Oxi-Shapes: Tropical geometric analysis of bounded redox proteomic state spaces
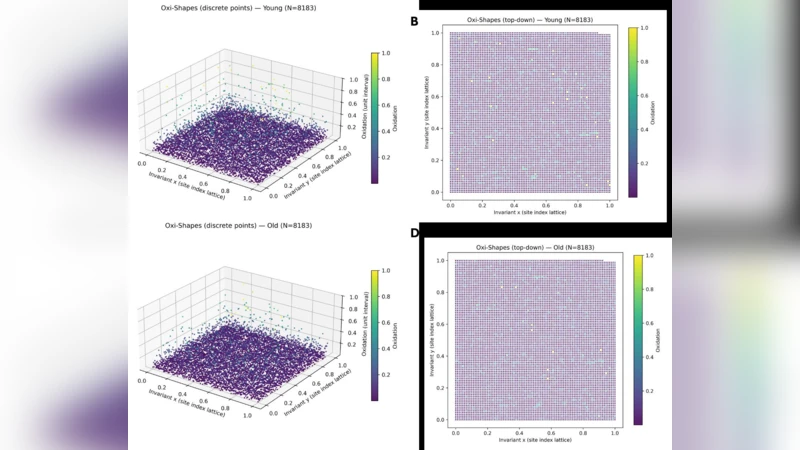
Redox proteomics generates bounded biochemical measurements that are categorically mismatched to conventional linear algebraic formalisms. This work introduces Oxi-Shapes, a tropical geometric framework for the measurement-native analysis of bounded redox proteomic data. Oxi-Shapes represents cysteine oxidation as a scalar field over a discrete lattice, enabling global and site-wise analysis without rescaling, interpolation, or kinetic assumptions. At the global level, the framework yields internal redox entropy, lattice curvature, and derived energy functionals that characterise the geometric structure of the redox proteome. At the site level, Oxi-Shapes defines a bounded change space that makes explicit hard geometric constraints on admissible redox transitions and enables a normalised signed representation of site-wise change as a fraction of available redox freedom. Applied to an ageing mouse brain dataset, Oxi-Shapes reveals that a small decrease in mean oxidation arises from a profound redistribution of site-wise redox states, with thousands of residues shifting toward the reduced absorbing boundary. These results demonstrate that categorically correct algebraic representations expose structure in proteomic data that is inaccessible to mean-centric or unbounded analyses.
💡 Research Summary
The paper introduces Oxi‑Shapes, a novel analytical framework that treats bounded redox proteomics data with tropical geometry rather than conventional linear algebra. Redox proteomics measures the oxidation state of cysteine residues on a scale that is intrinsically limited (typically 0 = fully reduced, 1 = fully oxidized). Traditional pipelines force these measurements into unbounded spaces by normalising, log‑transforming, or interpolating, thereby discarding the hard bounds that encode important biological constraints. Oxi‑Shapes preserves these limits by mapping each cysteine to a scalar field φ defined on a discrete lattice; the lattice axes correspond to sequence position and functional grouping (e.g., domains). In the tropical (max‑plus) algebra, addition becomes the maximum operator and multiplication becomes ordinary addition, which naturally respects the bounded interval.
Global Geometry
Three global descriptors are derived from the lattice field:
- Redox Entropy (S) – a tropical analogue of Shannon entropy computed from the distribution of φ values. It quantifies the diversity of oxidation states across the proteome.
- Lattice Curvature (K) – the range of the discrete Laplacian Δφ across the lattice, highlighting regions where oxidation changes abruptly.
- Energy Functional (E) – the tropical sum of weighted absolute differences between neighboring φ values, measuring the total “cost” of oxidation heterogeneity.
These descriptors provide a holistic view of the proteome’s redox architecture, revealing whether the system is ordered, highly variable, or energetically strained.
Site‑wise Bounded Change Space (BCS)
For each residue the authors define a bounded change interval Δφ ∈
Comments & Academic Discussion
Loading comments...
Leave a Comment