Simple Agents Outperform Experts in Biomedical Imaging Workflow Optimization
Adapting production-level computer vision tools to bespoke scientific datasets is a critical “last mile” bottleneck. Current solutions are impractical: fine-tuning requires large annotated datasets scientists often lack, while manual code adaptation costs scientists weeks to months of effort. We consider using AI agents to automate this manual coding, and focus on the open question of optimal agent design for this targeted task. We introduce a systematic evaluation framework for agentic code optimization and use it to study three production-level biomedical imaging pipelines. We demonstrate that a simple agent framework consistently generates adaptation code that outperforms human-expert solutions. Our analysis reveals that common, complex agent architectures are not universally beneficial, leading to a practical roadmap for agent design. We open source our framework and validate our approach by deploying agent-generated functions into a production pipeline, demonstrating a clear pathway for real-world impact.
💡 Research Summary
The paper tackles a pervasive “last‑mile” bottleneck in biomedical imaging: adapting high‑performance, production‑level computer‑vision tools to the idiosyncratic data generated in individual labs. Traditional remedies—large‑scale fine‑tuning or manual code engineering—are either data‑inefficient or labor‑intensive, often requiring weeks to months of expert effort. The authors propose a radically simpler solution: an LLM‑driven code‑generation agent that automatically writes the necessary preprocessing and post‑processing functions, evaluates them on a small gold‑standard validation set (10–100 images), and iteratively refines the code based on feedback.
To answer the open question of how complex an agent must be for this narrowly scoped task, the authors design a systematic evaluation framework. The “Base Agent” consists of three prompt components (Task Prompt, Data Prompt describing the imaging modality and channel semantics, and an API List of 98 relevant functions from OpenCV, scikit‑image, and SciPy) and two functional modules: a Coding Agent (the LLM that emits Python code) and an Execution Agent (which injects the generated code into the target pipeline, runs it, and returns a performance score). This minimal architecture already provides the context needed for the LLM to produce functional code.
The study then explores four axes of augmentation: (1) LLM type (GPT‑4.1, a reasoning‑oriented model “o3”, and a smaller open‑source Llama 3.3‑70B‑Instruct‑Turbo); (2) inclusion of expert‑written functions as in‑context examples; (3) a Function Bank that stores previously generated functions and feeds the top‑3 and bottom‑3 back into the prompt; and (4) an AutoML component that periodically performs hyper‑parameter search on the best‑scoring functions. For each configuration, the authors run 20 random seeds, each producing 60 trials (20 iterations × 3 function pairs), and finally select the top‑15 functions across runs to evaluate on a held‑out test set, thereby mitigating over‑fitting.
Three representative biomedical imaging pipelines are used as case studies, covering molecular (Polaris spot detection), cellular (Cellpose segmentation), and macroscopic (MedSAM medical‑image segmentation) scales. Expert baselines are taken from the official, production‑optimized codebases of the original tool authors—code that required weeks to months of manual tuning, as quantified via Git history and code‑complexity analysis. The validation objectives are: (i) F1 score for Polaris (95 images), (ii) average precision at IoU 0.5 for Cellpose (100 images), and (iii) the sum of Normalized Surface Dice and Dice Similarity Coefficient for MedSAM (25 images).
Results are striking. Even the simplest Base Agent outperforms the expert baselines across all three tasks: Polaris F1 improves from 0.841 to 0.867, Cellpose AP@IoU 0.5 from 0.402 to 0.409, and MedSAM NSD+DSC from 0.820 to 0.971. The largest gain appears in MedSAM, where the agent‑generated pipeline achieves a 0.151 absolute increase. When the authors replace the LLM with the smaller Llama 3.3‑70B model, performance drops below the expert baseline, highlighting the importance of model capacity for this code‑generation problem.
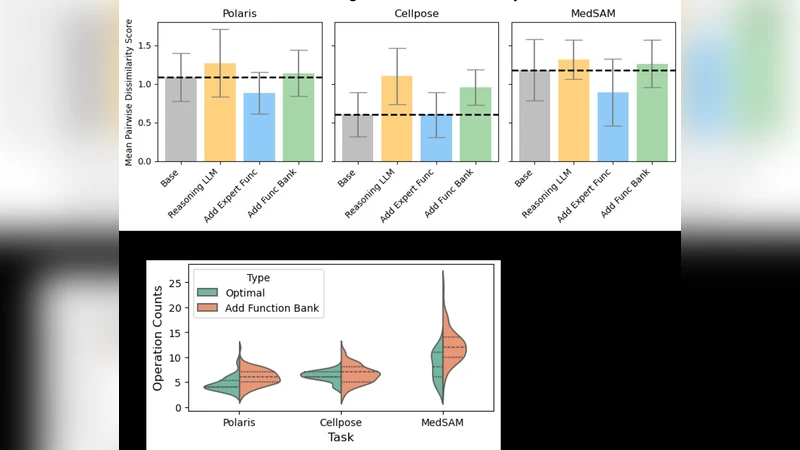
The impact of the additional components is mixed. Adding expert functions dramatically helps Polaris (F1 0.929) but yields modest or neutral gains for Cellpose and MedSAM. The Function Bank improves MedSAM (up to 1.037) but slightly harms Polaris and Cellpose. The AutoML hyper‑parameter tuner can push scores higher (e.g., MedSAM 1.037) but adds computational overhead and does not consistently beat the Base Agent alone. Overall, the study reveals that more sophisticated architectures—hierarchical planning, large tool‑spaces, or extensive memory—are not universally beneficial for targeted tool adaptation. Simplicity, combined with a well‑specified data and API context and a capable LLM, suffices to generate high‑quality adaptation code within 1–2 days of compute time, saving weeks or months of human effort.
Beyond empirical findings, the paper contributes a publicly released framework (GitHub) that enables other researchers to reproduce the experiments, extend the design space, or plug the agent into their own pipelines. The authors also demonstrate a real‑world deployment: an agent‑generated function was merged via an anonymous GitHub pull request into the official codebase of one of the studied tools, confirming that the approach can move from research to production.
In conclusion, the work establishes that for the specific problem of adapting pre‑trained biomedical imaging tools to new datasets, a “simple agent”—essentially a prompt‑rich LLM with a code‑execution feedback loop—is not only sufficient but often superior to expert‑crafted solutions. The systematic analysis provides a practical roadmap: start with a minimal Base Agent, select a strong LLM, and only add auxiliary components when task‑specific evidence suggests benefit. This insight reshapes how the community should think about AI‑agent design for low‑data, high‑impact scientific workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment