De Novo Molecular Generation from Mass Spectra via Many-Body Enhanced Diffusion
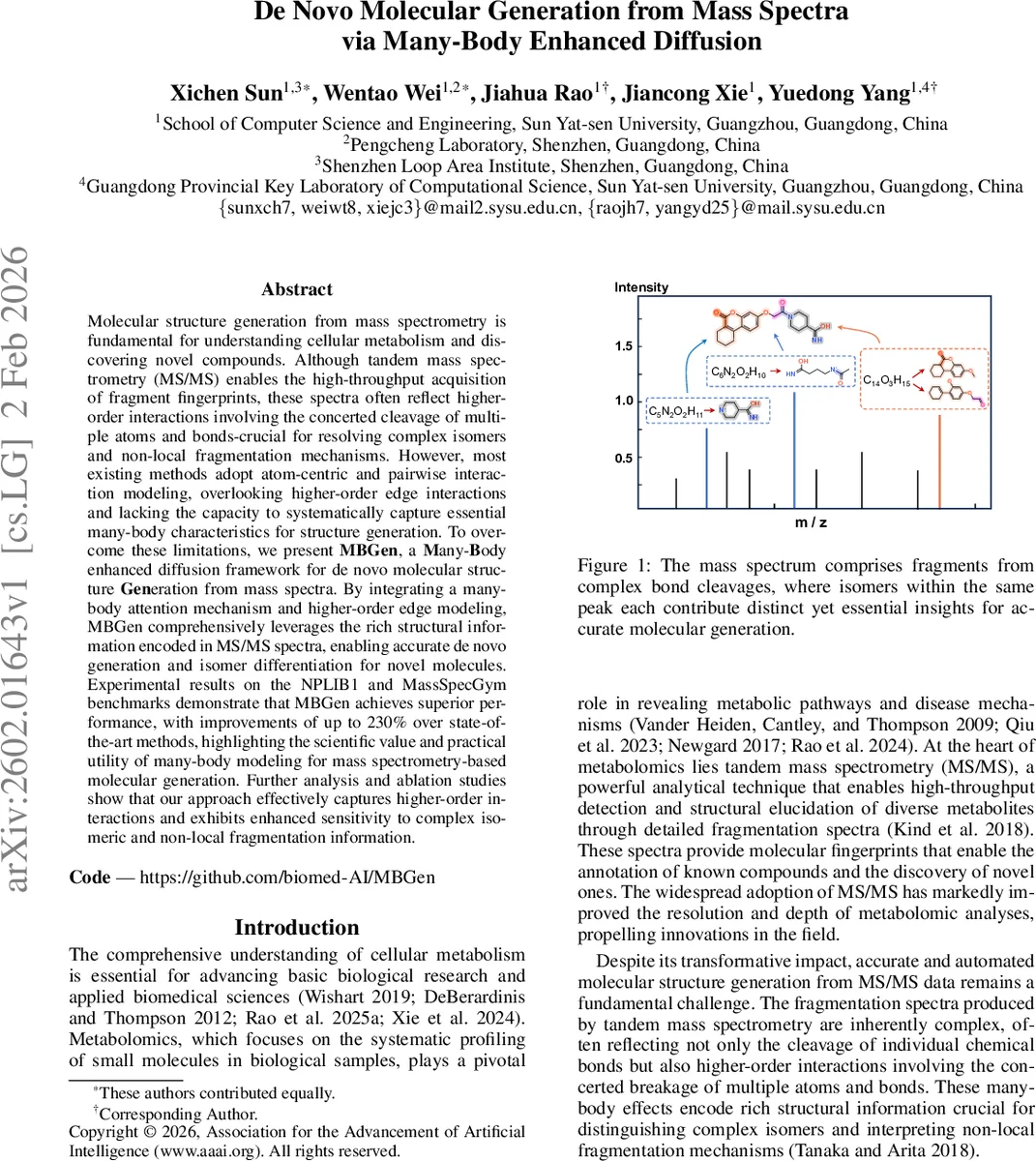
Molecular structure generation from mass spectrometry is fundamental for understanding cellular metabolism and discovering novel compounds. Although tandem mass spectrometry (MS/MS) enables the high-throughput acquisition of fragment fingerprints, these spectra often reflect higher-order interactions involving the concerted cleavage of multiple atoms and bonds-crucial for resolving complex isomers and non-local fragmentation mechanisms. However, most existing methods adopt atom-centric and pairwise interaction modeling, overlooking higher-order edge interactions and lacking the capacity to systematically capture essential many-body characteristics for structure generation. To overcome these limitations, we present MBGen, a Many-Body enhanced diffusion framework for de novo molecular structure Generation from mass spectra. By integrating a many-body attention mechanism and higher-order edge modeling, MBGen comprehensively leverages the rich structural information encoded in MS/MS spectra, enabling accurate de novo generation and isomer differentiation for novel molecules. Experimental results on the NPLIB1 and MassSpecGym benchmarks demonstrate that MBGen achieves superior performance, with improvements of up to 230% over state-of-the-art methods, highlighting the scientific value and practical utility of many-body modeling for mass spectrometry-based molecular generation. Further analysis and ablation studies show that our approach effectively captures higher-order interactions and exhibits enhanced sensitivity to complex isomeric and non-local fragmentation information.
