Learning to Look Closer: A New Instance-Wise Loss for Small Cerebral Lesion Segmentation
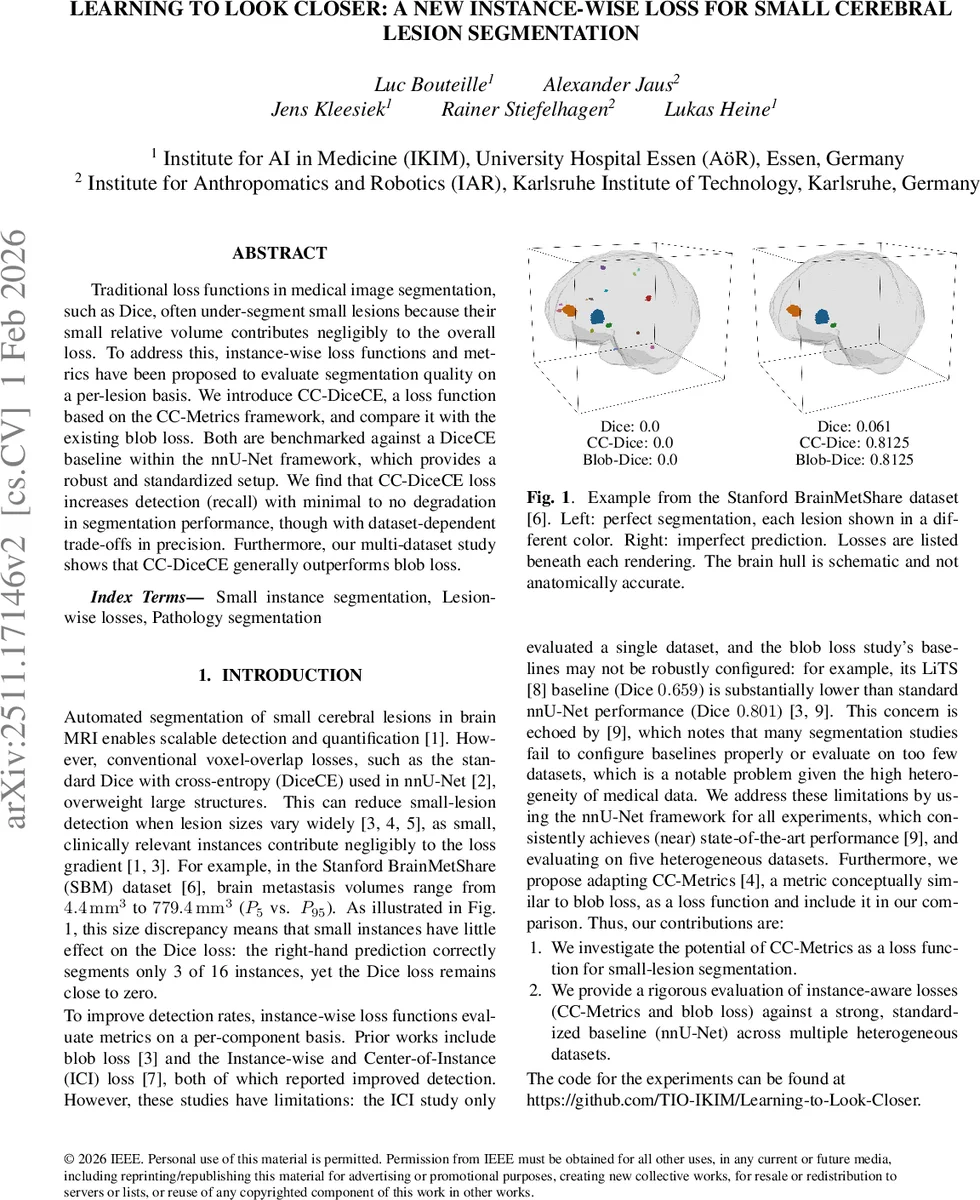
Traditional loss functions in medical image segmentation, such as Dice, often under-segment small lesions because their small relative volume contributes negligibly to the overall loss. To address this, instance-wise loss functions and metrics have been proposed to evaluate segmentation quality on a per-lesion basis. We introduce CC-DiceCE, a loss function based on the CC-Metrics framework, and compare it with the existing blob loss. Both are benchmarked against a DiceCE baseline within the nnU-Net framework, which provides a robust and standardized setup. We find that CC-DiceCE loss increases detection (recall) with minimal to no degradation in segmentation performance, though with dataset-dependent trade-offs in precision. Furthermore, our multi-dataset study shows that CC-DiceCE generally outperforms blob loss.
