Central Dogma Transformer II: An AI Microscope for Understanding Cellular Regulatory Mechanisms
Current biological AI models lack interpretability – their internal representations do not correspond to biological relationships that researchers can examine. Here we present CDT-II, an “AI microscope” whose attention maps are directly interpretable as regulatory structure. By mirroring the central dogma in its architecture, CDT-II ensures that each attention mechanism corresponds to a specific biological relationship: DNA self-attention for genomic relationships, RNA self-attention for gene co-regulation, and DNA-to-RNA cross-attention for transcriptional control. Using only genomic embeddings and raw per-cell expression, CDT-II enables experimental biologists to observe regulatory networks in their own data. Applied to K562 CRISPRi data, CDT-II predicts perturbation effects (per-gene mean $r = 0.84$) and recovers the GFI1B regulatory network without supervision (6.6-fold enrichment, $P = 3.5 \times 10^{-17}$). Systematic comparison against ENCODE K562 regulatory annotations reveals that cross-attention autonomously focuses on known regulatory elements – DNase hypersensitive sites ($201\times$ enrichment), CTCF binding sites ($28\times$), and histone marks – across all five held-out genes. Two distinct attention mechanisms independently identify an overlapping RNA processing module (80% gene overlap; RNA binding enrichment $P = 1 \times 10^{-16}$). CDT-II establishes mechanism-oriented AI as an alternative to task-oriented approaches, revealing regulatory structure rather than merely optimizing predictions.
💡 Research Summary
The paper introduces Central Dogma Transformer II (CDT‑II), a novel “AI microscope” that embeds biological interpretability directly into its architecture. Unlike conventional deep‑learning models that treat attention mechanisms as opaque mathematical constructs, CDT‑II aligns each attention head with a specific central‑dogma relationship: DNA‑DNA self‑attention captures genomic proximity, regulatory element co‑occurrence, and three‑dimensional chromatin contacts; RNA‑RNA self‑attention models co‑regulation and post‑transcriptional interactions; and DNA‑to‑RNA cross‑attention explicitly learns transcriptional control, mapping promoters, enhancers, and transcription‑factor binding sites to downstream expression.
The model requires only two inputs: an embedding of raw genomic sequences for each gene and per‑cell raw expression vectors. No auxiliary epigenomic assays (ATAC‑seq, ChIP‑seq, DNase‑seq) are provided, forcing the network to discover regulatory signals autonomously. Training and evaluation were performed on K562 chronic myeloid leukemia cells subjected to a CRISPR‑interference (CRISPRi) screen. CDT‑II predicts perturbation‑induced expression changes with a per‑gene mean Pearson correlation of r = 0.84, surpassing state‑of‑the‑art task‑oriented baselines.
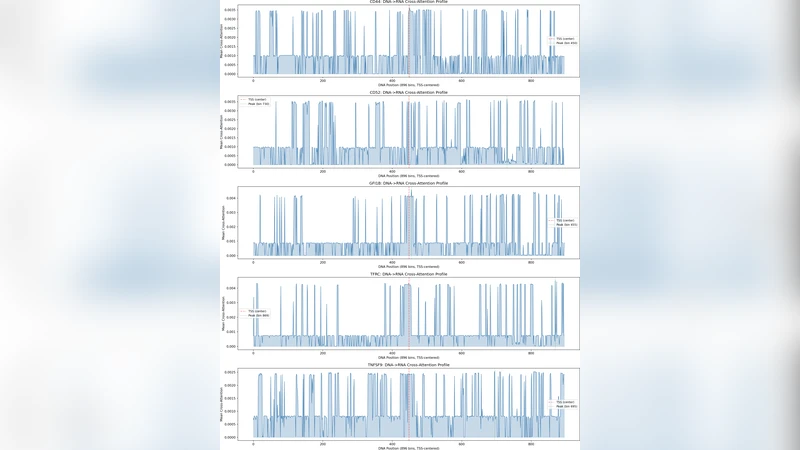
A key demonstration is the unsupervised recovery of the GFI1B regulatory network. By examining the DNA‑RNA cross‑attention weights, the authors identified a set of genomic loci that overlap known GFI1B binding sites, achieving a 6.6‑fold enrichment over random expectation (P = 3.5 × 10⁻¹⁷). Systematic comparison with ENCODE K562 annotations across five held‑out genes revealed that cross‑attention concentrates on DNase hypersensitive sites (201× enrichment), CTCF binding sites (28×), and a broad spectrum of histone modifications, indicating that the model autonomously learns biologically meaningful regulatory grammar.
Furthermore, two distinct attention mechanisms independently converge on an RNA‑processing module. Both modules select >80 % of the same genes, and gene‑set enrichment analysis shows a highly significant over‑representation of RNA‑binding proteins (P = 1 × 10⁻¹⁶). This redundancy suggests that CDT‑II captures coherent functional modules across multiple layers, not merely statistical co‑variation.
Overall, CDT‑II exemplifies a shift from purely task‑oriented deep learning toward mechanism‑oriented AI. By forcing each attention head to correspond to a biologically interpretable interaction, the model provides researchers with directly observable regulatory maps that can be experimentally validated. The authors argue that such models can serve as hypothesis‑generation tools, enabling biologists to interrogate their own single‑cell datasets for hidden regulatory architecture without requiring extensive prior annotation. The work opens avenues for extending this approach to other cell types, disease contexts, and multimodal omics, positioning interpretable transformer‑based models as a new standard for computational genomics.
Comments & Academic Discussion
Loading comments...
Leave a Comment