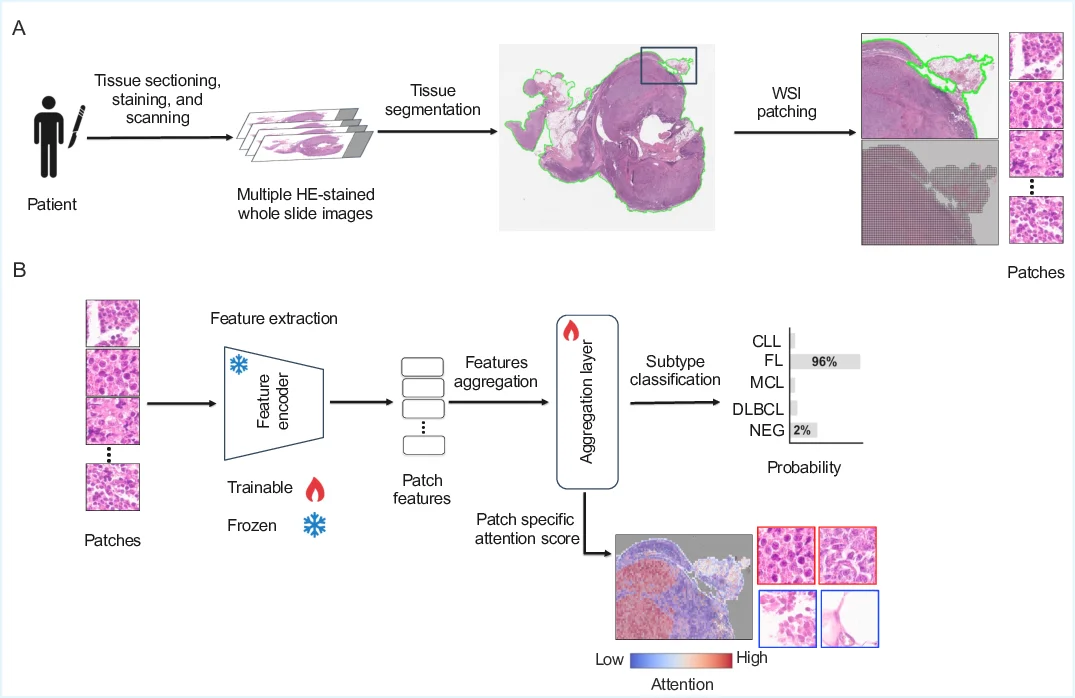
Timely and accurate lymphoma diagnosis is essential for guiding cancer treatment. Standard diagnostic practice combines hematoxylin and eosin (HE)-stained whole slide images with immunohistochemistry, flow cytometry, and molecular genetic tests to determine lymphoma subtypes, a process requiring costly equipment, skilled personnel, and causing treatment delays. Deep learning methods could assist pathologists by extracting diagnostic information from routinely available HE-stained slides, yet comprehensive benchmarks for lymphoma subtyping on multicenter data are lacking. In this work, we present the first multicenter lymphoma benchmarking dataset covering four common lymphoma subtypes and healthy control tissue. We systematically evaluate five publicly available pathology foundation models (H-optimus-1, H0-mini, Virchow2,
Deep Dive into A Multicenter Benchmark of Multiple Instance Learning Models for Lymphoma Subtyping from HE-stained Whole Slide Images.
