ReX-MLE: The Autonomous Agent Benchmark for Medical Imaging Challenges
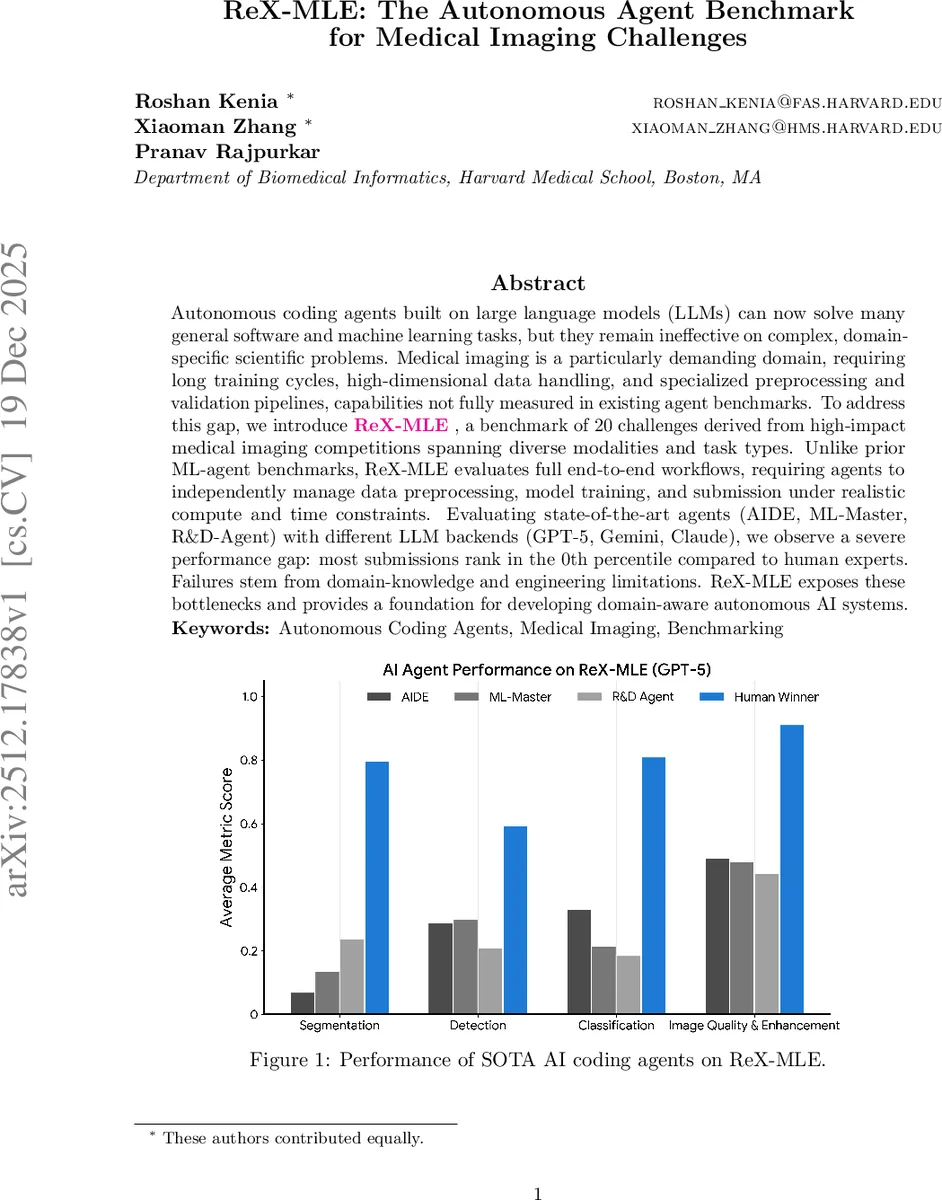
Autonomous coding agents built on large language models (LLMs) can now solve many general software and machine learning tasks, but they remain ineffective on complex, domain-specific scientific problems. Medical imaging is a particularly demanding domain, requiring long training cycles, high-dimensional data handling, and specialized preprocessing and validation pipelines, capabilities not fully measured in existing agent benchmarks. To address this gap, we introduce ReX-MLE, a benchmark of 20 challenges derived from high-impact medical imaging competitions spanning diverse modalities and task types. Unlike prior ML-agent benchmarks, ReX-MLE evaluates full end-to-end workflows, requiring agents to independently manage data preprocessing, model training, and submission under realistic compute and time constraints. Evaluating state-of-the-art agents (AIDE, ML-Master, R&D-Agent) with different LLM backends (GPT-5, Gemini, Claude), we observe a severe performance gap: most submissions rank in the 0th percentile compared to human experts. Failures stem from domain-knowledge and engineering limitations. ReX-MLE exposes these bottlenecks and provides a foundation for developing domain-aware autonomous AI systems.
