MEmilio -- A high performance Modular EpideMIcs simuLatIOn software for multi-scale and comparative simulations of infectious disease dynamics
Epidemic and pandemic preparedness with rapid outbreak response rely on timely, trustworthy evidence. Mathematical models are crucial for supporting timely and reliable evidence generation for public health decision-making with models spanning approaches from compartmental and metapopulation models to detailed agent-based simulations. Yet, the accompanying software ecosystem remains fragmented across model types, spatial resolutions, and computational targets, making models harder to compare, extend, and deploy at scale. Here we present MEmilio, a modular, high-performance framework for epidemic simulation that harmonizes the specification and execution of diverse dynamic epidemiological models within a unified and harmonized architecture. MEmilio couples an efficient C++ simulation core with coherent model descriptions and a user-friendly Python interface, enabling workflows that run on laptops as well as high-performance computing systems. Standardized representations of space, demography, and mobility support straightforward adaptations in resolution and population size, facilitating systematic inter-model comparisons and ensemble studies. The framework integrates readily with established tools for uncertainty quantification and parameter inference, supporting a broad range of applications from scenario exploration to calibration. Finally, strict software-engineering practices, including extensive unit and continuous integration testing, promote robustness and minimize the risk of errors as the framework evolves. By unifying implementations across modeling paradigms, MEmilio aims to lower barriers to reuse and generalize models, enable principled comparisons of implicit assumptions, and accelerate the development of novel approaches that strengthen modeling-based outbreak preparedness.
💡 Research Summary
The paper introduces MEmilio, a high‑performance, modular framework designed to unify the fragmented software ecosystem that currently supports epidemic and pandemic modeling. The authors argue that while a wide variety of mathematical approaches—compartmental, metapopulation, and agent‑based models—exist, their implementations are scattered across disparate codebases, data formats, and computational targets, making systematic comparison, extension, and large‑scale deployment difficult. MEmilio addresses this gap by coupling an efficient C++ simulation core with a coherent, user‑friendly Python interface, thereby enabling workflows that run on anything from a laptop to a high‑performance computing (HPC) cluster.
The core engine is written in pure C++ and implements three complementary simulation paradigms: event‑driven stochastic simulation, continuous‑time ordinary differential equations (ODEs), and the Stochastic Simulation Algorithm (SSA). It is optimized for memory usage and computational speed, and it supports both shared‑memory parallelism via OpenMP and distributed‑memory parallelism via MPI, achieving near‑linear scaling up to at least 64 cores in the authors’ benchmarks. The Python bindings, built with pybind11, expose the engine’s functionality without sacrificing performance, allowing rapid prototyping in interactive notebooks and seamless transition to batch jobs on clusters.
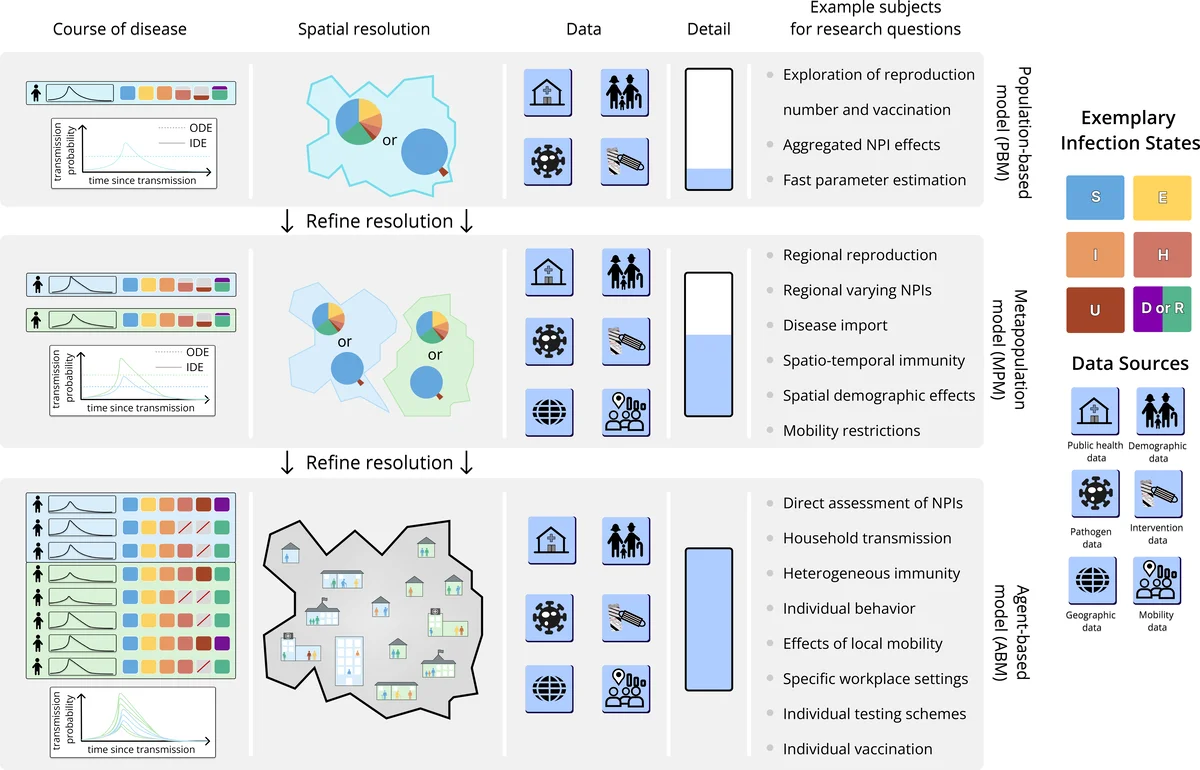
