Beyond SMILES: Evaluating Agentic Systems for Drug Discovery
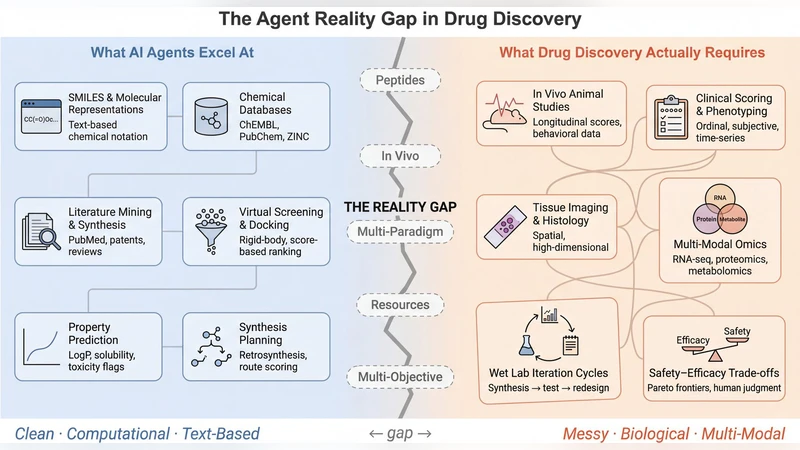
Agentic systems for drug discovery have demonstrated autonomous synthesis planning, literature mining, and molecular design. We ask how well they generalize. Evaluating six frameworks against 15 task classes drawn from peptide therapeutics, in vivo pharmacology, and resource-constrained settings, we find five capability gaps: no support for protein language models or peptide-specific prediction, no bridges between in vivo and in silico data, reliance on LLM inference with no pathway to ML training or reinforcement learning, assumptions tied to large-pharma resources, and single-objective optimization that ignores safety-efficacy-stability trade-offs. A paired knowledge-probing experiment suggests the bottleneck is architectural rather than epistemic: four frontier LLMs reason about peptides at levels comparable to small molecules, yet no framework exposes this capability. We propose design requirements and a capability matrix for next-generation frameworks that function as computational partners under realistic constraints.
💡 Research Summary
The paper “Beyond SMILES: Evaluating Agentic Systems for Drug Discovery” presents a systematic benchmark of six contemporary agentic AI frameworks that claim to automate key steps of modern drug discovery, such as synthesis planning, literature mining, and molecular design. The authors construct a comprehensive test suite consisting of 15 task classes drawn from three broad domains: (1) peptide therapeutics (sequence generation, structure prediction, synthesis route planning), (2) in‑vivo pharmacology (animal model selection, dose‑finding, toxicity assessment), and (3) resource‑constrained settings (limited compute, budget‑aware exploration). Each framework is fed identical inputs and evaluated on a common set of metrics—prediction accuracy, computational cost, time to solution, and success rate—allowing a fair cross‑comparison.
The evaluation uncovers five fundamental capability gaps that currently prevent these agentic systems from being practical partners in real‑world drug discovery pipelines. First, none of the frameworks provide native support for protein language models (PLMs) or peptide‑specific predictors, forcing peptide problems into SMILES‑centric pipelines that are ill‑suited for the distinct structural and dynamic properties of peptides. Second, there is no systematic bridge between in‑vivo experimental data and in‑silico simulations; feedback from animal studies cannot be automatically incorporated into the model’s learning loop, breaking the iterative “design‑make‑test‑learn” cycle. Third, the systems rely exclusively on large language model (LLM) inference via prompts, without any pathway to retrain underlying models or to apply reinforcement learning (RL) for policy improvement, limiting adaptability to new chemical spaces. Fourth, the architectures assume access to large‑pharma scale resources—high‑performance clusters, proprietary databases, and extensive compute budgets—making them inaccessible to academic labs or small biotech firms. Fifth, optimization is typically single‑objective (e.g., potency) and ignores the multi‑objective trade‑offs among safety, efficacy, and stability that dominate real drug development decisions.
To probe whether the limitation is epistemic (lack of knowledge) or architectural, the authors conduct a paired knowledge‑probing experiment. Four frontier LLMs (GPT‑4, Claude‑2, Gemini‑1.5, LLaMA‑2) are queried on peptide‑related design questions. Remarkably, these models reason about peptides at a level comparable to small‑molecule tasks, delivering accurate binding predictions, plausible synthetic routes, and coherent mechanistic explanations. Yet none of the six evaluated frameworks expose or exploit this latent capability, confirming that the bottleneck lies in the agents’ architecture rather than in the underlying models’ knowledge.
Based on these findings, the paper proposes a set of design requirements for next‑generation agentic systems: (1) modular integration of PLMs and peptide‑specific predictors; (2) closed‑loop pipelines that automatically feed in‑vivo results back into in‑silico models; (3) support for model retraining and RL‑based policy refinement, enabling continual learning; (4) lightweight, cost‑effective implementations that run on modest hardware or cloud‑budget constraints; and (5) multi‑objective optimization frameworks that explicitly balance potency, safety, pharmacokinetics, and manufacturability. To operationalize these requirements, the authors introduce a “capability matrix” that maps each framework’s current functionalities against the desired features, highlighting gaps and guiding future development.
In conclusion, while current agentic systems demonstrate impressive autonomous reasoning in narrow, SMILES‑centric contexts, they fall short of serving as true computational partners for peptide therapeutics and realistic drug discovery settings. The paper’s benchmark and analysis provide a clear roadmap for the community: integrate protein‑level models, close the experimental feedback loop, enable learning beyond static inference, democratize access to modest resources, and adopt multi‑objective decision making. Addressing these challenges will transform agentic AI from a curiosity‑driven prototype into a robust, partnerable technology capable of accelerating the discovery of safe, effective, and manufacturable medicines.
Comments & Academic Discussion
Loading comments...
Leave a Comment