Timely and accurate lymphoma diagnosis is essential for guiding cancer treatment. Standard diagnostic practice combines hematoxylin and eosin (HE)-stained whole slide images with immunohistochemistry, flow cytometry, and molecular genetic tests to determine lymphoma subtypes, a process requiring costly equipment, skilled personnel, and causing treatment delays. Deep learning methods could assist pathologists by extracting diagnostic information from routinely available HE-stained slides, yet comprehensive benchmarks for lymphoma subtyping on multicenter data are lacking. In this work, we present the first multicenter lymphoma benchmarking dataset covering four common lymphoma subtypes and healthy control tissue. We systematically evaluate five publicly available pathology foundation models (H-optimus-1, H0-mini, Virchow2,
Proceedings of Machine Learning Research – Under Review:1–19, 2025
Full Paper – MIDL 2025 submission
A Multicenter Benchmark of Multiple Instance Learning
Models for Lymphoma Subtyping from HE-stained Whole
Slide Images
Rao Muhammad Umer 1
umer.rao@helmholtz-munich.de
Daniel Sens 1
daniel.sens@helmholtz-munich.de
Jonathan Noll 1
jonathan.noll.leon@gmail.com
Sohom Dey 1
sohom21d@gmail.com
Christian Matek 1,2,7
christian.matek@uk-erlangen.de
Lukas Wolfseher 8
Lukas.Wolfseher@informatik.uni-Kiel.de
Rainer Spang 8
rainer.spang@klinik.uni-r.de
Ralf Huss 9
huss@bio-m.org
Johannes Raffler 9
Johannes.Raffler@uk-augsburg.de
Sarah Reinke 10
sreinke@path.uni-kiel.de
Ario Sadafi 1,6
ario.sadafi@helmholtz-munich.de
Wolfram Klapper 10
wklapper@path.uni-kiel.de
Katja Steiger 6
katja.steiger@tum.de
Kristina Schwamborn 6
kschwamborn@tum.de
Carsten Marr 1,2,3,4,5
carsten.marr@helmholtz-munich.de
1 Institute of AI for Health, Helmholtz Munich, Munich, Germany
2 Department of Medicine III, Ludwig-Maximilian-University Hospital, Munich, Germany
3 Computational Health Center & Helmholtz AI, Helmholtz Munich, Neuherberg, Germany
4 German Cancer Consortium (DKTK), partner site Munich, Germany
5 Munich Center for Machine Learning (MCML), Munich, Germany
6 Technical University of Munich, Munich, Germany
7 Institute of Pathology, Erlangen, Germany
8 University of Kiel, Kiel, Germany
9 Institute for Digital Medicine, University Hospital, Augsburg, Germany
10 Institute of Pathology, University Hospital, Kiel, Germany
Editors: Under Review for MIDL 2025
Abstract
Timely and accurate lymphoma diagnosis is essential for guiding cancer treatment.
Standard diagnostic practice combines hematoxylin and eosin (HE)-stained whole slide im-
ages with immunohistochemistry, flow cytometry, and molecular genetic tests to determine
lymphoma subtypes, a process requiring costly equipment, skilled personnel, and causing
treatment delays. Deep learning methods could assist pathologists by extracting diagnostic
information from routinely available HE-stained slides, yet comprehensive benchmarks for
lymphoma subtyping on multicenter data are lacking.
In this work, we present the first multicenter lymphoma benchmarking dataset cover-
ing four common lymphoma subtypes and healthy control tissue. We systematically evalu-
ate five publicly available pathology foundation models (H-optimus-1, H0-mini, Virchow2,
© 2025 CC-BY 4.0, R.M.U. et al.
arXiv:2512.14640v2 [cs.CV] 3 Feb 2026
UNI2, Titan) combined with attention-based (AB-MIL) and transformer-based (TransMIL)
multiple instance learning aggregators across three magnifications (10×, 20×, 40×). On in-
distribution test sets, models achieve multiclass balanced accuracies exceeding 80% across
all magnifications, with all foundation models performing similarly and both aggregation
methods showing comparable results. The magnification study reveals that 40× resolu-
tion is sufficient, with no performance gains from higher resolutions or cross-magnification
aggregation. However, on out-of-distribution test sets, performance drops substantially to
around 60%, highlighting significant generalization challenges. To advance the field, larger
multicenter studies covering additional rare lymphoma subtypes are needed. We provide
an automated benchmarking pipeline to facilitate such future research.
Keywords: Multicenter Lymphoma Benchmark, Multiple Instance Learning, Whole Slide
Images, Pathology Foundation Models.
1. Introduction
Cancer is one of the deadliest diseases and remains an insurmountable obstacle to advance
the quality and expectancy of life all over the world (Bray et al., 2021). Lymphoma is a
type of blood cancer that originates in the lymphatic system, which is a critical part of hu-
man body’s immune system. It specifically arises from lymphocytes, white blood cells that
play a key role in defending the body against infections. Lymphomas are broadly classified
into two main categories (Lewis et al., 2020): Hodgkin lymphoma (HL) and non-Hodgkin
lymphoma (NHL), with each category having numerous subtypes. The diagnosis of lym-
phoma involves a combination of clinical evaluation, medical imaging, and most importantly,
biopsy of the affected tissue. The biopsy is examined under a microscope (i.e., digitized
as gigapixel HE (Hematoxylin and Eosin) stained whole slide images), and additional tests
like immunohistochemical (IHC) stains, flow cytometry, cytogenetic, and molecular analysis
help to determine the specific subtype of lymphoma (Lewis et al., 2020). These auxiliary
tests require costly equipment, expensive reagents, and trained personnel. Treatment varies
depending on the lymphoma subtype, stage, and other factors such as the patient’s overall
health. Common treatment options (Lewis et al., 2020) include chemotherapy, radiation
therapy, targeted therapy, immunotherapy, and stem cell transplantation.
Histopathology plays a central role in clinical medicine for tissue-based diagnostics and
in biomedical
…(Full text truncated)…
This content is AI-processed based on ArXiv data.