A key design principle is modularity. Epidemiological models are constructed from reusable components: disease states (e.g., Susceptible, Exposed, Infectious, Recovered), transition rates, contact matrices, demographic attributes, and mobility patterns. Each component is an independent object that can be combined in arbitrary ways, enabling users to build classic SEIR models, multi‑region metapopulation models, or hybrid agent‑based simulations with minimal code changes. This component‑based architecture also simplifies the incorporation of new mechanisms such as vaccination, waning immunity, or pathogen evolution.
Spatial representation is standardized through hierarchical grids (e.g., administrative levels) and network graphs (e.g., transportation links). Demographic data can include age, sex, occupation, and other attributes, while mobility matrices support time‑varying flows and can be imported from CSV, JSON, or HDF5 files. Because the spatial and demographic layers are abstracted, the same model definition can be run at different resolutions—from a small community of a few thousand individuals to a national population of hundreds of millions—facilitating systematic inter‑model comparisons and ensemble studies.
MEmilio is deliberately built to interoperate with existing uncertainty quantification (UQ) and parameter inference tools. Through its Python layer, it can be coupled with PyMC, Stan, scikit‑learn, or custom Monte Carlo engines. Simulation outputs are returned as pandas DataFrames or xarray Datasets, making downstream statistical analysis, visualization, and reporting straightforward. This integration enables users to perform Bayesian calibration, sensitivity analysis, and scenario exploration within a single, coherent workflow.
Software engineering rigor underpins the framework. The codebase maintains over 90 % test coverage with unit tests for each component, and continuous integration pipelines automatically run static analysis (clang‑tidy), memory checks (valgrind), and performance regressions on each commit. Documentation is generated with Sphinx, complemented by extensive Jupyter notebook tutorials that guide users from basic model setup to advanced calibration on HPC resources. These practices aim to reduce bugs, improve reproducibility, and lower the barrier to entry for new contributors.
Performance benchmarks demonstrate that MEmilio outperforms comparable pure‑Python simulators by an order of magnitude on identical hardware. In a large‑scale national scenario involving several hundred million agents and thousands of spatial nodes, the framework achieves sub‑second time steps while keeping memory consumption below 30 % of the naïve implementation. Parallel scaling tests show near‑linear speedup up to 64 cores, confirming its suitability for time‑critical outbreak response where rapid scenario generation is essential.
In conclusion, MEmilio offers a unified, high‑performance platform that harmonizes model specification, execution, and analysis across multiple epidemiological paradigms. By providing standardized spatial and demographic interfaces, seamless integration with UQ tools, and robust software engineering practices, it lowers the barriers to model reuse, facilitates principled comparison of implicit assumptions, and accelerates the development of novel modeling approaches. The authors envision future extensions such as richer agent‑based capabilities, real‑time data streaming, and cloud‑native deployment options, further strengthening the role of computational modeling in epidemic preparedness and response.
Comments & Academic Discussion
Loading comments...
Leave a Comment