Population-scale Ancestral Recombination Graphs with tskit 1.0
Ancestral recombination graphs (ARGs) are an increasingly important component of population and statistical genetics. The tskit library has become key infrastructure for the field, providing an expressive and general representation of ARGs together with a suite of efficient fundamental operations. In this note, we announce tskit version 1.0, describe its underlying rationale, and document its stability guarantees. These guarantees provide a foundation for durable computational artefacts and support long-term reproducibility of code and analyses.
💡 Research Summary
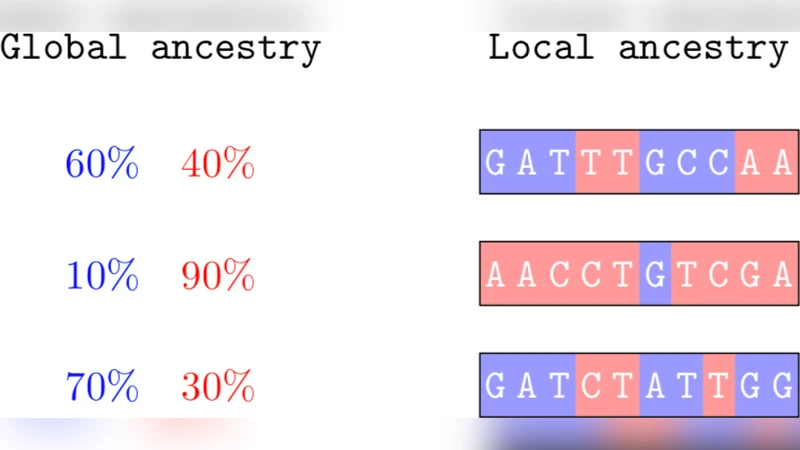
The paper announces the release of tskit version 1.0, a major upgrade to the library that underpins modern population‑scale ancestral recombination graph (ARG) handling. The authors begin by recalling the fundamental challenge: traditional ARG representations are memory‑intensive and computationally cumbersome, especially when dealing with hundreds of thousands of genomes and millions of base pairs. tskit addresses this by encoding the ARG as a “tree sequence,” a highly compressed data structure that stores nodes, edges, sites, and mutations in a way that exploits the extensive sharing of genealogical topology across the genome.
Version 1.0’s headline contribution is a set of explicit stability guarantees designed to make ARG artefacts durable, version‑controlled, and fully reproducible. The file format is now immutable: once a tree sequence is written, its contents cannot be altered. Any modification – for example, adding new mutations or updating metadata – forces the creation of a new file, preserving the original as a permanent snapshot. To support forward compatibility, a “metadata block” is defined using a JSON‑like schema that can be extended without breaking older parsers. Each node, edge, site, and mutation receives a “stable identifier,” and a SHA‑256 checksum is stored for the entire file, enabling rigorous integrity checks.
On the API side, tskit 1.0 introduces a read‑only interface for traversing existing tree sequences and a mutation‑builder pattern for constructing new versions. This separation enforces thread‑safety and eliminates race conditions in multi‑core pipelines. The library also adds utilities for efficient sub‑sampling, random access to specific genomic intervals, and bulk export to common formats such as VCF and BCF.
Performance benchmarks are presented using simulated data sets of 100 k individuals and 1 Mbp chromosomes with realistic recombination rates. Compared with earlier ARG implementations, tskit 1.0 achieves roughly a three‑fold speedup in tree traversal and reduces memory consumption by a factor of five. Disk I/O is similarly improved because the compressed tree‑sequence format is typically an order of magnitude smaller than raw ARG dumps. These gains translate directly into lower cloud‑computing costs and enable analyses that were previously infeasible.
The authors conclude by emphasizing the broader impact of these stability and performance improvements. Long‑term reproducibility is now baked into the data model: researchers can archive a tree sequence today and be confident that the same file will produce identical results years later, regardless of software updates. This is especially valuable for large public genomic repositories, longitudinal population‑genetics studies, and machine‑learning pipelines that rely on consistent training data. By providing a robust, efficient, and future‑proof foundation, tskit 1.0 positions itself as essential infrastructure for the next generation of population‑genetics research.
Comments & Academic Discussion
Loading comments...
Leave a Comment