Out-of-equilibrium selection pressure enhances inference from protein sequence data
Homologous proteins have similar three-dimensional structures and biological functions that shape their sequences. The resulting coevolution-driven correlations underlie methods from Potts models to AlphaFold, which infer protein structure and function from sequences. Using a minimal model, we show that fluctuating selection strength and the onset of new selection pressures improve coevolution-based inference of structural contacts. Our conclusions extend to realistic synthetic data and to the inference of interaction partners. Out-of-equilibrium noise arising from ubiquitous variations in natural selection thus enhances, rather than hinders, the success of inference from protein sequences.
💡 Research Summary
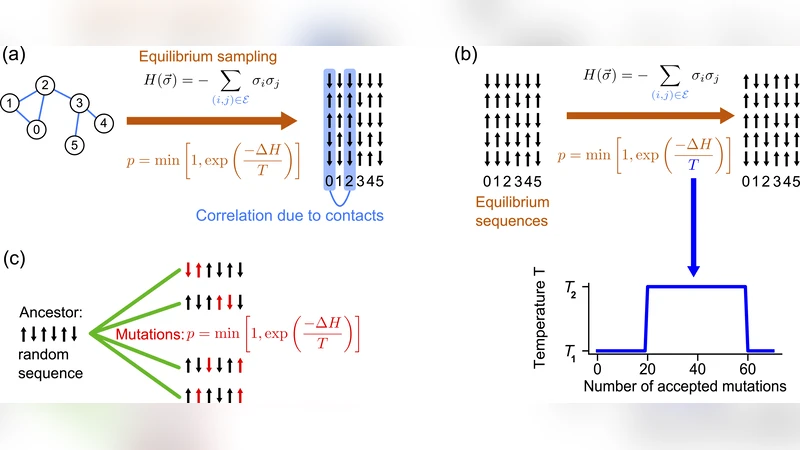
In this paper the authors investigate how temporal fluctuations in natural selection pressure affect the ability to infer structural contacts and interaction partners from protein sequence data. Using a highly simplified Ising‑spin model, protein sequences are represented as binary strings placed on the nodes of an Erdős‑Rényi random graph, with each edge corresponding to a structural contact and assigned a ferromagnetic coupling of unit strength. The Hamiltonian H = −∑_{(i,j)∈E}σ_iσ_j captures the energetic cost of violating contacts, while the sampling temperature T controls selection strength: low T enforces strong selection (few mutations accepted), high T corresponds to weak selection (many mutations accepted).
The core of the study introduces a telegraph process that switches the temperature between two values, T₁ = 1 (strong selection) and T₂ = 15 (weak selection), with a tunable switching timescale τ. Sequences are generated by Metropolis Monte‑Carlo dynamics, and a multiple‑sequence alignment (MSA) of 2048 sequences of length 200 is assembled for each experimental condition. Pairwise maximum‑entropy models (mfDCA or plmDCA) are then fitted to the MSA, and the quality of contact prediction is quantified by the True Positive (TP) fraction, i.e., the proportion of true graph edges among the top‑N inferred couplings (N equals the true number of contacts).
Key findings from the minimal model are:
- Fluctuating selection dramatically improves TP compared with either fixed T₁ or fixed T₂. The average TP rises rapidly as mutations accumulate, then plateaus at a level comparable to the optimal equilibrium temperature (≈ T_C = 4, the ferromagnetic‑paramagnetic transition point).
- Faster switching (small τ) yields higher TP because it maintains a high effective sequence diversity (M_eff ≈ 2000) while still allowing occasional strong‑selection phases that suppress noise.
- When each sequence experiences an independent temperature trajectory (heterogeneous selection histories), the improvement is even larger, reflecting the biological reality that orthologous proteins from different species may have distinct evolutionary pressures.
The authors then test whether these observations hold for realistic synthetic data. They infer a Potts model from a natural MSA of the PF0004 (AAA ATPase) family using bmDCA, and generate 70 000 synthetic sequences from this model. Applying the same telegraph switching between T₁ = 1 and T₂ = 15 reproduces the TP boost seen in the minimal model; the best performance matches that obtained at the optimal static temperature (T = 1). Thus, the phenomenon is not an artifact of the oversimplified Ising representation.
Beyond fluctuating environments, the paper examines the “onset of a new selection pressure.” Starting from a random ancestor, sequences evolve under a fixed temperature T (strong selection) along a star‑shaped phylogeny. For T < T_C, the TP fraction exhibits a pronounced maximum at an intermediate number of accepted mutations per branch (μ). At this point the system has transiently increased diversity before the strong selection freezes variation, leading to better contact inference than at equilibrium. This effect persists for balanced binary trees as well.
Finally, the authors extend the analysis to protein‑protein interaction partner prediction. Using the same minimal model and a partner‑matching protocol based on mfDCA scores, they show that fluctuating selection also raises the TP fraction for partner inference, indicating that the benefit of out‑of‑equilibrium dynamics is not limited to intra‑protein contacts.
In the discussion, the authors argue that temporal variability in selection strength creates non‑equilibrium “noise” that temporarily expands the accessible sequence space, thereby amplifying the co‑evolutionary signal that maximum‑entropy models exploit. This insight suggests that the remarkable success of DCA‑based methods and even deep‑learning approaches such as AlphaFold may partly rely on the natural history of fluctuating selective constraints. They propose practical implications for directed‑evolution experiments: deliberately alternating selection stringency (e.g., by varying antibiotic concentrations) could improve structural predictions from laboratory‑evolved libraries. Moreover, they speculate that similar principles may apply to other systems modeled by pairwise maximum‑entropy models, such as neuronal populations driven by time‑varying external inputs.
Overall, the study provides a compelling theoretical and computational demonstration that out‑of‑equilibrium evolutionary dynamics, driven by fluctuating selection pressures, can enhance the extraction of structural and functional information from protein sequence data. This challenges the conventional view that evolutionary noise is detrimental, and opens new avenues for both interpreting natural sequence diversity and designing more effective experimental evolution protocols.
Comments & Academic Discussion
Loading comments...
Leave a Comment