Adaptive Frequency Domain Alignment Network for Medical image segmentation
High-quality annotated data plays a crucial role in achieving accurate segmentation. However, such data for medical image segmentation are often scarce due to the time-consuming and labor-intensive nature of manual annotation. To address this challenge, we propose the Adaptive Frequency Domain Alignment Network (AFDAN)–a novel domain adaptation framework designed to align features in the frequency domain and alleviate data scarcity. AFDAN integrates three core components to enable robust cross-domain knowledge transfer: an Adversarial Domain Learning Module that transfers features from the source to the target domain; a Source-Target Frequency Fusion Module that blends frequency representations across domains; and a Spatial-Frequency Integration Module that combines both frequency and spatial features to further enhance segmentation accuracy across domains. Extensive experiments demonstrate the effectiveness of AFDAN: it achieves an Intersection over Union (IoU) of 90.9% for vitiligo segmentation in the newly constructed VITILIGO2025 dataset and a competitive IoU of 82.6% on the retinal vessel segmentation benchmark DRIVE, surpassing existing state-of-the-art approaches.
💡 Research Summary
The paper introduces the Adaptive Frequency Domain Alignment Network (AFDAN), a novel domain‑adaptation framework designed to mitigate the scarcity of annotated medical images by aligning source and target features in the frequency domain and fusing them with spatial representations. AFDAN consists of three core modules. The Adversarial Domain Learning (ADL) module applies an adversarial game to the amplitude spectra of source and target images, encouraging a generator to produce source amplitudes indistinguishable from target amplitudes, thereby reducing style‑related domain shift. The Source‑Target Frequency Fusion (STFF) module decomposes source and target features via 2‑D FFT, retains the source phase (which encodes structural information), and stochastically mixes low‑frequency amplitude components using a random coefficient α∈U(0,1). This creates dense intermediate amplitude distributions that accelerate adaptation while preserving high‑frequency details essential