💡 Research Summary
The paper introduces MBGen, a novel framework for de‑novo molecular structure generation conditioned on tandem mass spectrometry (MS/MS) data. Recognizing that MS/MS spectra encode not only single‑bond cleavages but also concerted, higher‑order fragmentation events involving multiple atoms and bonds, the authors argue that existing atom‑centric and pairwise graph models are insufficient for capturing this rich information. MBGen addresses these shortcomings through two main innovations: (1) an edge‑centric diffusion decoder that treats chemical bonds as the primary modeling units, and (2) a many‑body attention module that explicitly models interactions among triplets (and potentially higher‑order groups) of edges.
The overall pipeline consists of three stages. First, a spectrum encoder based on the pretrained MIST Formula Transformer converts a set of (m/z, intensity) peaks, each annotated with a molecular formula via SIRIUS, into a global fingerprint vector y using a Set‑Transformer with pairwise attention. Second, the graph decoder initializes node features from atomic types and constructs initial edge embeddings from pairs of node features and their relational descriptors. Edge embeddings are iteratively refined through node‑edge interaction layers that incorporate the global fingerprint via FiLM modulation and attention‑weighted aggregation of node and edge information. Third, the many‑body attention module updates each edge embedding by aggregating information from neighboring edge pairs, effectively allowing information to flow across triplets (i, j, k) without passing through the central node j. This design mitigates the bottleneck of traditional message‑passing and enriches the representation with higher‑order chemical context.
Training proceeds in three steps: (i) pretraining the spectrum encoder, (ii) pretraining the diffusion decoder on synthetic graphs, and (iii) end‑to‑end fine‑tuning on paired spectra‑graph data. The diffusion process progressively denoises a randomly initialized adjacency tensor, guided at each step by the edge‑centric message passing and many‑body attention, until a chemically plausible molecular graph is produced.
Experimental evaluation on two benchmarks—NPLIB1 (synthetic compounds) and MassSpecGym (real metabolomics spectra)—shows that MBGen substantially outperforms state‑of‑the‑art methods such as MADGEN and DiffMS. Reported gains include up to 230 % improvement in top‑k accuracy and markedly higher rates of correctly distinguishing structural isomers that generate very similar spectra. Ablation studies confirm that both the edge‑centric formulation and the many‑body attention are essential: removing either component degrades performance, especially in isomer discrimination. Visualization of attention weights reveals that the model assigns high importance to chemically meaningful triplets (e.g., C‑O‑H, N‑C‑C), aligning with known fragmentation pathways.
The authors acknowledge limitations: the approach relies on high‑resolution spectra and accurate formula annotation (via SIRIUS), making it less robust to low‑resolution or highly noisy data. The many‑body attention incurs cubic computational complexity with respect to the number of atoms, which may hinder scalability to very large molecules. Future work is suggested on improving efficiency, extending to lower‑quality spectra, and integrating uncertainty estimation for formula annotation.
In summary, MBGen demonstrates that incorporating many‑body interactions and an edge‑centric perspective into diffusion‑based molecular generation can dramatically enhance the exploitation of MS/MS data, yielding more accurate structure predictions and superior isomer resolution. This advance holds promise for metabolite identification, novel drug discovery, and broader applications where de‑novo elucidation of unknown compounds from mass spectra is required.
Comments & Academic Discussion
Loading comments...
Leave a Comment