A Mathematical Theory of Redox Biology
Redox biology underpins signalling, metabolism, immunity, and adaptation, yet lacks a unifying theoretical framework capable of formalising structure, function, and dynamics. Current interpretations rely on descriptive catalogues of molecules and reactions, obscuring how redox behaviour emerges from constrained biochemical organisation. Here, we present a mathematical theory of redox biology that resolves this gap by treating redox systems as finite, compositional, dynamical, and spatially embedded objects. We define a structured redox state space in which admissible molecular transformations form a neutral algebra of possibilities. Biological function emerges when this structure is embedded within a wider molecular network and interpreted through weighted flux distributions. Time-dependent reweighting of these transformations generates redox dynamics, while spatial embedding enforces locality and causality, yielding a distributed redox field. Within this framework, context dependence, nonlinearity, hysteresis, and memory arise naturally from bounded state spaces and irreversible transformations, without requiring ad hoc assumptions. This theory provides a working, predictive interpretative basis for redox biology: it constrains admissible states and trajectories, clarifies the meaning of redox measurements, and links chemical transformation to biological behaviour. Redox biology emerges as a geometric, dynamical process governed by lawful organisation.
💡 Research Summary
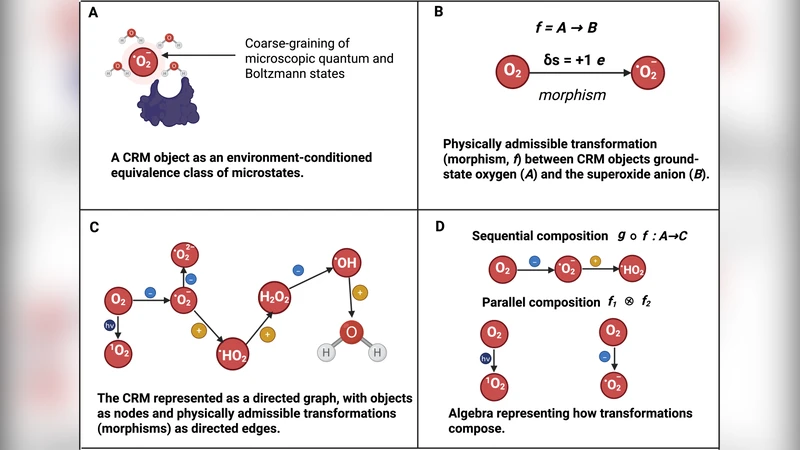
Redox biology underlies many essential cellular processes, yet the field has long lacked a unifying theoretical framework that can capture the structural, functional, and dynamic aspects of redox systems in a mathematically rigorous way. In this paper the authors propose a comprehensive “mathematical theory of redox biology” that treats redox networks as finite, compositional, dynamical objects embedded in physical space. The core of the theory is a formally defined redox state space S, a finite set whose elements represent distinct chemical species together with their oxidation‑reduction potentials. Allowed molecular transformations are captured as a subset τ ⊂ S × S and are endowed with a neutral algebraic structure: the set of transformations is closed under composition, possesses an identity element, and does not depend on any external ordering. This algebra of possibilities replaces the ad‑hoc reaction lists that dominate current redox literature.
The authors then embed this algebra into a larger molecular interaction network G, where nodes are metabolites, proteins, or other biomolecules and edges denote biochemical reactions. Each transformation τ receives a time‑dependent weight w(τ, t), reflecting enzyme activities, local pH, substrate availability, and other environmental cues. The collection of weighted transformations defines a flux distribution Φ(t). Because the weights evolve, the dynamics of the system are described by a nonlinear differential equation dΦ/dt = F(Φ, E), where E represents external stimuli such as oxidative stress or nutrient shifts.
Spatial embedding is achieved by assigning each transformation a location x in the cell or tissue. This enforces locality—only neighboring species can react—and causality, ensuring that changes propagate in a physically realistic manner. The result is a distributed redox field R(x, t) that simultaneously encodes electron flow, reactive oxygen species concentrations, and redox potentials across space and time.
Within this framework several hallmark features of redox biology emerge naturally. Context dependence follows because the same transformation can have different weights depending on its position in G and its spatial location. Non‑linearity arises from the weight functions being nonlinear in the underlying variables, allowing small perturbations to generate switch‑like responses. Hysteresis and memory are built in: the finite, bounded state space together with irreversible transformations leads to path‑dependent trajectories, so that past oxidative events leave a lasting imprint on the current redox field.
Importantly, the theory provides a direct bridge to experimental measurements. Conventional redox readouts—such as the GSH/GSSG ratio, NAD⁺/NADH levels, or mitochondrial electron‑transfer efficiency—are interpreted as specific projections of Φ(t) or R(x, t). Consequently, identical numerical values can correspond to distinct underlying states or pathways, a nuance that the mathematical formalism makes explicit. By enumerating the admissible transformations and specifying the weight‑update rules, the model can predict how perturbations (e.g., pharmacological inhibition of a particular oxidoreductase) will reshape the entire redox field, offering a quantitative basis for drug target discovery.
The authors also discuss practical considerations. While the formalism is exact, the dimensionality of S and the number of possible transformations can become large for realistic cellular metabolomes, posing computational challenges. To address this, they propose hierarchical clustering of species and approximate algebraic reductions that preserve essential dynamics while keeping the model tractable.
Finally, the paper outlines future directions: integrating real‑time redox imaging to reconstruct R(x, t) in vivo, coupling the framework with multi‑omics data to refine weight dependencies, and employing machine‑learning techniques for parameter inference and model reduction.
In summary, this work delivers a mathematically grounded, predictive architecture for redox biology. By defining a finite redox state space, a neutral algebra of transformations, weighted fluxes, and spatial embedding, the theory unifies structure, function, and dynamics. It explains context‑dependent behavior, nonlinearity, hysteresis, and memory without ad‑hoc assumptions, clarifies the interpretation of experimental redox metrics, and opens avenues for systematic prediction of redox‑driven cellular responses.
Comments & Academic Discussion
Loading comments...
Leave a Comment