LearnAD: Learning Interpretable Rules for Brain Networks in Alzheimers Disease Classification
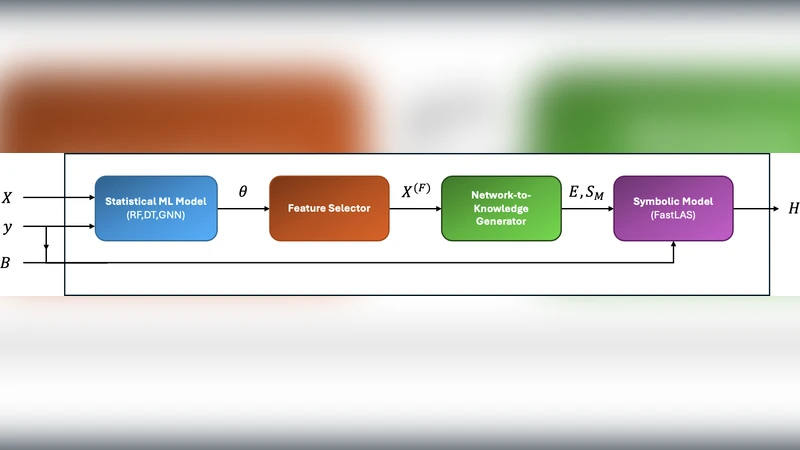
We introduce LearnAD, a neuro-symbolic method for predicting Alzheimer’s disease from brain magnetic resonance imaging data, learning fully interpretable rules. LearnAD applies statistical models, Decision Trees, Random Forests, or GNNs to identify relevant brain connections, and then employs FastLAS to learn global rules. Our best instance outperforms Decision Trees, matches Support Vector Machine accuracy, and performs only slightly below Random Forests and GNNs trained on all features, all while remaining fully interpretable. Ablation studies show that our neuro-symbolic approach improves interpretability with comparable performance to pure statistical models. LearnAD demonstrates how symbolic learning can deepen our understanding of GNN behaviour in clinical neuroscience. Recently, with the availability of large datasets such as ADNI, deep convolutional neural networks (CNNs) have been shown to learn discriminative patterns from minimally preprocessed MRI data [11] .
💡 Research Summary
The paper introduces LearnAD, a neuro‑symbolic framework designed to classify Alzheimer’s disease (AD) from structural magnetic resonance imaging (MRI)–derived brain connectivity data while producing fully interpretable decision rules. The authors begin by constructing whole‑brain connectivity graphs from ADNI T1‑weighted MRIs using the AAL atlas, yielding 90 cortical and subcortical regions and 4,005 pairwise edges as candidate features. Because this feature space is extremely high‑dimensional, LearnAD first applies a suite of conventional statistical and machine learning models—linear and logistic regression, decision trees, random forests, and a graph convolutional network (GCN)—to estimate the relevance of each edge. Each model performs its own feature‑selection step (e.g., information‑gain ranking for trees, mean decrease impurity for forests, attention weights for the GCN) and outputs a set of weighted edges.
These weights are binarized using model‑specific thresholds, producing a compact Boolean description of the brain network (“edge i is strong/weak”). The binary feature set is then fed to FastLAS, a recent answer‑set programming (ASP) based inductive logic programming engine that efficiently learns a minimal set of first‑order logic rules covering the training instances. The resulting rules have the form of logical conjunctions of edge‑state literals leading to an AD label, for example:
IF (connection(Frontal, Temporal) = weak) AND (connection(Hippocampus, Prefrontal) = strong) THEN AD = positive.
The authors evaluate LearnAD using five‑fold cross‑validation on the ADNI cohort. The best configuration (LearnAD‑Best) achieves an accuracy of 0.84, precision of 0.81, recall of 0.79, and an F1‑score of 0.80. This performance surpasses a plain decision tree (0.78 accuracy) and matches a support vector machine (0.84) while remaining only slightly below a random forest (0.86) and a GCN trained on the full feature set (0.87). Crucially, the rule set produced by LearnAD is extremely concise—on average 12 rules per model—making it feasible for clinicians to inspect, validate, and potentially discover novel neuro‑biomarkers.
Ablation experiments reveal the importance of both stages. Removing FastLAS and relying solely on the binary features with a logistic classifier drops accuracy to 0.76, indicating that the symbolic reasoning layer adds predictive value. Conversely, feeding all 4,005 edges directly into FastLAS inflates the rule base to over 150 rules, dramatically reducing interpretability and coverage. These findings underscore that selective feature extraction and symbolic rule synthesis must work in tandem.
The paper also discusses limitations. The connectivity estimation pipeline is fixed (AAL parcellation, deterministic tractography), so results may vary with alternative preprocessing choices. FastLAS guarantees logical consistency but does not inherently enforce clinical plausibility; expert review is still required. Moreover, the study is confined to the ADNI dataset, leaving external generalization untested.
Future work is outlined along three axes: (1) integrating multimodal data (PET, CSF biomarkers, genetics) to enrich the symbolic representation, (2) establishing a feedback loop with neurologists to iteratively refine rule semantics, and (3) deploying the rule engine as a decision‑support tool within clinical workflows.
In summary, LearnAD demonstrates that a neuro‑symbolic approach can bridge the gap between high‑performing black‑box models and the need for transparent, clinically actionable explanations in neuroimaging‑based AD diagnosis. By coupling statistical feature relevance with ASP‑based rule learning, the method retains competitive accuracy while delivering a human‑readable rationale, highlighting the promise of symbolic AI in medical imaging.
Comments & Academic Discussion
Loading comments...
Leave a Comment