📝 Original Info
- Title: NEURO-GUARD: Neuro-Symbolic Generalization and Unbiased Adaptive Routing for Diagnostics – Explainable Medical AI
- ArXiv ID: 2512.18177
- Date: 2025-12-20
- Authors: Researchers from original ArXiv paper
📝 Abstract
Accurate yet interpretable image-based diagnosis remains a central challenge in medical AI, particularly in settings characterized by limited data, subtle visual cues, and high-stakes clinical decision-making. Most existing vision models rely on purely data-driven learning and produce black-box predictions with limited interpretability and poor cross-domain generalization, hindering their real-world clinical adoption. We present NEURO-GUARD, a novel knowledge-guided vision framework that integrates Vision Transformers (ViTs) with language-driven reasoning to improve performance, transparency, and domain robustness. NEURO-GUARD employs a retrieval-augmented generation (RAG) mechanism for self-verification, in which a large language model (LLM) iteratively generates, evaluates, and refines feature-extraction code for medical images. By grounding this process in clinical guidelines and expert knowledge, the framework progressively enhances feature detection and classification beyond purely data-driven baselines. Extensive experiments on diabetic retinopathy classification across four benchmark datasets APTOS, EyePACS, Messidor-1, and Messidor-2 demonstrate that NEURO-GUARD improves accuracy by 6.2% over a ViT-only baseline (84.69% vs. 78.4%) and achieves a 5% gain in domain generalization. Additional evaluations on MRI-based seizure detection further confirm its cross-domain robustness, consistently outperforming existing methods.
Overall, NEURO-GUARD bridges symbolic medical reasoning with subsymbolic visual learning, enabling interpretable, knowledge-aware, and generalizable medical image diagnosis while achieving state-of-the-art performance across multiple datasets.
💡 Deep Analysis
Deep Dive into NEURO-GUARD: Neuro-Symbolic Generalization and Unbiased Adaptive Routing for Diagnostics -- Explainable Medical AI.
Accurate yet interpretable image-based diagnosis remains a central challenge in medical AI, particularly in settings characterized by limited data, subtle visual cues, and high-stakes clinical decision-making. Most existing vision models rely on purely data-driven learning and produce black-box predictions with limited interpretability and poor cross-domain generalization, hindering their real-world clinical adoption. We present NEURO-GUARD, a novel knowledge-guided vision framework that integrates Vision Transformers (ViTs) with language-driven reasoning to improve performance, transparency, and domain robustness. NEURO-GUARD employs a retrieval-augmented generation (RAG) mechanism for self-verification, in which a large language model (LLM) iteratively generates, evaluates, and refines feature-extraction code for medical images. By grounding this process in clinical guidelines and expert knowledge, the framework progressively enhances feature detection and classification beyond purel
📄 Full Content
NEURO-GUARD: Neuro-Symbolic Generalization and Unbiased Adaptive
Routing for Diagnostics - Explainable Medical AI
Midhat Urooj
Arizona State University
Tempe, AZ, USA
murooj@asu.edu
Ayan Banerjee
Arizona State University
Tempe, AZ, USA
abanerj3@asu.edu
Sandeep Gupta
Arizona State University
Tempe, AZ, USA
Sandeep.Gupta@asu.edu
Abstract
Accurate yet interpretable image-based diagnosis remains
a central challenge in medical AI, particularly in settings
with limited data, subtle visual patterns, and high-stakes
clinical decisions.
However, most current vision models
produce black-box predictions with limited generalizabil-
ity and poor real-world usability. We present
NEURO-
GUARD, a novel framework that combines Vision Trans-
formers (ViTs) with knowledge-guided reasoning to en-
hance performance, transparency, and cross-domain gen-
eralization.
