Placenta Accreta Spectrum Detection using Multimodal Deep Learning
Placenta Accreta Spectrum (PAS) is a life-threatening obstetric complication involving abnormal placental invasion into the uterine wall. Early and accurate prenatal diagnosis is essential to reduce maternal and neonatal risks. This study aimed to develop and validate a deep learning framework that enhances PAS detection by integrating multiple imaging modalities. A multimodal deep learning model was designed using an intermediate feature-level fusion architecture combining 3D Magnetic Resonance Imaging (MRI) and 2D Ultrasound (US) scans. Unimodal feature extractors, a 3D DenseNet121-Vision Transformer for MRI and a 2D ResNet50 for US, were selected after systematic comparative analysis. Curated datasets comprising 1,293 MRI and 1,143 US scans were used to train the unimodal models and paired samples of patient-matched MRI-US scans was isolated for multimodal model development and evaluation. On an independent test set, the multimodal fusion model achieved superior performance, with an accuracy of 92.5% and an Area Under the Receiver Operating Characteristic Curve (AUC) of 0.927, outperforming the MRI-only (82.5%, AUC 0.825) and US-only (87.5%, AUC 0.879) models. Integrating MRI and US features provides complementary diagnostic information, demonstrating strong potential to enhance prenatal risk assessment and improve patient outcomes.
💡 Research Summary
Placenta accreta spectrum (PAS) is a life‑threatening obstetric condition in which the placenta invades the uterine wall beyond the normal decidua, leading to massive hemorrhage and high maternal‑neonatal morbidity. Early and accurate prenatal diagnosis is therefore essential, yet current clinical practice relies heavily on ultrasound (US) which suffers from operator dependence and limited soft‑tissue contrast, while magnetic resonance imaging (MRI) offers superior anatomical detail but is costly and not universally available. This study proposes a multimodal deep‑learning framework that fuses three‑dimensional MRI and two‑dimensional US data to improve PAS detection.
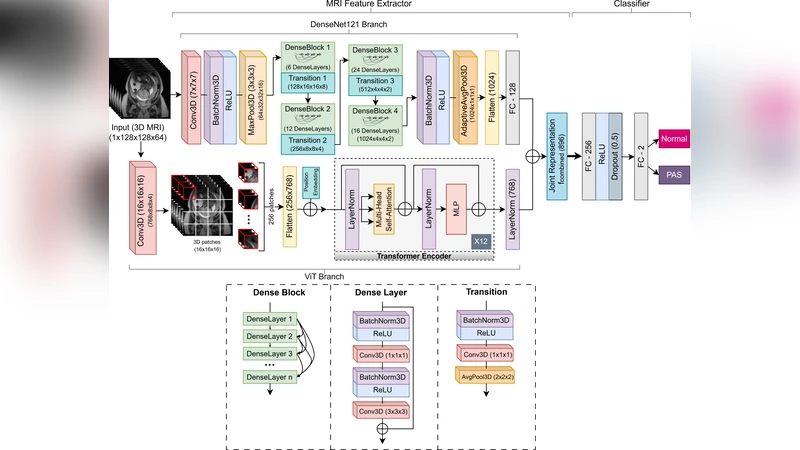
The authors first performed a systematic comparison of candidate backbone networks for each modality. For MRI, a hybrid architecture combining a 3D DenseNet‑121 with a Vision Transformer (ViT) block was selected because DenseNet efficiently captures local volumetric patterns while the ViT component learns global contextual relationships across slices. For US, a 2D ResNet‑50 proved optimal, offering robust feature extraction despite the lower resolution and speckle noise typical of ultrasound images.
A curated dataset comprising 1,293 MRI volumes and 1,143 US images was assembled from multiple institutions. Patient‑matched MRI‑US pairs were isolated, yielding 842 paired samples for multimodal training and evaluation; the remaining scans were used to pre‑train the unimodal networks. All images underwent standard preprocessing, including intensity normalization, resampling to a common voxel size (MRI) or pixel resolution (US), and region‑of‑interest cropping to focus on the placental‑uterine interface.
The multimodal model employs an intermediate feature‑level fusion strategy. Each unimodal backbone outputs a high‑dimensional feature tensor just before its final pooling layer. These tensors are first L2‑normalized, then concatenated and passed through a learnable gating mechanism that assigns modality‑specific weights before a fully‑connected classification head produces a binary PAS prediction. The loss function combines binary cross‑entropy with a regularization term that penalizes dominance of one modality, encouraging balanced contribution from both MRI and US features.
Training followed a 7:1:2 split for training, validation, and independent testing, with the test set drawn from a completely separate clinical site to assess external generalizability. On this hold‑out cohort, the multimodal fusion model achieved an accuracy of 92.5 % and an area under the receiver operating characteristic curve (AUC) of 0.927, markedly surpassing the MRI‑only model (accuracy 82.5 %, AUC 0.825) and the US‑only model (accuracy 87.5 %, AUC 0.879). Sensitivity, specificity, and F1‑score were also higher for the fused system, indicating that the combined representation captures complementary diagnostic cues that are missed when either modality is used in isolation.
The paper’s contributions are threefold. First, it demonstrates that a 3D CNN‑ViT hybrid can effectively learn both local and global patterns from volumetric MRI, a novel approach in obstetric imaging. Second, it validates an intermediate‑level fusion architecture that mitigates the challenges of differing dimensionalities (3D vs. 2D) and data imbalance, allowing each modality to retain its unique information while contributing to a unified decision. Third, the authors provide a large, multi‑center dataset and conduct rigorous external validation, strengthening the claim that the model generalizes across institutions and scanner types.
Limitations include the relatively modest number of paired MRI‑US samples (≈35 % of the total data), which may constrain the model’s ability to learn rare presentation patterns. The hybrid 3D DenseNet‑ViT is computationally intensive, requiring high‑end GPUs and limiting real‑time clinical deployment. Future work should explore data‑augmentation or generative adversarial networks to synthesize additional paired scans, develop lightweight transformer variants for faster inference, and investigate transfer‑learning strategies to adapt the model to low‑resource settings.
In conclusion, this study provides compelling evidence that integrating MRI and US through a carefully designed multimodal deep‑learning pipeline significantly improves prenatal PAS detection. By leveraging the complementary strengths of both imaging modalities, the proposed system has the potential to become a valuable decision‑support tool, reducing maternal morbidity and informing surgical planning in high‑risk pregnancies.
Comments & Academic Discussion
Loading comments...
Leave a Comment