oculomix: Hierarchical Sampling for Retinal-Based Systemic Disease Prediction
Oculomics - the concept of predicting systemic diseases, such as cardiovascular disease and dementia, through retinal imaging - has advanced rapidly due to the data efficiency of transformer-based foundation models like RETFound. Image-level mixed sample data augmentations, such as CutMix and MixUp, are frequently used for training transformers, yet these techniques perturb patient-specific attributes, such as medical comorbidity and clinical factors, since they only account for images and labels. To address this limitation, we propose a hierarchical sampling strategy, Oculomix, for mixed sample augmentations. Our method is based on two clinical priors. First (exam level), images acquired from the same patient at the same time point share the same attributes. Second (patient level), images acquired from the same patient at different time points have a soft temporal trend, as morbidity generally increases over time. Guided by these priors, our method constrains the mixing space to the patient and exam levels to better preserve patient-specific characteristics and leverages their hierarchical relationships. The proposed method is validated using ViT models on a five-year prediction of major adverse cardiovascular events (MACE) in a large ethnically diverse population (Alzeye). We show that Oculomix consistently outperforms image-level CutMix and MixUp by up to 3% in AUROC, demonstrating the necessity and value of the proposed method in oculomics.
💡 Research Summary
The paper tackles a critical shortcoming of conventional mixed‑sample data augmentations (MSDA) such as CutMix and MixUp when applied to retinal‑based systemic disease prediction (oculomics). Standard MSDA treats each image as an independent unit, blending pixel regions and class labels without regard for patient‑specific clinical information (age, sex, comorbidities, socioeconomic status, etc.). This can corrupt the very attributes that make retinal images predictive of systemic outcomes. To remedy this, the authors introduce Oculomix, a hierarchical sampling strategy that respects the natural patient‑exam‑image hierarchy present in longitudinal ophthalmic datasets.
Clinical priors
- Exam‑level prior – All images captured during a single clinical visit (typically both eyes, macula‑centered and disc‑centered) share the same clinical context (demographics, disease labels, risk factors).
- Patient‑level prior – Across visits, a patient’s disease burden evolves gradually; later exams generally reflect higher morbidity.
Methodology
- Exam‑level sampling restricts mixing to images from the same exam of the same patient. The mixed label is simply the original binary label for that exam, preserving all exam‑level covariates.
- Patient‑level sampling allows mixing images from different exams of the same patient. Because a binary label cannot meaningfully represent a blend of two time points, the authors replace direct label interpolation with a pairwise ranking loss. For two exams at times t₁ < t₂, the loss enforces a margin‑based ordering on the predicted logits (logitₜ₂ – logitₜ₁ ≥ m). This encodes temporal progression without requiring an explicit mixed label.
The overall training objective combines the standard cross‑entropy loss for supervised learning with the ranking loss for patient‑level mixes:
L = L_sup(y_ti, ŷ_ti) + L_ranking(ŷ_ti, ŷ_mix).
Experimental setup
- Datasets: The large, ethnically diverse AlzEye cohort (≈353k patients) with linked retinal images and 5‑year major adverse cardiovascular events (MACE) labels; an external Korean HYU MACE cohort for validation.
- Models: Vision Transformers (ViT‑Small and ViT‑Base) pretrained on ImageNet‑21k, fine‑tuned with AdamW (lr = 2e‑4, batch = 512, 400 epochs).
- Comparisons: Three sampling strategies (Image‑level, Exam‑level, Patient+Exam‑level) combined with CutMix, MixUp, and CutMix+MixUp; also a no‑augmentation baseline.
