Rethinking Drug-Drug Interaction Modeling as Generalizable Relation Learning
Drug-drug interaction (DDI) prediction is central to drug discovery and clinical development, particularly in the context of increasingly prevalent polypharmacy. Although existing computational methods achieve strong performance on standard benchmarks, they often fail to generalize to realistic deployment scenarios, where most candidate drug pairs involve previously unseen drugs and validated interactions are scarce. We demonstrate that proximity in the embedding spaces of prevailing molecule-centric DDI models does not reliably correspond to interaction labels, and that simply scaling up model capacity therefore fails to improve generalization. To address these limitations, we propose GenRel-DDI, a generalizable relation learning framework that reformulates DDI prediction as a relation-centric learning problem, in which interaction representations are learned independently of drug identities. This relation-level abstraction enables the capture of transferable interaction patterns that generalize to unseen drugs and novel drug pairs. Extensive experiments across multiple benchmark demonstrate that GenRel-DDI consistently and significantly outperforms state-of-the-art methods, with particularly large gains on strict entity-disjoint evaluations, highlighting the effectiveness and practical utility of relation learning for robust DDI prediction. The code is available at https://github.com/SZU-ADDG/GenRel-DDI.
💡 Research Summary
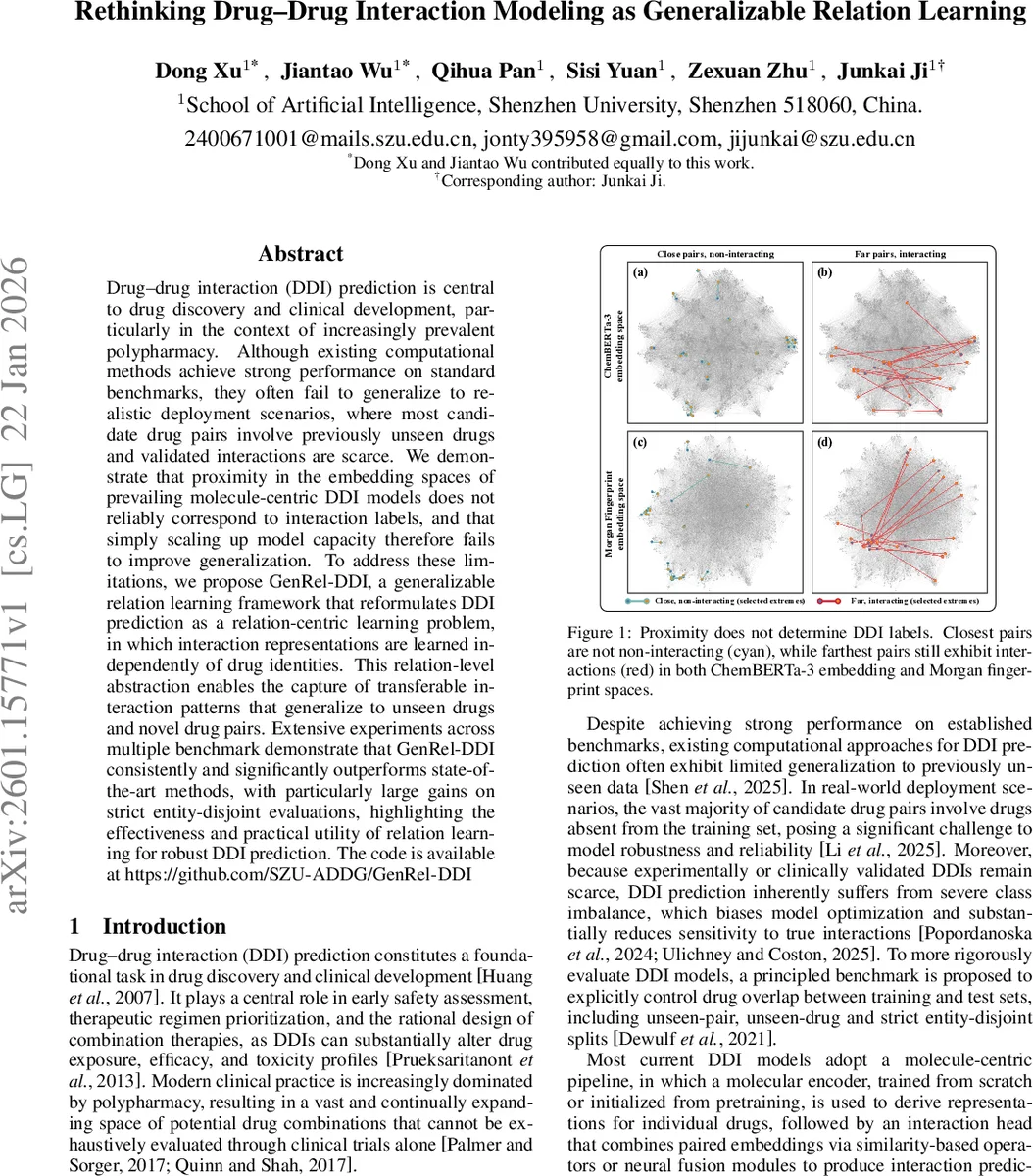
Drug‑drug interaction (DDI) prediction is a cornerstone of modern drug discovery and clinical development, yet current computational approaches struggle to generalize beyond the training data. Most existing methods follow a molecule‑centric paradigm: each drug is encoded independently by a graph neural network, a language model, or another molecular encoder, and a downstream interaction head fuses the two embeddings using similarity measures or neural fusion modules. The authors demonstrate that, in both ChemBERTa‑3 and Morgan fingerprint spaces, proximity of drug embeddings does not reliably correspond to interaction labels—closest pairs can be non‑interacting while farthest pairs may still interact. Consequently, simply increasing model capacity does not improve out‑of‑distribution performance, especially when test pairs involve unseen drugs or when labeled interactions are scarce.
To address these shortcomings, the paper introduces GenRel‑DDI, a generalizable relation‑learning framework that reframes DDI prediction as a relation‑centric task. The key idea is to decouple interaction modeling from drug identities. Multiple pretrained molecular encoders (graph‑based, language‑based, or multimodal) are kept frozen as “anchors.” For each drug, two token streams are generated: a reference stream (fixed) and an adaptive stream (trainable). Within‑drug conditioning uses cross‑attention to let the adaptive stream attend to the reference stream (and vice‑versa), producing a single drug‑level representation that is already conditioned on its partner.