💡 Research Summary
The paper addresses a well‑known limitation of conventional voxel‑overlap loss functions such as Dice (often combined with cross‑entropy, DiceCE) in medical image segmentation: small lesions contribute negligibly to the loss because the loss is dominated by larger structures. Consequently, neural networks trained with DiceCE tend to under‑segment or completely miss tiny pathological entities, which is clinically unacceptable for many brain pathologies (e.g., metastases, lacunes, microbleeds).
To mitigate this, the authors investigate instance‑wise loss functions that evaluate each connected component (lesion) separately. They focus on two approaches: the previously proposed “blob loss” and a novel adaptation of the CC‑Metrics framework, which they call CC‑DiceCE. CC‑Metrics computes a Voronoi partition of the image based on the ground‑truth components; each Voronoi region is then scored with a base metric (here DiceCE). By averaging over all components, every lesion receives equal weight regardless of size, and missing a lesion incurs a maximal penalty (1/N contribution set to the worst possible value). False positives affect only the Voronoi region they occupy, limiting their impact on the overall score.
All experiments are performed within the highly standardized nnU‑Net pipeline, retaining default hyper‑parameters except for using the non‑smooth Dice variant (ε = 0) to avoid instability on the CMB dataset. The instance‑wise losses are combined 1:1 with the global DiceCE loss, yielding Blob‑DiceCE and CC‑DiceCE objectives. Five heterogeneous brain MRI datasets are used: Lacunes (LAC), Cerebral Microbleeds (CMB), Stanford BrainMetShare (SBM), White Matter Hyperintensities (WMH), and the BraTS glioma challenge. These datasets differ markedly in lesion size distribution, multiplicity, and imaging modalities, providing a robust testbed for the proposed loss.
Evaluation metrics include global Dice, CC‑Dice (the instance‑wise Dice version of CC‑Metrics), instance‑wise F1, and recall stratified by lesion‑volume quartiles. Five‑fold cross‑validation is applied, and results are reported as mean ± standard deviation across folds.
Key findings:
-
Global Dice stability – CC‑DiceCE maintains Dice scores within the typical 5‑fold variation for all datasets, with the worst observed drop of only –0.011. In contrast, Blob‑DiceCE reduces Dice noticeably on LAC and WMH, confirming earlier reports of Dice degradation when using blob loss.
-
Improved detection – CC‑DiceCE consistently raises CC‑Dice and recall in four of the five datasets (LAC, CMB, SBM, BraTS). The increase is not confined to the smallest lesions; recall gains are distributed across all volume quartiles, indicating that the loss acts as a general instance‑regularizer rather than a specialized small‑object detector.
-
Precision trade‑offs – Precision remains largely unchanged, with a modest decline only on BraTS where large tumor masses dominate and small peripheral annotation artifacts become false positives. This aligns with the theoretical expectation that false positives only penalize the Voronoi region they occupy.
-
Comparison with blob loss – CC‑DiceCE outperforms Blob‑DiceCE in 22 out of 25 metric‑dataset combinations. Blob‑DiceCE often yields larger Dice reductions while delivering comparable or inferior instance‑wise metrics.
-
Effect of training schedule – On WMH, a shorter 150‑epoch schedule (versus the default 1000 epochs) improves both Dice and CC‑Dice for all three loss functions, suggesting that the default schedule may cause over‑fitting that masks the benefits of instance‑wise losses.
The authors hypothesize that the heightened penalty for missed components drives the network to be more aggressive in lesion detection, which is clinically advantageous when the cost of a false negative outweighs that of a false positive. They also note that the loss’s simplicity (a straightforward combination of two well‑understood terms) makes it easy to integrate into existing pipelines without extensive hyper‑parameter tuning.
In conclusion, the study demonstrates that adapting CC‑Metrics into a loss function (CC‑DiceCE) provides a practical, effective means to boost small‑lesion detection while preserving overall segmentation quality. The approach outperforms the previously introduced blob loss across diverse brain MRI datasets and offers a balanced trade‑off suitable for clinical deployment, where missing a subtle pathology is often far more detrimental than flagging an extra candidate for radiologist review. Future work is suggested to extend the method to other organs, imaging modalities, and to explore lightweight implementations for real‑time clinical use.
Comments & Academic Discussion
Loading comments...
Leave a Comment