Results
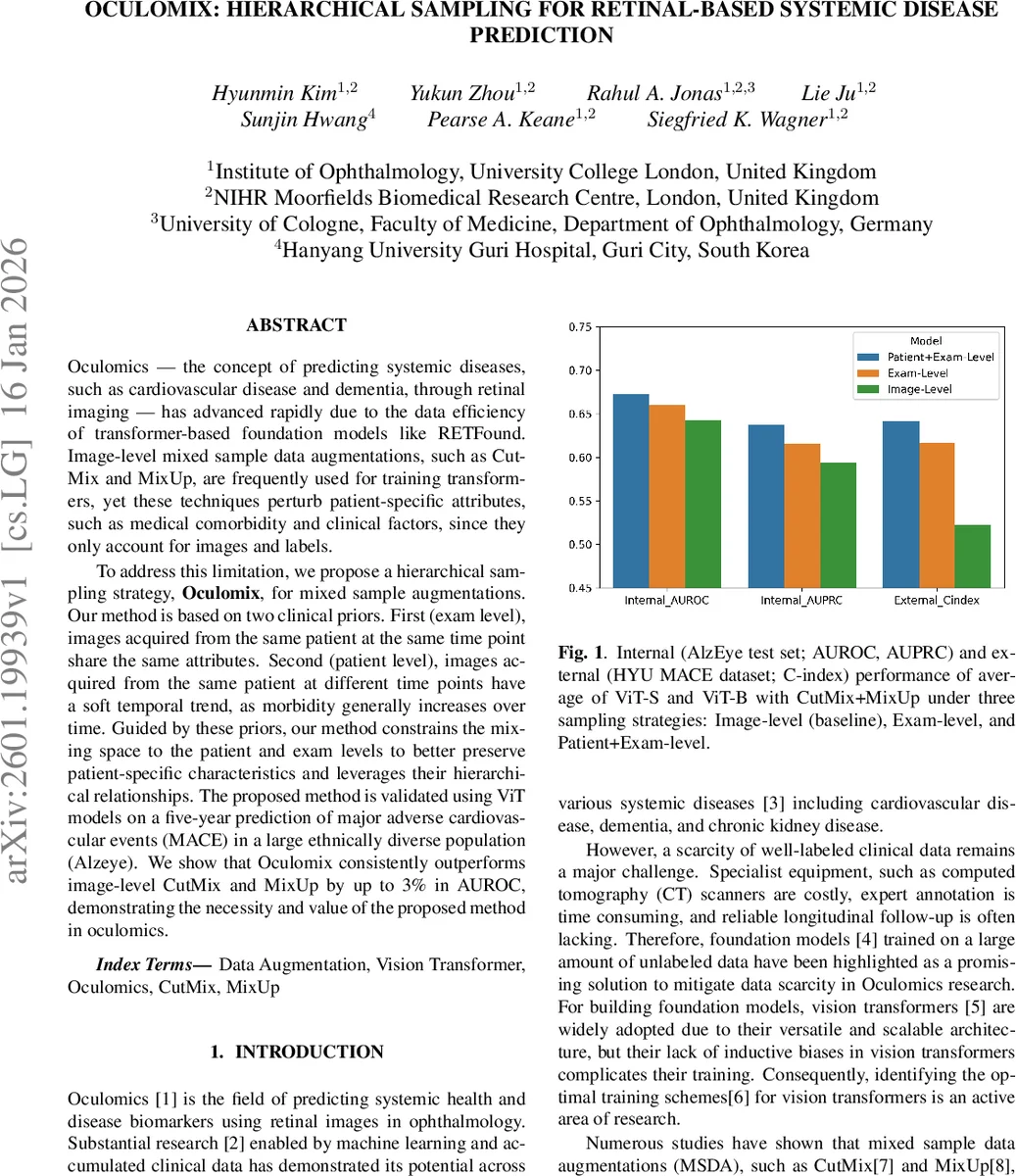
- Internal test: Patient+Exam‑level sampling achieved the highest AUROC (67.22 % for ViT‑Small, 67.29 % for ViT‑Base), outperforming Image‑level by ~3.6 percentage points and Exam‑level by ~1 point. AUPRC improved by up to 4.4 percentage points.
- External test: The C‑index rose from 49.62 % (Image‑level) to 64.98 % (Patient+Exam‑level), a 12 % absolute gain, demonstrating robust generalisation across populations.
- Ablation: When CutMix or MixUp were applied alone, Patient+Exam still outperformed Exam‑level, but the gap widened dramatically when both augmentations were combined, indicating synergistic benefits from richer mixing combinations.
- Supervision type: Relative supervision (ranking loss) consistently beat direct label supervision (linear interpolation of binary labels) for patient‑level mixes, confirming that temporal ordering is a more reliable proxy than ambiguous mixed labels.
Interpretation
The hierarchical constraint reduces the combinatorial space of mixes that would otherwise pair images from unrelated patients, thereby preserving patient‑specific risk factors. Patient‑level mixing expands the diversity of training pairs, mitigating over‑fitting observed with Exam‑level only. The ranking loss elegantly sidesteps the ill‑posed problem of defining a mixed label across time, while still providing a learning signal that respects disease progression.
Limitations and future work
- The study focuses on binary MACE outcomes; extending to multi‑label or continuous risk scores may require more sophisticated temporal modeling.
- Only retinal fundus images were used; integrating OCT or multimodal data could further boost predictive power.
- The ranking margin m was fixed; adaptive margins or learned temporal embeddings could refine supervision.
Conclusion
Oculomix introduces a clinically informed, hierarchy‑aware augmentation framework that markedly improves transformer‑based retinal oculomics. By aligning data mixing with the intrinsic patient‑exam structure and employing a temporal ranking loss, the method preserves critical clinical attributes, enhances model robustness, and achieves state‑of‑the‑art performance on both internal and external cohorts. This work paves the way for more nuanced data augmentation strategies in medical imaging where patient context is paramount.
Comments & Academic Discussion
Loading comments...
Leave a Comment