📝 Original Info
- Title: ForamDeepSlice: A High-Accuracy Deep Learning Framework for Foraminifera Species Classification from 2D Micro-CT Slices
- ArXiv ID: 2512.00912
- Date: 2025-11-30
- Authors: Researchers from original ArXiv paper
📝 Abstract
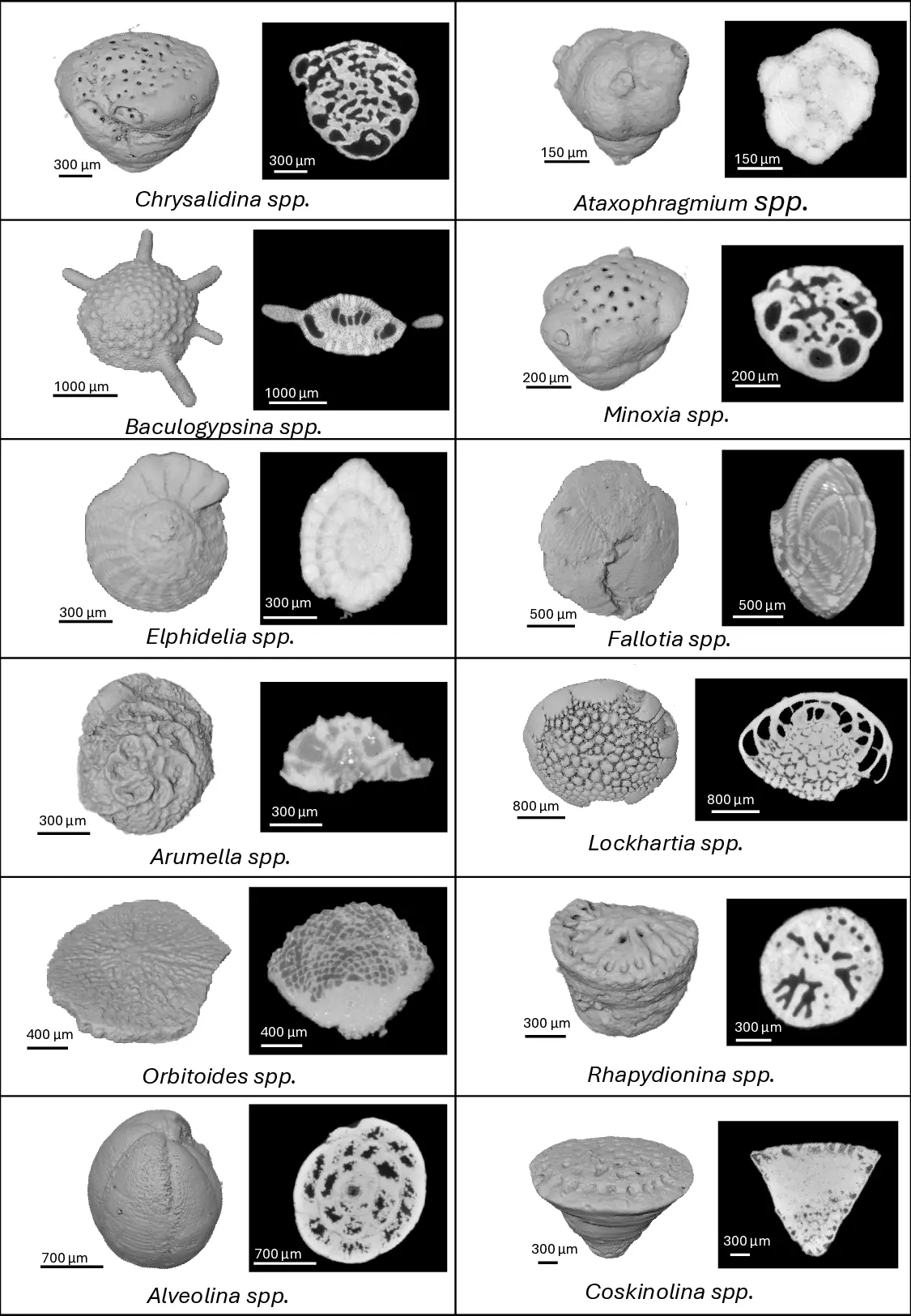
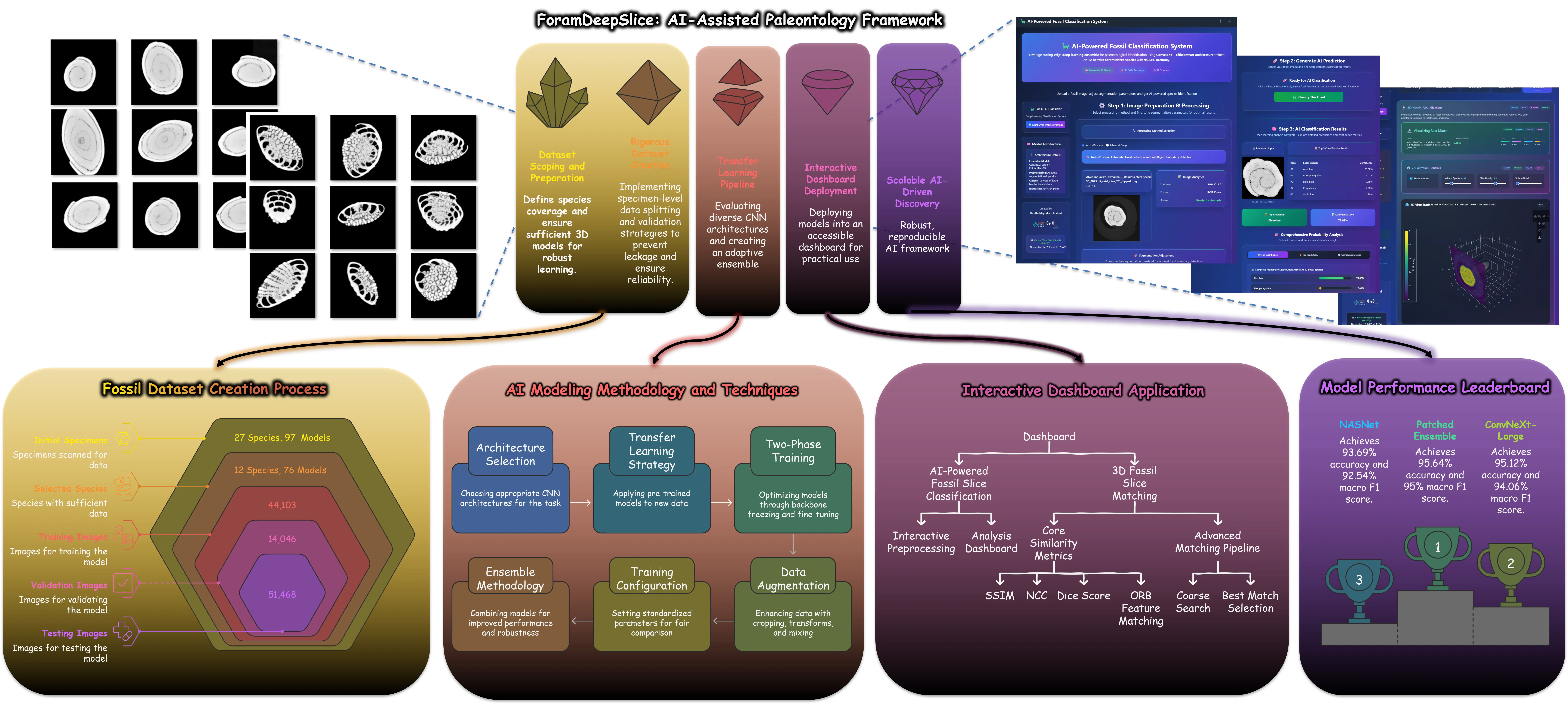
This study presents a comprehensive deep learning pipeline for the automated classification of 12 foraminifera species using 2D micro-CT slices derived from 3D scans. We curated a scientifically rigorous dataset comprising 97 micro-CT scanned specimens across 27 species, selecting 12 species with sufficient representation for robust machine learning. To ensure methodological integrity and prevent data leakage, we employed specimen-level data splitting, resulting in 109,617 high-quality 2D slices (44,103 for training, 14,046 for validation, and 51,468 for testing). We evaluated seven state-of-the-art 2D convolutional neural network (CNN) architectures using transfer learning. Our final ensemble model, combining ConvNeXt-Large and EfficientNetV2-Small, achieved a test accuracy of 95.64%, with a top-3 accuracy of 99.6% and an area under the ROC curve (AUC) of 0.998 across all species. To facilitate practical deployment, we developed an interactive advanced dashboard that supports real-time slice classification and 3D slice matching using advanced similarity metrics, including SSIM, NCC, and the Dice coefficient. This work establishes new benchmarks for AI-assisted micropaleontological identification and provides a fully reproducible framework for foraminifera classification research, bridging the gap between deep learning and applied geosciences.
