LmPT: Conditional Point Transformer for Anatomical Landmark Detection on 3D Point Clouds
Accurate identification of anatomical landmarks is crucial for various medical applications. Traditional manual landmarking is time-consuming and prone to inter-observer variability, while rule-based methods are often tailored to specific geometries or limited sets of landmarks. In recent years, anatomical surfaces have been effectively represented as point clouds, which are lightweight structures composed of spatial coordinates. Following this strategy and to overcome the limitations of existing landmarking techniques, we propose Landmark Point Transformer (LmPT), a method for automatic anatomical landmark detection on point clouds that can leverage homologous bones from different species for translational research. The LmPT model incorporates a conditioning mechanism that enables adaptability to different input types to conduct cross-species learning. We focus the evaluation of our approach on femoral landmarking using both human and newly annotated dog femurs, demonstrating its generalization and effectiveness across species. The code and dog femur dataset will be publicly available at: https://github.com/Pierreoo/LandmarkPointTransformer.
💡 Research Summary
The paper introduces Landmark Point Transformer (LmPT), a novel conditional deep‑learning architecture designed to automatically detect anatomical landmarks on 3‑dimensional point‑cloud representations of bones. Traditional manual landmarking is labor‑intensive and suffers from inter‑observer variability, while rule‑based algorithms are often limited to specific geometries or a small set of landmarks. Recent advances have shown that point clouds are an efficient way to model anatomical surfaces, but few methods exploit modern transformer‑based networks for this task, especially across species.
LmPT builds on the Point Transformer (PT) family, which uses self‑attention on local neighborhoods to capture complex geometric relationships. Two PT backbones are evaluated: PTv2, which relies on k‑nearest‑neighbor (k‑NN) attention windows and a classic encoder‑decoder with pooling/unpooling, and PTv3, which replaces neighborhood queries with a space‑filling curve serialization for higher computational efficiency. The key innovation is the integration of a Feature‑wise Linear Modulation (FiLM) conditioning module. A small embedding that encodes the “input type” (e.g., human femur vs. dog femur) is passed through a linear layer to generate per‑channel scale and shift parameters. These FiLM parameters are applied only to the bottleneck features of the encoder, allowing the same network to adapt its internal representations to different anatomical morphologies without increasing the overall parameter count.
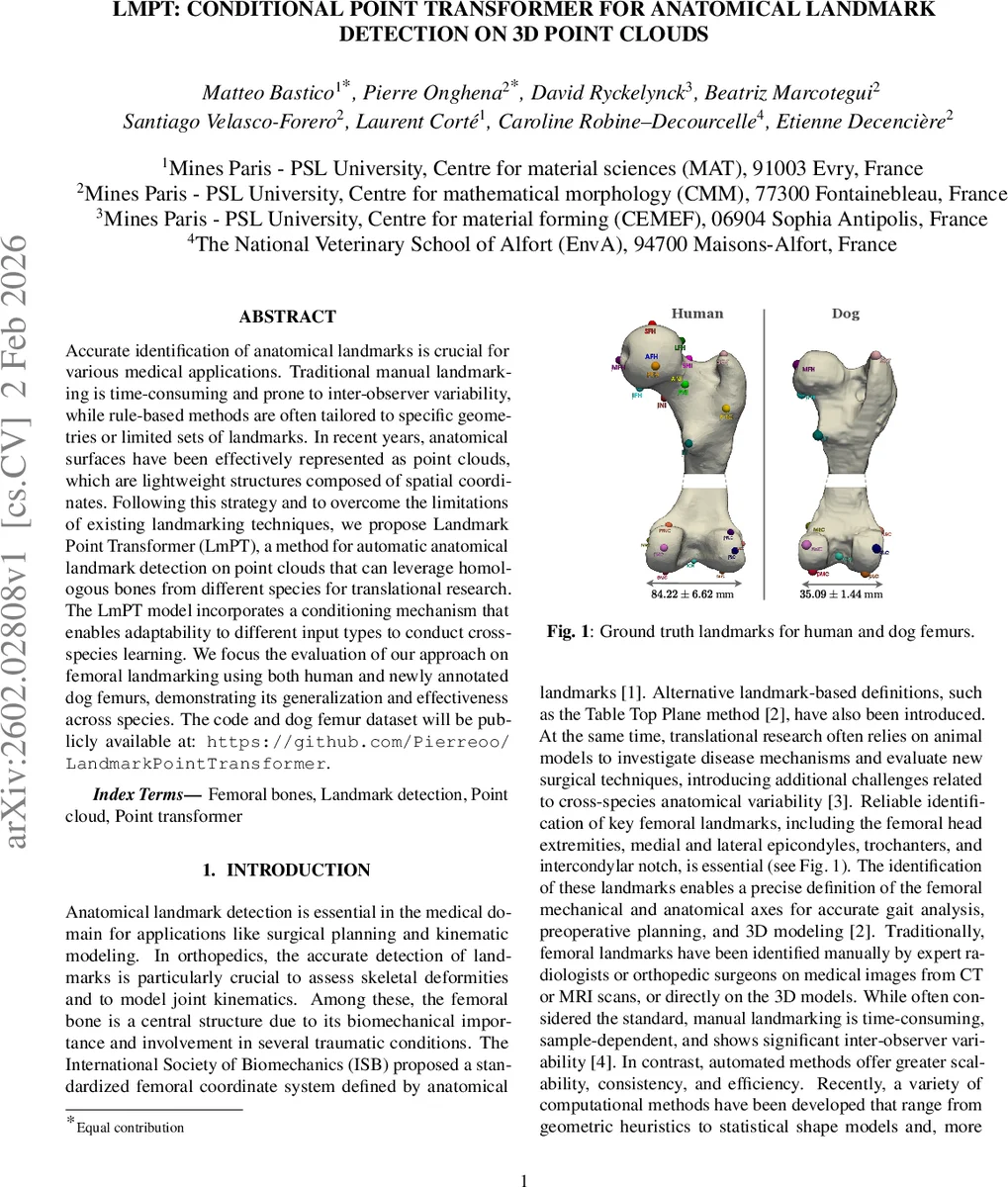
The authors assembled two datasets. The human dataset is a publicly available collection of 20 femur meshes (10 left, 10 right) annotated with 22 landmarks by five experts, each performing four repetitions; the medoid of the 20 annotations per landmark serves as ground truth. The dog dataset is newly created for this work, comprising 14 femur meshes (7 left, 7 right) from various breeds, annotated by a veterinary specialist with 11 landmarks that largely correspond to a subset of the human set. For cross‑species experiments the two datasets are merged, yielding 26 training and 8 test samples. All point clouds are uniformly down‑sampled to 8,192 points, normalized, and augmented with random rotations, scaling, and left‑right flipping to improve robustness to side symmetry.