A role‑separated relation trunk then operates on the conditioned token sequences of the two drugs. Cross‑attention between the sequences yields partner‑conditioned interaction factors, which are pooled to form a compact relation vector zₐb. This vector captures the interaction pattern without referencing the specific drug IDs, enabling the same trunk to be reused for any pair, including those containing drugs never seen during training. A task‑specific head maps zₐb to class logits, and the whole system is trained with standard cross‑entropy loss while only the adaptive streams and the relation trunk are updated.
The authors provide a theoretical analysis showing that freezing the anchor streams bounds prediction drift under a Lipschitz assumption, ensuring that the model’s behavior on unseen drugs is controlled. Empirically, GenRel‑DDI is evaluated on three rigorously defined splits: unseen‑pair (both drugs seen but the pair is new), unseen‑drug (exactly one drug unseen), and entity‑disjoint (neither drug seen). Across multiple public DDI benchmarks, GenRel‑DDI consistently outperforms state‑of‑the‑art baselines such as DeepDDI, DeepC, and recent graph‑based models. The most pronounced gains appear on the entity‑disjoint split, where AUROC and AUPR improvements of 5–10 percentage points are reported. These results confirm that learning transferable interaction patterns at the relation level dramatically improves robustness to distribution shift.
Beyond performance, GenRel‑DDI offers practical advantages. By leveraging frozen pretrained encoders, it avoids catastrophic forgetting and reduces the need for extensive feature engineering. The lightweight adapter and relation trunk are highly scalable, allowing easy integration of additional modalities or larger pretrained models. The codebase is publicly released, facilitating reproducibility and adoption in real‑world pipelines.
In summary, the paper makes three major contributions: (1) empirical evidence that scaling up molecule‑centric DDI models does not solve generalization failures; (2) the introduction of a novel relation‑centric framework that decouples interaction learning from drug identities; and (3) extensive experiments demonstrating superior, statistically significant performance across diverse, realistic evaluation settings. GenRel‑DDI thus represents a significant step toward reliable, deployable DDI prediction systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment