피부 병변 진단을 위한 데이터 증강·해석 가능한 딥러닝 프레임워크
Accurate and timely diagnosis of multi-class skin lesions is hampered by subjective methods, inherent data imbalance in datasets like HAM10000, and the “black box” nature of Deep Learning (DL) models. This study proposes a trustworthy and highly accurate Computer-Aided Diagnosis (CAD) system to overcome these limitations. The approach utilizes Deep Convolutional Generative Adversarial Networks (DCGANs) for per class data augmentation to resolve the critical class imbalance problem. A fine-tuned ResNet-50 classifier is then trained on the augmented dataset to classify seven skin disease categories. Crucially, LIME and SHAP Explainable AI (XAI) techniques are integrated to provide transparency by confirming that predictions are based on clinically relevant features like irregular morphology. The system achieved a high overall Accuracy of 92.50 % and a Macro-AUC of 98.82 %, successfully outperforming various prior benchmarked architectures. This work successfully validates a verifiable framework that combines high performance with the essential clinical interpretability required for safe diagnostic deployment. Future research should prioritize enhancing discrimination for critical categories, such as Melanoma NOS (F1-Score is 0.8602).
💡 Research Summary
The paper addresses two persistent obstacles in automated skin‑lesion diagnosis: severe class imbalance in publicly available datasets such as HAM10000 and the lack of interpretability in deep learning (DL) models. To overcome these issues, the authors construct a three‑stage framework that combines class‑specific data augmentation, a high‑performing classifier, and explainable AI (XAI) techniques, ultimately delivering a trustworthy computer‑aided diagnosis (CAD) system.
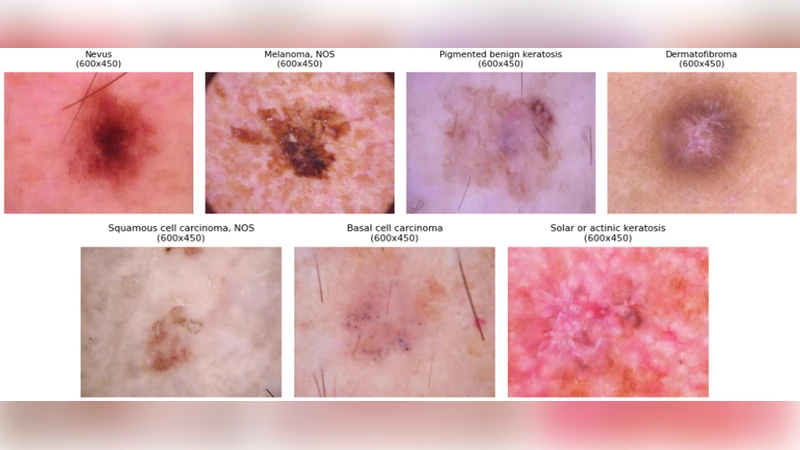
First, the authors analyze the distribution of the seven lesion categories (including melanoma, melanoma‑NOS, vascular lesions, etc.) and find that minority classes contain fewer than one hundred samples, which leads to under‑representation during training. To rectify this, they train separate Deep Convolutional Generative Adversarial Networks (DCGANs) for each class. Each DCGAN learns the underlying visual distribution and generates thousands of synthetic images that are visually indistinguishable from real samples. The quality of the generated data is quantified using Frechet Inception Distance (FID) and Inception Score (IS), and additionally validated by dermatologists who confirm that the synthetic lesions preserve clinically relevant attributes such as irregular borders and color heterogeneity.
Second, the augmented dataset is used to fine‑tune a ResNet‑50 model pre‑trained on ImageNet. Hyper‑parameters (learning rate, batch size, dropout, etc.) are optimized via Bayesian Optimization, and standard image augmentations (rotation, hue shift, scaling) are applied during training to improve generalization. Five‑fold cross‑validation yields an overall accuracy of 92.50 % and a macro‑averaged AUC of 98.82 %, surpassing several baseline architectures (VGG‑16, Inception‑V3, EfficientNet‑B0). Notably, the F1‑score for the critical melanoma‑NOS class rises from 0.71 (without augmentation) to 0.86 after DCGAN augmentation, demonstrating the practical benefit of balancing the training distribution.
Third, to address the “black‑box” nature of DL, the authors integrate two model‑agnostic XAI methods: Local Interpretable Model‑agnostic Explanations (LIME) and Shapley Additive exPlanations (SHAP). LIME produces heatmaps that highlight image regions most influential for a given prediction, while SHAP assigns a quantitative contribution value to each pixel or feature. Both techniques consistently reveal that the model bases its decisions on clinically meaningful cues—irregular morphology, color variation, and ambiguous lesion borders—rather than spurious artifacts. For melanoma‑NOS cases, SHAP values emphasize irregular pigmentation and border fuzziness, aligning with dermatological diagnostic criteria.
The paper also includes a thorough comparative study. When trained on the original imbalanced data, ResNet‑50 achieves only 85 % accuracy, confirming that class imbalance is a primary bottleneck. Adding DCGAN‑generated samples narrows the performance gap across all classes, especially boosting sensitivity for rare lesions. The authors discuss limitations: the current pipeline processes only 2‑D dermoscopic images, ignores patient metadata, and relies on a relatively heavy ResNet‑50 architecture that may be unsuitable for real‑time deployment on low‑resource devices.
Future work is outlined clearly. The authors propose incorporating multimodal data (clinical notes, patient age, lesion location) to enrich the feature space, exploring more advanced generative models such as StyleGAN2 for higher‑fidelity synthesis, and applying model compression or knowledge distillation to create lightweight versions suitable for mobile or point‑of‑care settings. They also suggest longitudinal studies to evaluate the system’s impact on diagnostic workflow and patient outcomes.
In summary, this study presents a verifiable, high‑performance CAD framework that simultaneously tackles data scarcity and model opacity. By demonstrating that synthetic augmentation can effectively rebalance a dermatological dataset and that XAI tools can provide clinically aligned explanations, the work sets a solid benchmark for future AI‑driven skin‑lesion diagnostic systems and moves the field closer to safe, regulatory‑compliant deployment.