Training is performed for 500 epochs with a batch size of 4, using a channel‑wise cross‑entropy loss that ignores unlabeled points, the AdamW optimizer (learning rate = 3 × 10⁻⁴), and a one‑cycle learning‑rate scheduler. Evaluation metrics are mean absolute error (MAE) in millimetres and percentage of correct keypoints (PCK) computed over distance thresholds from 1 mm to 8 mm.
Results on the human femur test set show that LmPT‑v2 (PTv2 backbone) achieves an average MAE of 1.58 mm, substantially lower than the average manual error (AME = 4.10 mm) and outperforming the prior A&A atlas‑based method. The PTv3 variant performs worse (average MAE ≈ 3.4 mm), indicating that the precise k‑NN attention of PTv2 is more suitable for fine‑grained landmark localisation. PCK curves confirm that LmPT‑v2 reaches >80 % correct keypoints already at a 1 mm threshold, whereas PTv3 lags behind.
On the dog femur test set, LmPT‑v2 again leads with an average MAE of 1.23 mm, while the A&A method fails entirely because it is tailored to human anatomy. PTv3 again shows higher error (≈ 2.85 mm). The PCK analysis mirrors the human results, with LmPT‑v2 achieving near‑perfect scores at low thresholds.
Cross‑species training (human + dog) demonstrates the benefit of the FiLM conditioning. For human femurs, the MAE drops marginally (≈ 0.1 mm) compared to single‑species training, suggesting that the additional dog data provides useful geometric priors. For dog femurs, the MAE rises slightly (≈ 0.1 mm), likely because the dog set contains fewer landmarks and includes only a subset of the human ones; the extra human‑only landmarks may introduce irrelevant features. Nevertheless, the PCK curves for both species improve, reaching perfect scores at lower distance thresholds, confirming that the model successfully leverages homologous structures across species.
The authors discuss several limitations. The datasets are small, which restricts statistical power and may not capture the full variability of femoral anatomy across populations and breeds. PTv3’s efficiency gains come at the cost of reduced localisation precision, indicating that future work should aim to combine fast serialization with accurate neighborhood attention. Moreover, the current study focuses exclusively on femurs; extending the approach to other bones, joint surfaces, or whole‑body scans would test the generality of the conditional transformer concept.
In conclusion, LmPT introduces a conditional point‑transformer architecture that achieves state‑of‑the‑art performance for anatomical landmark detection on 3‑D point clouds. By incorporating FiLM‑based conditioning, the model can learn from heterogeneous, cross‑species data, opening the door to translational research that bridges human and animal models. The release of the annotated dog femur dataset and the open‑source code further supports reproducibility and future extensions in medical imaging and computational anatomy.
Comments & Academic Discussion
Loading comments...
Leave a Comment