💡 Deep Analysis
Deep Dive into ForamDeepSlice: A High-Accuracy Deep Learning Framework for Foraminifera Species Classification from 2D Micro-CT Slices.
This study presents a comprehensive deep learning pipeline for the automated classification of 12 foraminifera species using 2D micro-CT slices derived from 3D scans. We curated a scientifically rigorous dataset comprising 97 micro-CT scanned specimens across 27 species, selecting 12 species with sufficient representation for robust machine learning. To ensure methodological integrity and prevent data leakage, we employed specimen-level data splitting, resulting in 109,617 high-quality 2D slices (44,103 for training, 14,046 for validation, and 51,468 for testing). We evaluated seven state-of-the-art 2D convolutional neural network (CNN) architectures using transfer learning. Our final ensemble model, combining ConvNeXt-Large and EfficientNetV2-Small, achieved a test accuracy of 95.64%, with a top-3 accuracy of 99.6% and an area under the ROC curve (AUC) of 0.998 across all species. To facilitate practical deployment, we developed an interactive advanced dashboard that supports real-tim
📄 Full Content
ForamDeepSlice: A High-Accuracy Deep Learning Framework
for Foraminifera Species Classification from 2D Micro-CT
Slices
A Preprint
Abdelghafour Halimi
Visualization Core Lab
Thuwal, 23955, Saudi Arabia
abdelghafour.halimi@kaust.edu.sa
Ali Alibrahim
Physical Sciences and Engineering Division
Thuwal, 23955, Saudi Arabia
ali.alibrahim@kaust.edu.sa
Didier Barradas-Bautista
Visualization Core Lab
Thuwal, 23955, Saudi Arabia
didier.barradas@kaust.edu.sa
Ronell Sicat
Visualization Core Lab
Thuwal, 23955, Saudi Arabia
ronell.sicat@kaust.edu.sa
Abdulkader M. Afifi
Physical Sciences and Engineering Division
Thuwal, 23955, Saudi Arabia
abdulkader.alafifi@kaust.edu.sa
December 2, 2025
Abstract
This study presents a comprehensive deep learning pipeline for the automated classification of 12
foraminifera species using 2D micro-CT slices derived from 3D scans. We curated a scientifically
rigorous dataset comprising 97 micro-CT scanned specimens across 27 species, selecting 12 species
with sufficient representation for robust machine learning. To ensure methodological integrity and
prevent data leakage, we employed specimen-level data splitting, resulting in 109,617 high-quality
2D slices (44,103 for training, 14,046 for validation, and 51,468 for testing). We evaluated seven
state-of-the-art 2D convolutional neural network (CNN) architectures using transfer learning. Our final
ensemble model, combining ConvNeXt-Large and EfficientNetV2-Small, achieved a test accuracy of
95.64%, with a top-3 accuracy of 99.6% and an area under the ROC curve (AUC) of 0.998 across
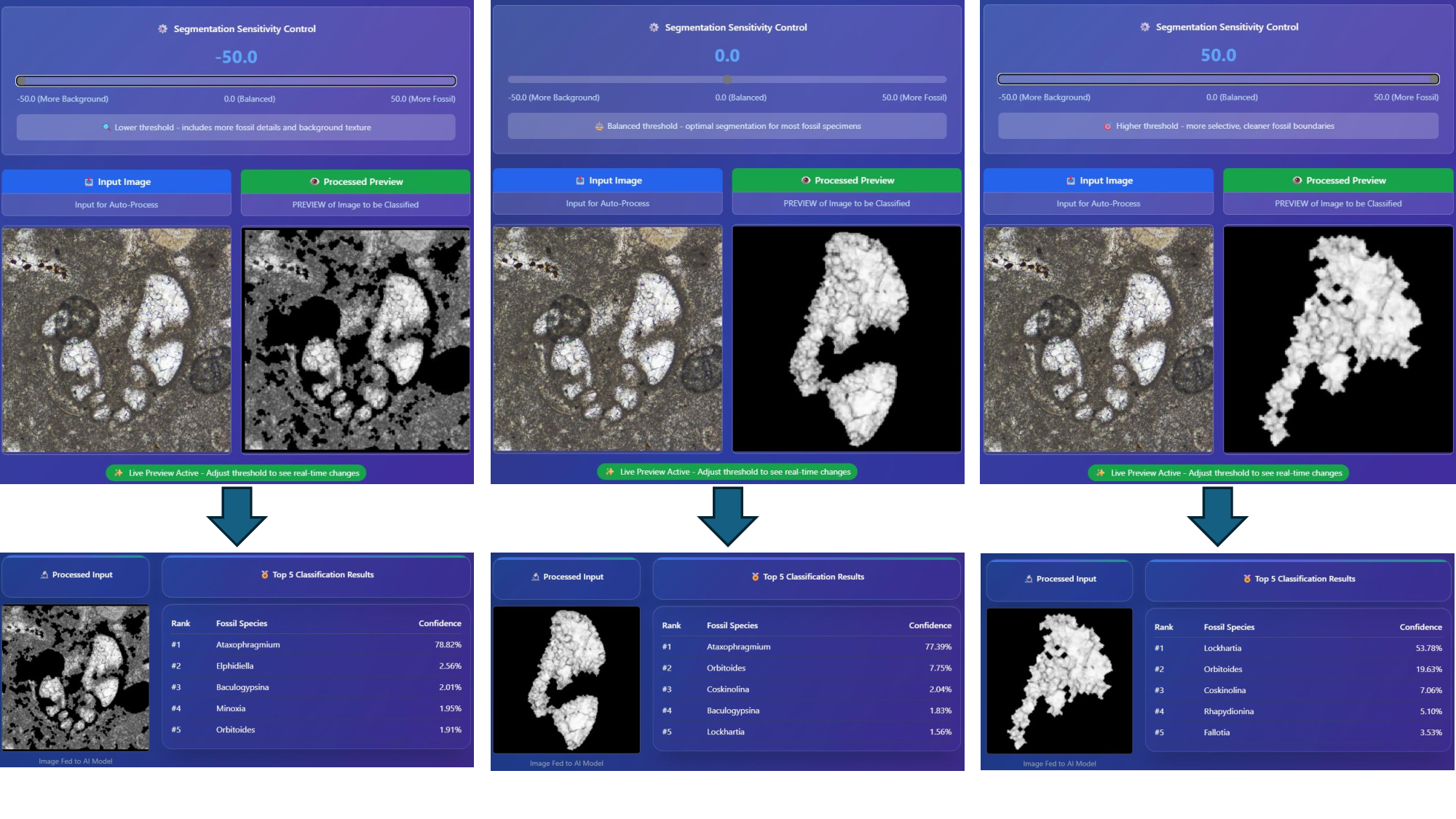
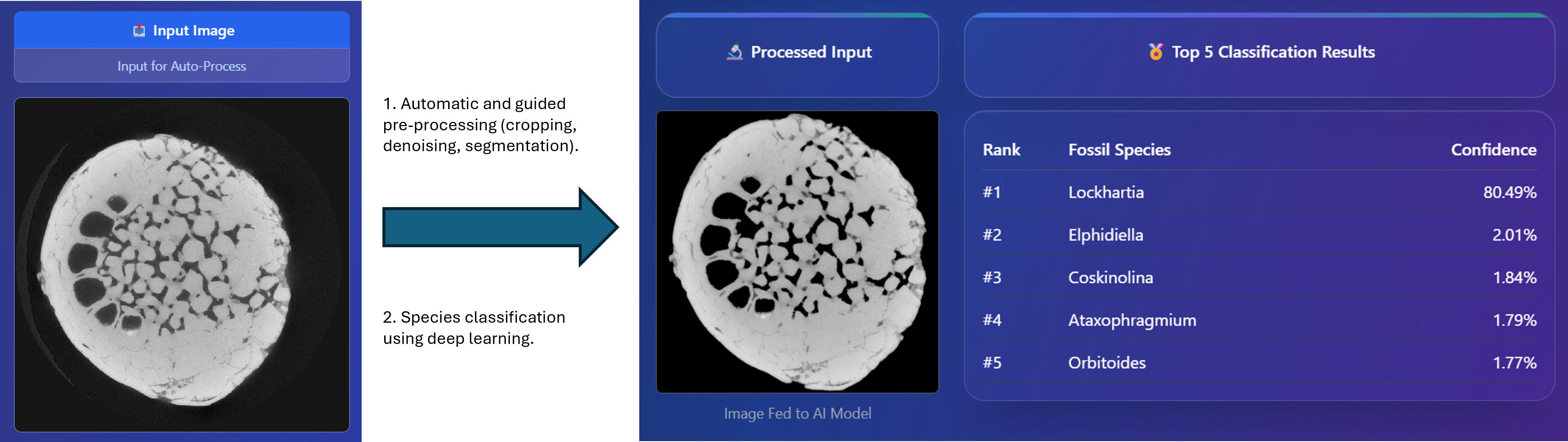
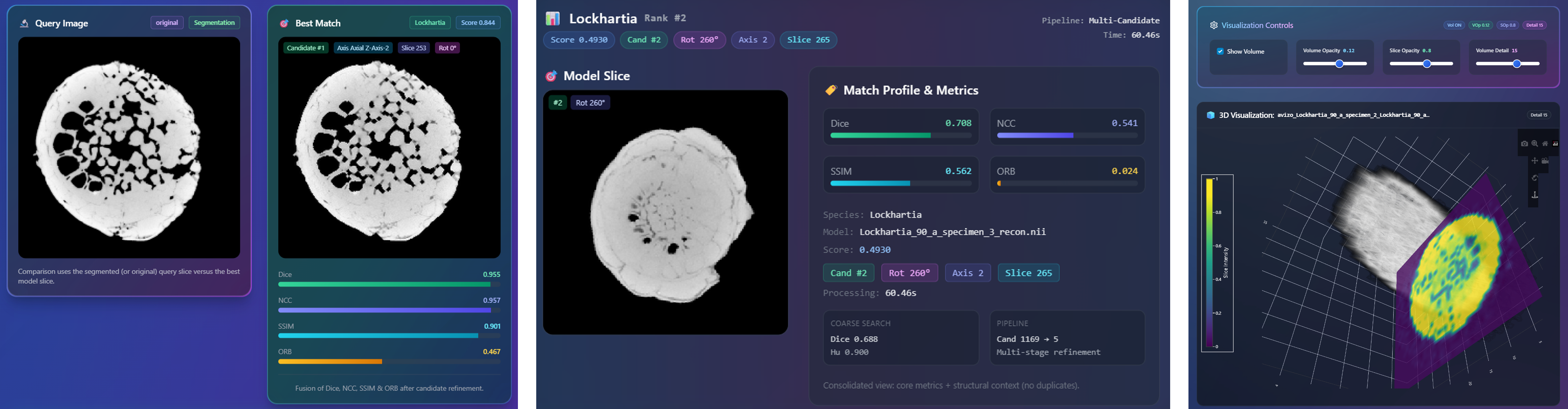
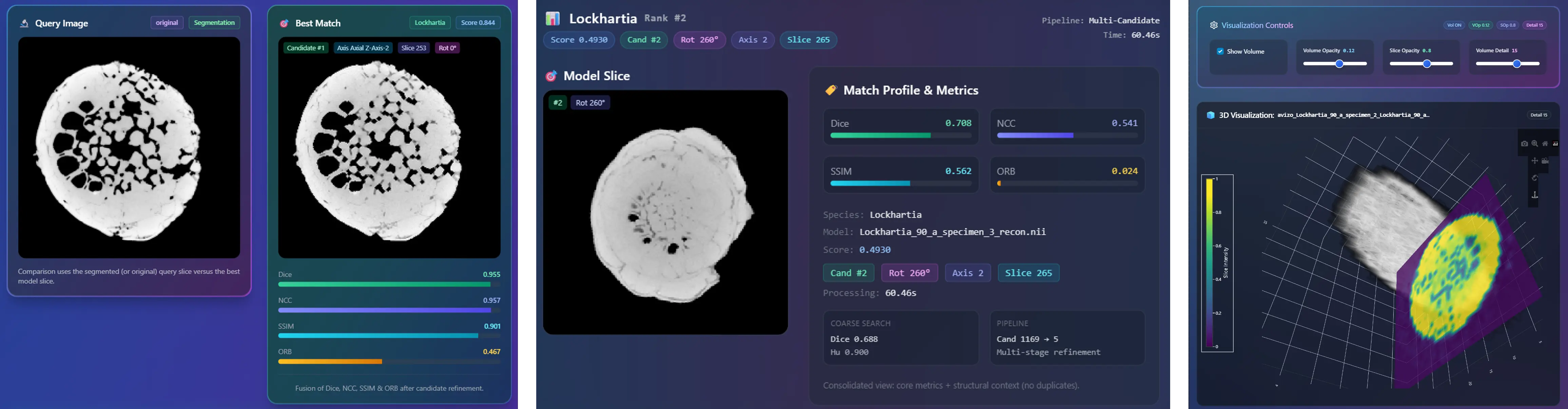
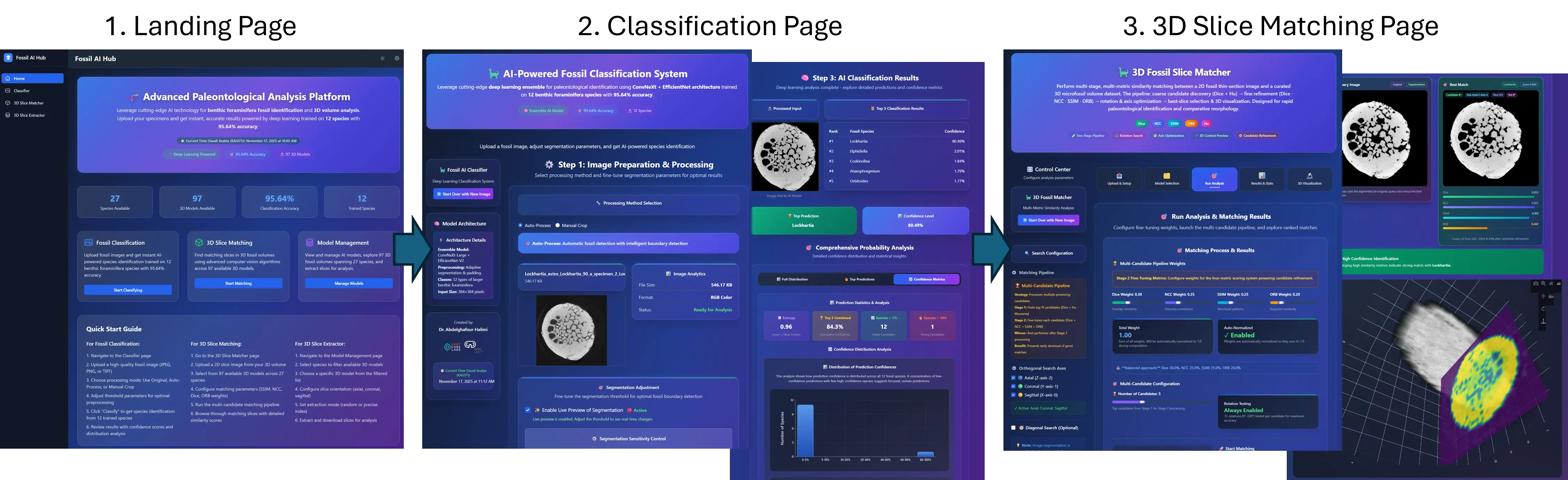
all species. To facilitate practical deployment, we developed an interactive advanced dashboard
that supports real-time slice classification and 3D slice matching using advanced similarity metrics,
including SSIM, NCC, and the Dice coefficient. This work establishes new benchmarks for AI-assisted
micropaleontological identification and provides a fully reproducible framework for foraminifera
classification research, bridging the gap between deep learning and applied geosciences.
Keywords foraminifera classification, deep learning, transfer learning, 2D slices, 3D micro-CT data
1
Introduction
Paleontology is undergoing a transformative data revolution, catalyzed by the widespread adoption of high-resolution,
non-destructive imaging technologies such as micro-Computed Tomography (micro-CT) [Knutsen and Konovalov,
2024, Edie et al., 2023]. These imaging methods have become indispensable for exploring the internal morphology of