NEURO-GUARD incorporates a retrieval-
augmented generation (RAG) mechanism for language-
driven self-verification, in which a large language model
(LLM) iteratively generates, evaluates, and refines feature
extraction code for medical images. By leveraging clini-
cal guidelines and expert knowledge, this LLM-guided mod-
ule progressively improves feature detection and classifica-
tion, outperforming purely data-driven baselines. Extensive
evaluations on diabetic retinopathy classification across
four benchmark datasets (APTOS, EyePACS, Messidor-1,
Messidor-2) show that NEURO-GUARD improves accuracy
by 6.2% over a ViT-only model (84.69% vs. 78.4% [3])
and achieves a 5% gain in domain generalization. Fur-
ther experiments on MRI-based seizure detection confirm its
cross-domain robustness, consistently surpassing existing
baselines. Notably, NEURO-GUARD bridges the gap be-
tween symbolic medical reasoning and subsymbolic feature
learning, demonstrating robust generalization across multi-
ple datasets while achieving state-of-the-art performance.
1. Introduction
Medical imaging plays a crucial role in disease diagnosis
and treatment planning, particularly in conditions such as
diabetic retinopathy (DR), tumor detection, and neurode-
generative disorders.
Recent advances in deep learning,
Figure 1.
Performance comparison of existing models versus
the NEURO-GUARD framework for 5-stage Diabetic Retinopa-
thy classification.