for fine anatomical structures. The Spatial‑Frequency Integration (SFI) module resamples frequency‑domain features to match spatial feature dimensions, concatenates them channel‑wise, and applies a spatial‑frequency attention mechanism to suppress noise and highlight domain‑invariant regions. The combined representation is decoded to produce the final segmentation mask.
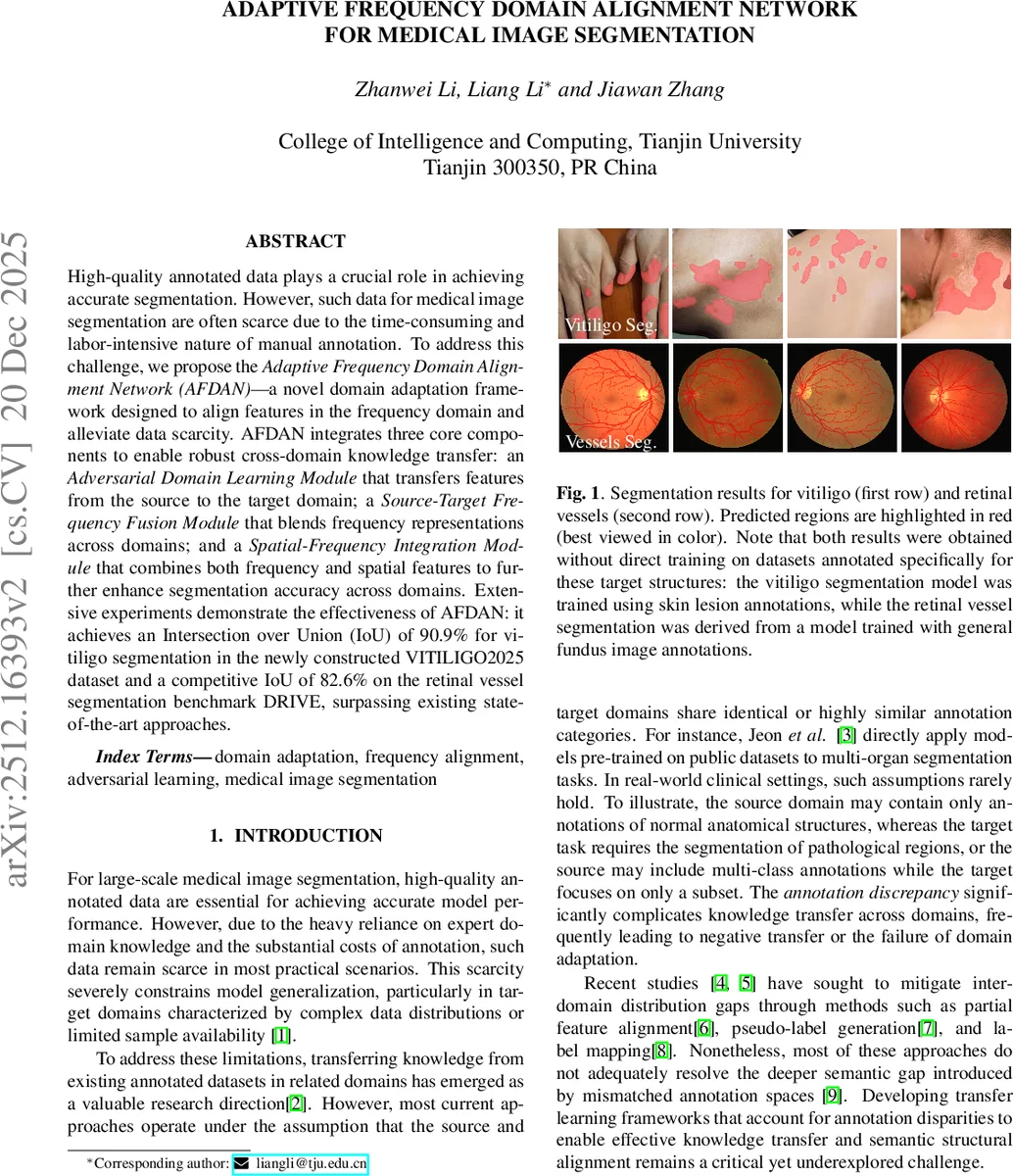
Experiments were conducted on two cross‑domain tasks: (1) vitiligo segmentation, where ISIC2018 (skin lesion) images serve as the source and the newly constructed VITILIGO2025 dataset as the target; (2) retinal vessel segmentation, where Fundus‑A VSeg is the source and DRIVE is the target. AFDAN achieved state‑of‑the‑art performance, reaching 90.9 % IoU (95.2 % Dice) on VITILIGO2025 and 82.6 % IoU (90.5 % Dice) on DRIVE, surpassing strong baselines such as PSPNet, YOLO11, EHTDI, and DSTC‑SSD A. Ablation studies demonstrated that each module contributes positively: STFF alone adds up to +4.1 %p IoU on vitiligo and +2.1 %p on DRIVE; ADL contributes +3.6 %p and +0.9 %p respectively; SFI adds +4.9 %p and +1.7 %p. Combinations of modules yield synergistic gains, with the full model delivering the highest improvements (+10.6 %p on vitiligo, +6.1 %p on DRIVE).
The authors argue that operating in the frequency domain, particularly focusing on amplitude alignment, reduces the semantic gap caused by mismatched annotation spaces (e.g., normal anatomy vs. pathological lesions) and preserves structural fidelity via phase retention. The modular design, built on a SegFormer backbone, allows easy integration with other architectures. However, the study is limited to two target domains, lacks extensive computational cost analysis, and does not explore joint amplitude‑phase alignment or multi‑source adaptation. Future work is suggested to extend the approach to multi‑class, multi‑source scenarios, investigate lightweight frequency transforms, and validate clinical applicability across diverse imaging modalities. Overall, AFDAN presents a compelling strategy for leveraging frequency‑domain information to achieve robust medical image segmentation under annotation scarcity.
Comments & Academic Discussion
Loading comments...
Leave a Comment