fossil specimens, offering unprecedented access to delicate anatomical structures that would otherwise be compromised
by traditional mechanical preparation techniques [Heřmanová et al., 2020, Cunningham et al., 2014]. However, this
arXiv:2512.00912v1 [cs.CV] 30 Nov 2025
Foraminifera Classification using Deep Learning
A Preprint
technological leap has introduced a significant bottleneck in post-processing: the manual segmentation of fossils from
surrounding rock matrices. As imaging resolutions increase, the volumetric datasets grow exponentially, making
segmentation—especially for low-contrast specimens like calcareous fossils in carbonate-rich matrices—the most
labor-intensive and error-prone step in the workflow [Edie et al., 2023].
To address this challenge, the field has increasingly turned to deep learning, a subset of machine learning that excels in
automated image analysis. Early applications demonstrated that convolutional neural networks (CNNs), particularly
U-Net architectures, could segment fossils with accuracies rivaling manual annotation, but in a fraction of the time
[Knutsen and Konovalov, 2024, Edie et al., 2023, Itaki et al., 2020a]. These breakthroughs have laid the foundation for
a broader integration of artificial intelligence (AI) into paleontological research, extending beyond segmentation to
include fossil classification and reconstruction, as summarized in Table 1.
One of the most promising directions in this domain is the classification of microfossils using transfer learning. Several
studies have shown that pretrained CNN architectures—such as VGG16, ResNet50, MobileNetV2, InceptionV3, and
Xception—can be fine-tuned to classify microfossils from low-resolution optical images with remarkable accuracy. For
instance, classification of the genus Globotruncanita achieved up to 96.97% accuracy and an AUC of 0.978 [Ozer et al.,
2024], while broader genus-level classification reached 99.78% accuracy [Ozer et al., 2023a]. These approaches are
particularly valuable for laboratories with limited computational resources, offering high performance with minimal
infrastructure.
Beyond transfer learning, researchers have explored custom architectures built from scratch, combining CNNs with
recurrent neural networks (LSTM, BiLSTM) to classify species within the genus Globotruncana, achieving up to
97.35% accuracy and an AUC of 0.968 [Ozer et al., 2023b]. Classical machine learning models such as SVM, LDA, and
kNN have also demonstrat
…(Full text truncated)…
📸 Image Gallery
Reference
This content is AI-processed based on ArXiv data.