💡 Research Summary
The paper introduces ReX‑MLE, a new benchmark designed to evaluate autonomous coding agents on realistic medical imaging challenges. While large language model (LLM)‑based agents have shown impressive results on generic software and machine‑learning tasks, they struggle with the high‑dimensional data, long training cycles, and specialized preprocessing pipelines that characterize medical imaging. To fill this evaluation gap, the authors curated 20 tasks drawn from ten high‑impact Grand Challenge competitions, covering eight imaging modalities (CT, MRI, MRA, ultrasound, pathology, microscopy, etc.) and four task types (segmentation, detection, classification, image generation). Each task is provided with the exact data formats, sample submissions, and official evaluation scripts used in the original competitions, forcing agents to produce submission‑ready outputs such as NIfTI volumes, mask files, and JSON detections.
Three state‑of‑the‑art autonomous agents are evaluated: AIDE (greedy tree‑search code optimizer), ML‑Master (Monte‑Carlo Tree Search with multi‑trajectory exploration), and R&D‑Agent (dual‑module researcher/developer design). All agents run on a standardized hardware configuration (NVIDIA H100 GPU, 64‑core CPU, 128 GB RAM) and are limited to a 24‑hour wall‑clock budget per challenge, mirroring realistic research constraints. The primary LLM backend is GPT‑5, with additional runs using Gemini 3 Pro and Claude 4.5 Sonnet for comparison. Agents receive five inputs: competition description, full training data with annotations, test data without ground truth, a sample submission, and a local evaluation script. No human intervention is allowed.
Performance is measured both by absolute metric scores (Dice, mAP, AUC, LNCC, etc.) and by relative percentile rank against the top human competitors on each leaderboard. The results are stark: across the 20 challenges, agents achieve median percentiles below 10 %, with many tasks scoring 0 %—essentially the bottom of the human leaderboard. Simple 2‑D pathology classification (HCD) is the only exception where ML‑Master reaches an AUC of 0.992, comparable to human experts. In contrast, high‑dimensional 3‑D segmentation (e.g., ISLES’22, PANTHER‑T1/T2, TopCoW‑CT A‑Seg) and detection tasks yield Dice or IoU scores near zero, indicating fundamental failures in data handling, normalization, augmentation, or model training.
To probe why agents fail, the authors conduct a capability analysis using 13 “Winning Strategies” identified in prior work (e.g., data exploration, preprocessing design, hyper‑parameter tuning, debugging, reproducibility checks). An automated LLM‑as‑judge pipeline evaluates each agent’s logs for evidence of these strategies, assigning binary indicators. The aggregated profiles reveal that agents rarely exhibit critical engineering practices such as resource‑aware planning, iterative validation, or systematic error recovery. Even when provided with the winning solution reports (“with solution” condition), performance does not improve appreciably, suggesting that the bottleneck lies not only in domain knowledge but also in the ability to translate high‑level descriptions into functional, optimized pipelines.
The paper draws two major implications. First, autonomous agents need enhanced mechanisms for handling large, heterogeneous medical imaging data, including memory‑efficient loading, modality‑specific preprocessing, and long‑duration training orchestration. Second, true scientific autonomy requires integrating self‑debugging, resource‑aware scheduling, and iterative validation loops into the agent architecture, moving beyond mere code generation to full experimental workflow management.
ReX‑MLE itself is released publicly with all datasets, preprocessing scripts, and evaluation code, providing a reproducible testbed for future research. The authors envision extending the benchmark with additional modalities, multimodal fusion tasks, and larger-scale challenges, positioning ReX‑MLE as a cornerstone for developing domain‑aware autonomous AI systems capable of tackling the most demanding scientific problems.
Comments & Academic Discussion
Loading comments...
Leave a Comment