurses reported the date of symptom onset, and testing ceased after the first positive result. The model identified a plausible transmission network among staff, highlighted a subset of nurses who were likely infected but never reported symptoms (asymptomatic carriers), and provided posterior distributions for the basic reproduction number within the ward. These insights extended beyond the original epidemiological report, which had only described aggregate case counts.
The paper’s contributions are threefold. First, it relaxes the restrictive assumption that detection equals infection or removal, allowing analysts to work with the sparse data that are often the only information available. Second, by incorporating an autoregressive detection probability, the model accommodates realistic testing protocols where observations are not independent and may stop after the first positive. Third, the Bayesian formulation yields full posterior uncertainty for all latent quantities, facilitating risk‑based decision making.
Limitations are acknowledged. The MCMC algorithm, while exact, can be computationally intensive for large populations, suggesting a need for scalable alternatives such as variational inference or Hamiltonian Monte Carlo. Moreover, the functional form of ψ must be specified a priori; misspecification could bias estimates if the true detection mechanism is highly nonlinear or involves unobserved covariates. The authors propose future work on non‑parametric or machine‑learning‑based detection models, integration of environmental transmission pathways, and real‑time updating for outbreak surveillance.
In summary, this study introduces a flexible, theoretically sound, and empirically validated framework for inferring hidden epidemic dynamics from minimal detection data. By bridging the gap between idealized continuous testing assumptions and the fragmented reality of field data, the AR‑coupled hidden Markov model offers a powerful tool for epidemiologists, public health officials, and researchers seeking to understand and control infectious disease spread at the individual level.
Comments & Academic Discussion
Loading comments...
Leave a Comment