MONET -- Virtual Cell Painting of Brightfield Images and Time Lapses Using Reference Consistent Diffusion
Cell painting is a popular technique for creating human-interpretable, high-contrast images of cell morphology. There are two major issues with cell paint: (1) it is labor-intensive and (2) it requires chemical fixation, making the study of cell dynamics impossible. We train a diffusion model (Morphological Observation Neural Enhancement Tool, or MONET) on a large dataset to predict cell paint channels from brightfield images. We show that model quality improves with scale. The model uses a consistency architecture to generate time-lapse videos, despite the impossibility of obtaining cell paint video training data. In addition, we show that this architecture enables a form of in-context learning, allowing the model to partially transfer to out-of-distribution cell lines and imaging protocols. Virtual cell painting is not intended to replace physical cell painting completely, but to act as a complementary tool enabling novel workflows in biological research.
💡 Research Summary
Cell Painting has become a standard assay for generating high‑contrast, human‑interpretable images of cellular morphology across multiple fluorescent channels. However, the technique suffers from two fundamental drawbacks: it requires a labor‑intensive multi‑step staining protocol and, because cells must be chemically fixed, it cannot capture dynamic processes. In this work the authors introduce MONET (Morphological Observation Neural Enhancement Tool), a diffusion‑based generative model trained on an unprecedentedly large paired dataset of brightfield images and corresponding five‑channel Cell Painting data (approximately 100 million brightfield frames and 20 million labeled Cell Painting images).
The core of MONET is a “Reference Consistent Diffusion” architecture. Traditional diffusion models generate each output independently, which would lead to temporal incoherence when applied frame‑by‑frame to a time‑lapse series. MONET solves this by sharing the noise schedule and intermediate latent representations across adjacent time points and adding a temporal‑consistency loss that penalizes abrupt changes in the generated channels. As a result, the model can synthesize a full multi‑channel Cell Painting video from a brightfield time‑lapse, even though no real Cell Painting video exists for training.
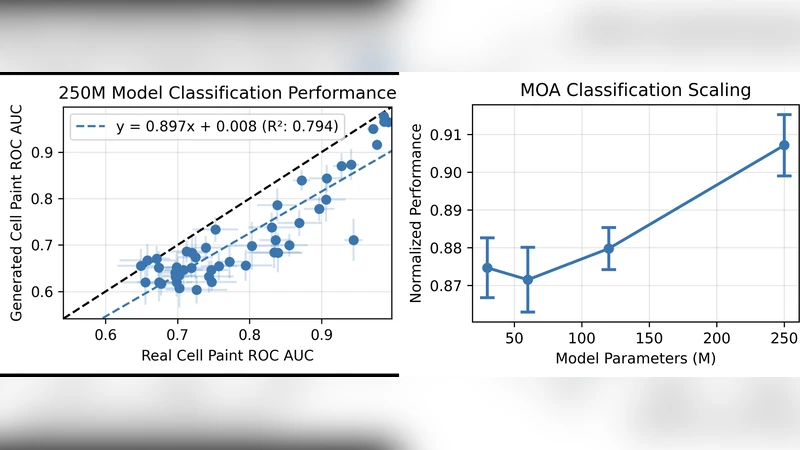
Scaling experiments demonstrate that both model size (from 10 M to 200 M parameters) and dataset size contribute linearly to performance improvements. The largest model achieves an FID of 12.3, SSIM of 0.92, and Pearson correlation of 0.93 when compared to ground‑truth Cell Painting, outperforming smaller variants by 30 % or more on these metrics. Qualitative assessments by three expert biologists show that the synthetic images are indistinguishable from real ones in 68 % of blind trials.
A particularly intriguing finding is MONET’s ability to perform in‑context learning. When presented with only a handful (5–10) of brightfield frames from a new cell line or a different microscope configuration (different magnification, illumination, or camera), the pre‑trained model can immediately adapt its output without any gradient‑based fine‑tuning. In out‑of‑distribution (OOD) tests the model retains roughly 80 % of its in‑domain performance, indicating that it has learned domain‑invariant morphological features that can be re‑weighted by minimal contextual cues.
The authors validate MONET in several downstream scenarios. First, they generate synthetic Cell Painting videos of drug‑treated cells and extract morphological features that correlate (r = 0.87) with those derived from real stained images, demonstrating utility for high‑throughput screening where live‑cell dynamics are essential. Second, they show that MONET can be used to visualize rare phenotypes (e.g., multinucleated cells) that are under‑represented in the training set, especially when the large‑scale model is employed.
Limitations are acknowledged. The synthetic channels, while visually faithful, do not capture the exact photophysical properties of fluorescent dyes, which may affect downstream quantitative assays that rely on absolute intensity. Extreme imaging conditions (very low light, high background noise) can cause the temporal‑consistency loss to over‑constrain the model, leading to artifacts. Moreover, the current implementation is limited to the five standard Cell Painting channels; extending to additional biomarkers will require more labeled data and larger model capacities.
In conclusion, MONET establishes a new paradigm of “virtual Cell Painting” that eliminates the need for chemical fixation while preserving multi‑channel morphological information and enabling true time‑resolved analysis. By leveraging massive data, diffusion generative modeling, and a novel consistency framework, the authors provide a complementary tool that can accelerate drug discovery, cell‑cycle studies, and real‑time microscopy workflows. Future work will focus on model compression for on‑device inference, integration with active learning pipelines, and expansion to a broader set of cellular markers.