Integrating Color Histogram Analysis and Convolutional Neural Network for Skin Lesion Classification
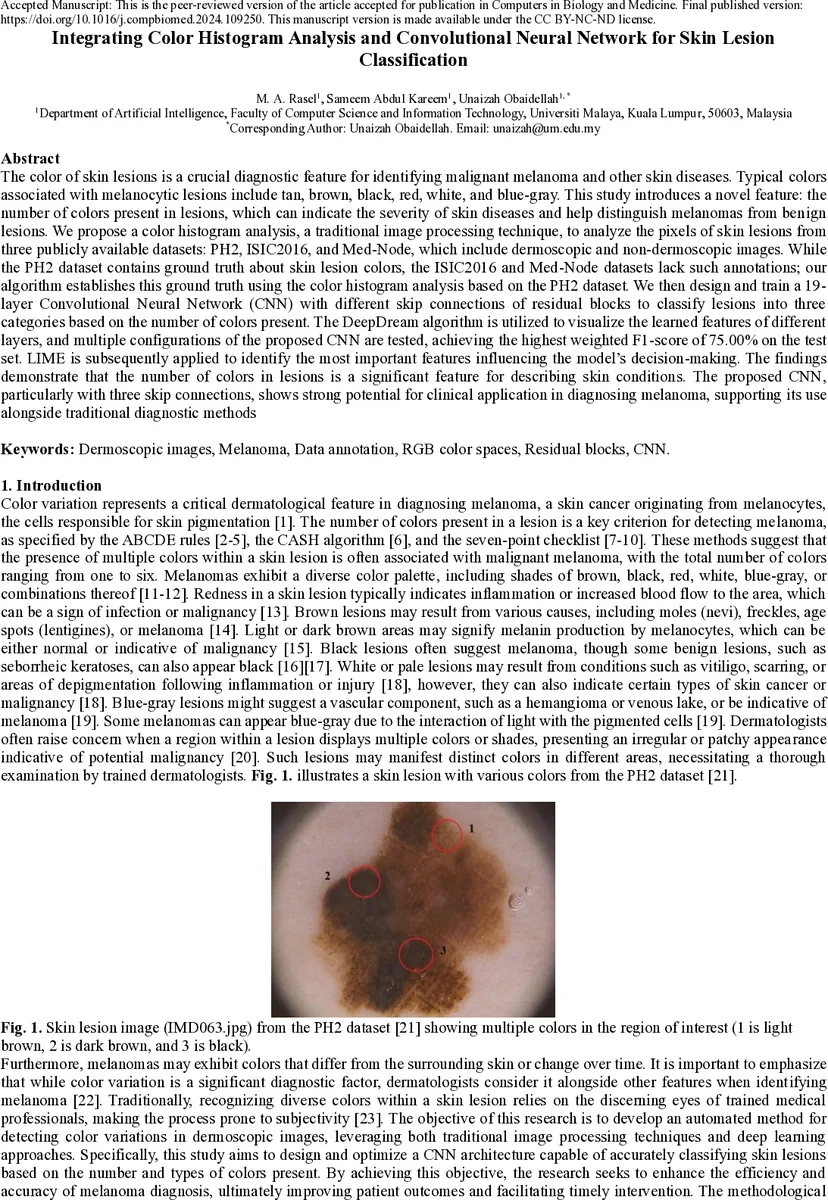
The color of skin lesions is an important diagnostic feature for identifying malignant melanoma and other skin diseases. Typical colors associated with melanocytic lesions include tan, brown, black, red, white, and blue gray. This study introduces a novel feature: the number of colors present in a lesion, which can indicate the severity of disease and help distinguish melanomas from benign lesions. We propose a color histogram analysis method to examine lesion pixel values from three publicly available datasets: PH2, ISIC2016, and Med Node. The PH2 dataset contains ground truth annotations of lesion colors, while ISIC2016 and Med Node do not; our algorithm estimates the ground truth using color histogram analysis based on PH2. We then design and train a 19 layer Convolutional Neural Network (CNN) with residual skip connections to classify lesions into three categories based on the number of colors present. DeepDream visualization is used to interpret features learned by the network, and multiple CNN configurations are tested. The best model achieves a weighted F1 score of 75 percent. LIME is applied to identify important regions influencing model decisions. The results show that the number of colors in a lesion is a significant feature for describing skin conditions, and the proposed CNN with three skip connections demonstrates strong potential for clinical diagnostic support.
💡 Research Summary
The paper proposes a novel automated pipeline that uses the number of distinct colors present in a skin lesion as a diagnostic feature for melanoma detection. The authors first exploit the PH2 dataset, which contains pixel‑level annotations for six clinically relevant colors (white, red, light brown, dark brown, blue‑gray, black). By analyzing the RGB/HSV distribution of these colors they derive quantitative thresholds and construct a color‑histogram algorithm that can count how many of the six colors appear in any given lesion image. This algorithm is then applied to two larger, unannotated datasets—ISIC2016 (379 selected images) and Med‑Node (170 images)—to generate pseudo‑ground‑truth labels indicating whether a lesion contains one, two, or three colors. Post‑processing steps such as morphological cleaning and removal of tiny regions are used to reduce noise and resolve overlapping color regions.
With these labels, the authors design a 19‑layer convolutional neural network (CNN) that incorporates three residual skip connections. Each block consists of a 3×3 convolution followed by batch normalization and ReLU activation; the skip connections feed the input of a block directly to a deeper layer, mitigating vanishing gradients and allowing the network to combine low‑level edge information with high‑level semantic cues. Input images are resized to 224×224×3, and the final classification layer uses a softmax over three classes (1‑color, 2‑color, 3‑color). Hyper‑parameter tuning identifies a learning rate of 1e‑4, batch size of 32, Adam optimizer, and a dropout rate of 0.3 as optimal. Cross‑validation shows that the residual‑skip architecture outperforms a plain CNN by roughly 6 % in weighted F1 score, with the most pronounced gains in distinguishing lesions that contain three colors.
To interpret the learned representations, the authors employ DeepDream visualizations. Early layers highlight edges and simple color boundaries, middle layers focus on specific color contrasts (e.g., black versus dark brown), and the deepest layers emphasize complex multi‑color patterns that drive the final decision. Additionally, LIME (Local Interpretable Model‑agnostic Explanations) is applied to individual predictions, revealing that the model consistently relies on regions where color variation is pronounced—exactly the clinical cues dermatologists use when applying the ABCDE or CASH criteria.
Performance on the combined test set reaches a weighted F1 score of 75 %, with per‑class precision and recall ranging from 0.71 to 0.78. The model demonstrates robust generalization across datasets that differ in resolution, imaging device, and illumination conditions, indicating that the color‑histogram derived labels are sufficiently reliable for training deep models. Limitations include dependence on fixed HSV thresholds, which can be sensitive to lighting variations, and reduced labeling accuracy when more than three colors are present (the algorithm collapses to the most dominant three).
The authors suggest future work such as adopting perceptually uniform color spaces (e.g., CIELAB), incorporating multi‑label learning to predict both the specific colors and their spatial distribution, and expanding the architecture to handle a larger number of color categories. Overall, the study demonstrates that integrating traditional color‑histogram analysis with a residual‑skip CNN yields an effective, interpretable system for quantifying lesion color diversity—a feature that aligns with established dermatological diagnostic rules and holds promise for clinical decision support.
Comments & Academic Discussion
Loading comments...
Leave a Comment