A Survey on Medical Image Compression: From Traditional to Learning-Based Approaches
The exponential growth of medical imaging has created significant challenges in data storage, transmission, and management for healthcare systems. In this vein, efficient compression becomes increasingly important. Unlike natural image compression, medical image compression prioritizes preserving diagnostic details and structural integrity, imposing stricter quality requirements and demanding fast, memory-efficient algorithms that balance computational complexity with clinically acceptable reconstruction quality. Meanwhile, the medical imaging family includes a plethora of modalities, each possessing different requirements. For example, 2D medical image (e.g., X-rays, histopathological images) compression focuses on exploiting intra-slice spatial redundancy, while volumetric medical image faces require handling intra-slice and inter-slice spatial correlations, and 4D dynamic imaging (e.g., time-series CT/MRI, 4D ultrasound) additionally demands processing temporal correlations between consecutive time frames. Traditional compression methods, grounded in mathematical transforms and information theory principles, provide solid theoretical foundations, predictable performance, and high standardization levels, with extensive validation in clinical environments. In contrast, deep learning-based approaches demonstrate remarkable adaptive learning capabilities and can capture complex statistical characteristics and semantic information within medical images. This comprehensive survey establishes a two-facet taxonomy based on data structure (2D vs 3D/4D) and technical approaches (traditional vs learning-based), thereby systematically presenting the complete technological evolution, analyzing the unique technical challenges, and prospecting future directions in medical image compression.
💡 Research Summary
**
The paper presents a comprehensive survey of medical image compression, organized around two fundamental dimensions: data structure (2‑D versus 3‑D/4‑D) and methodological approach (traditional signal‑processing techniques versus learning‑based methods). It begins by highlighting the unique constraints of medical imaging—strict diagnostic‑fidelity requirements, DICOM standards, high bit‑depth, and the need for ROI‑aware processing—distinguishing it from natural‑image compression where perceptual loss is often acceptable.
Traditional methods are reviewed in depth, covering transform‑based codecs (DCT, wavelet, JPEG‑2000), predictive coding schemes (lossless 3‑D codecs such as JP3D, Med‑CAE), and entropy coding (Huffman, arithmetic). These approaches offer solid theoretical guarantees, deterministic behavior, and widespread clinical adoption, but they struggle to capture complex anatomical structures and inter‑slice or temporal redundancies inherent in volumetric data.
The survey then shifts to learning‑based techniques, categorizing them into three streams: (1) auto‑encoder, variational auto‑encoder, and generative adversarial network models that learn compact latent representations; (2) deep neural networks (2‑D, 3‑D, and 4‑D CNNs, RNN/LSTM, transformers) that directly predict bitstreams or improve entropy models; and (3) implicit neural representations (INR) that encode an image as a continuous function, enabling ultra‑low‑bit storage while preserving reconstruction quality. Empirical evidence shows that these methods often surpass traditional codecs in rate‑distortion performance, especially when semantic information is important. However, they demand large annotated datasets, high computational resources, and face regulatory hurdles due to limited interpretability.
Hybrid solutions are identified as a promising direction, combining the reliability of classical pipelines with the adaptability of neural networks—for example, inserting a neural entropy coder after a DCT stage or using deep models for ROI‑aware bit allocation.
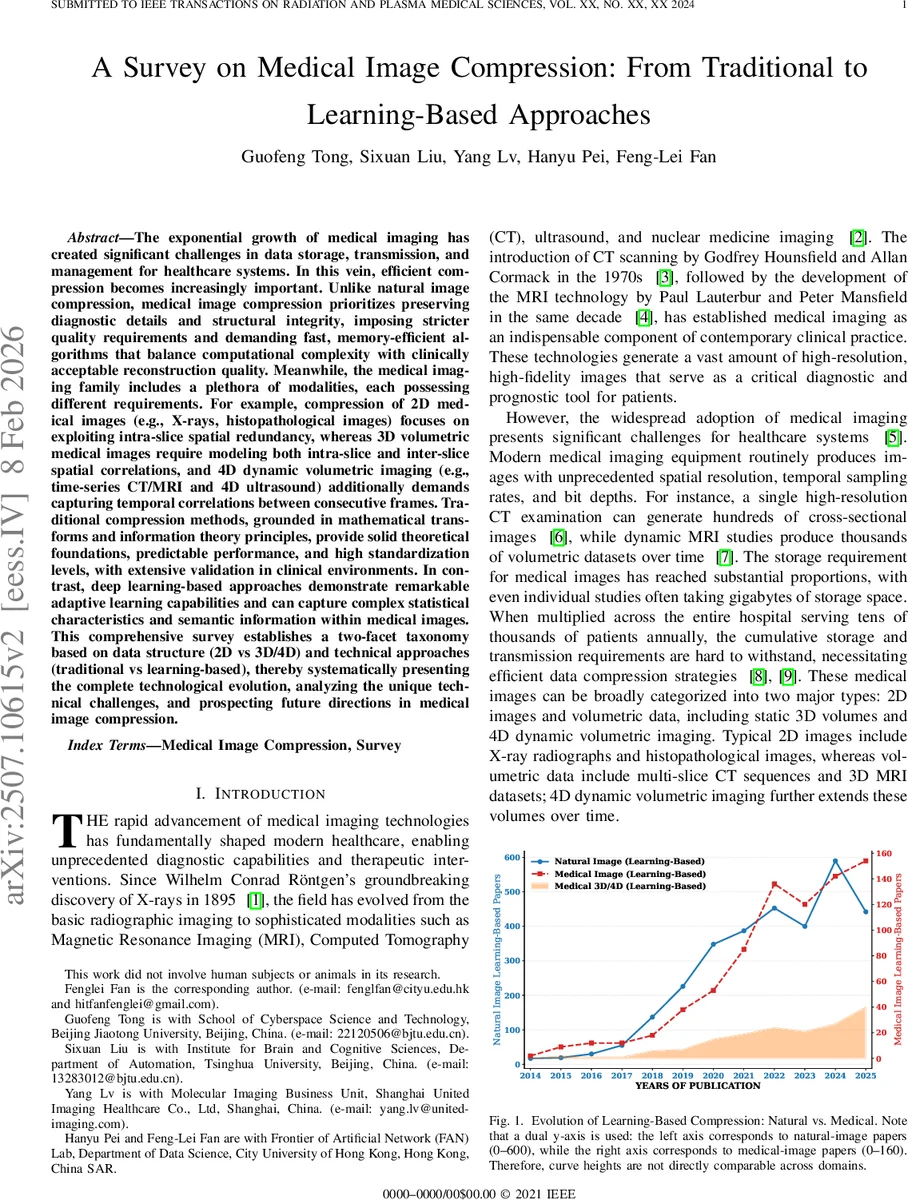
A bibliometric analysis reveals a rapid increase in learning‑based medical image compression publications from 2014 to 2025, with 3‑D/4‑D works doubling between 2023 and 2025. The authors note a 2‑3‑year lag between advances in natural‑image compression and their adoption in the medical domain, underscoring the need for domain‑specific adaptation.
The paper’s contributions are threefold: (1) a unified two‑axis taxonomy that aligns historical milestones with methodological categories; (2) an exhaustive technical coverage that bridges classical transform/predictive coding, modern deep and generative models, hybrid schemes, and INR‑based techniques; and (3) a focused discussion of medical‑specific constraints such as diagnostic fidelity, DICOM compliance, and clinical workflow integration. By providing this holistic perspective, the survey serves as a roadmap for future research aiming to develop compression algorithms that maintain clinical image quality while achieving substantial storage and transmission savings.
Comments & Academic Discussion
Loading comments...
Leave a Comment