particularly Vision Transformers (ViTs) and Convolutional
Neural Networks (CNNs), have significantly improved di-
agnostic accuracy [3, 14]. However, their black-box na-
ture limits clinical adoption due to a lack of interpretability,
making it challenging for clinicians to validate AI-driven
decisions. Additionally, these models suffer from domain
shift vulnerabilities, struggling to generalize across imag-
ing datasets with diverse acquisition protocols and patient
demographics [22, 26]. Given these challenges, an ideal
medical AI framework should not only provide high accu-
racy but also generate clinically interpretable decisions by
integrating structured domain knowledge into its reasoning
process.
Existing explainability techniques, such as Gradient-
weighted Class Activation Mapping (Grad-CAM) [13] and
Shapley Additive Explanations (SHAP) [8], provide post-
hoc feature attribution but remain static, heuristic-based,
and disconnected from the model’s decision logic.
Hy-
brid approaches incorporating attention mechanisms and
uncertainty estimation attempt to improve interpretabil-
arXiv:2512.18177v1 [cs.AI] 20 Dec 2025
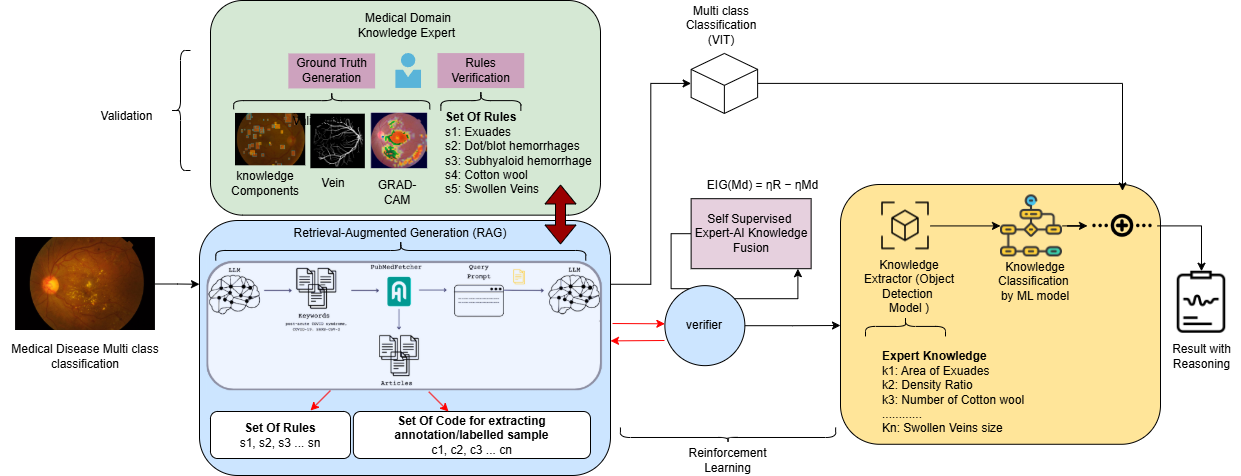
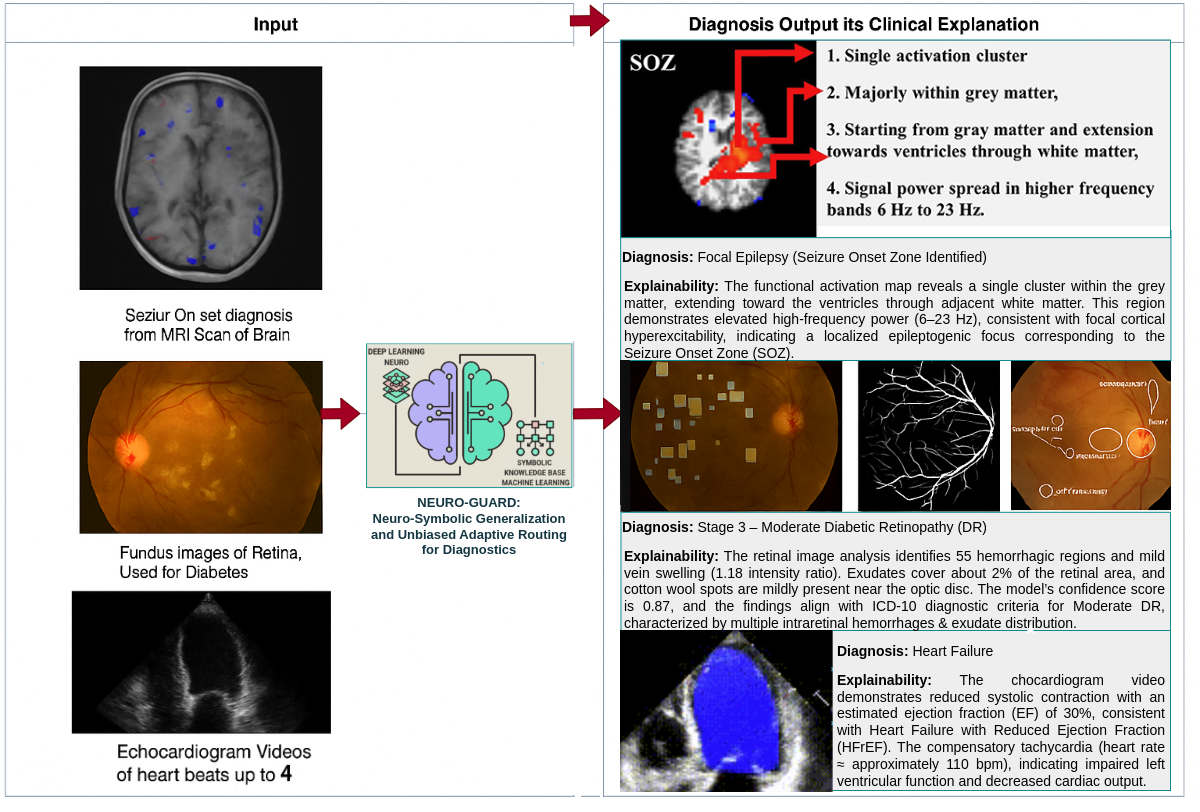
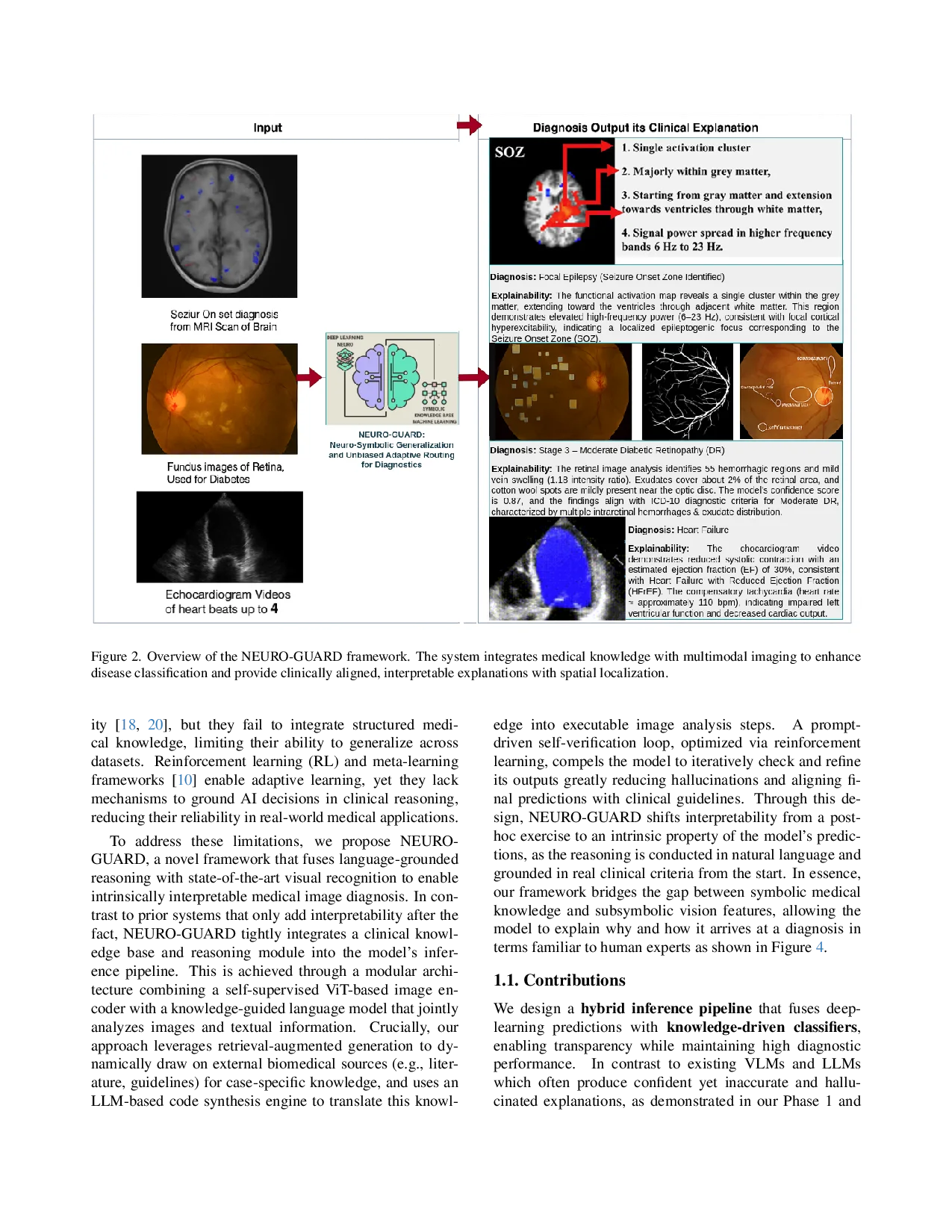
Figure 2. Overview of the NEURO-GUARD framework. The system integrates medical knowledge with multimodal imaging to enhance
disease classification and provide clinically aligned, interpretable explanations with spatial localization.
ity [18, 20], but they fail to integrate structured medi-
cal knowledge, limiting their ability to generalize across
datasets. Reinforcement learning (RL) and meta-learning
frameworks [10] enable adaptive learning, yet they lack
mechanisms to ground AI decisions in clinical reasoning,
reducing their reliability in real-world medical applications.
To address these limitations, we propose NEURO-
GUARD, a novel framework that fuses language-grounded
reasoning with state-of-the-art visual recognition to enable
intrinsically interpretable medical image diagnosis. In con-
trast to prior systems that only add interpretability after the
fact, NEURO-GUARD tightly integrates a clinical knowl-
edge base and reasoning module into the model’s infer-
ence pipeline. This is achieved through a modular archi-
tecture combining a self-supervised ViT-based image en-
coder with a knowledge-guided language model that jointly
analyzes images and textual information.
Crucially, our
approach leverages retrieval-augmented generation to dy-
namically draw on external biomedical sources (e.g., liter-
ature, guidelines) for case-specific knowledge, and uses an
LLM-based code synthesis engine to translate this knowl-
edge into executable image analysis steps.
A prompt-
driven self-verification loop, optimized via reinforcement
learni
…(Full text truncated)…
📸 Image Gallery
Reference
This content is AI-processed based on ArXiv data.