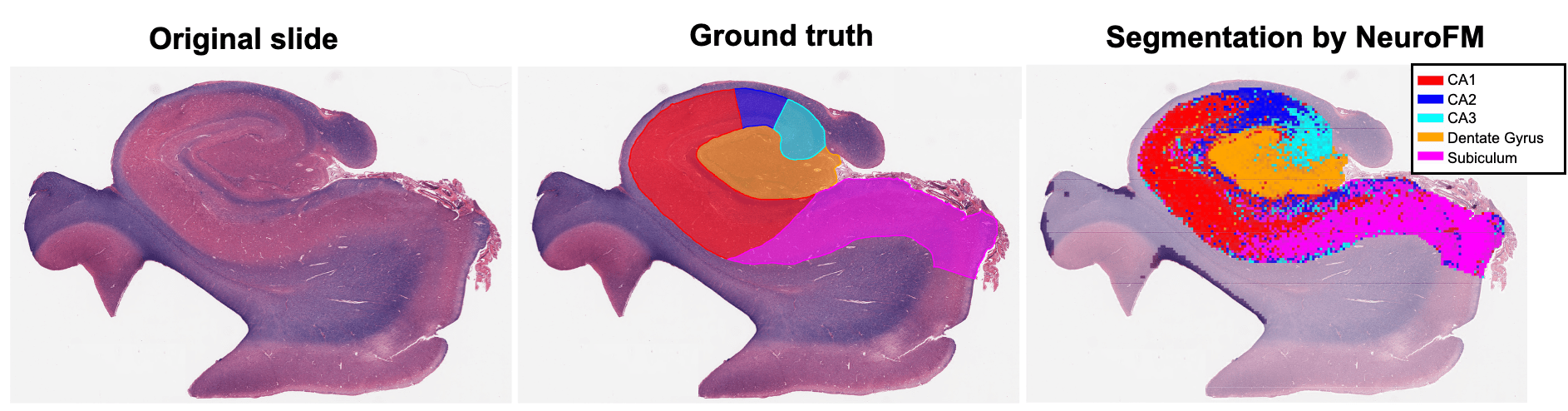
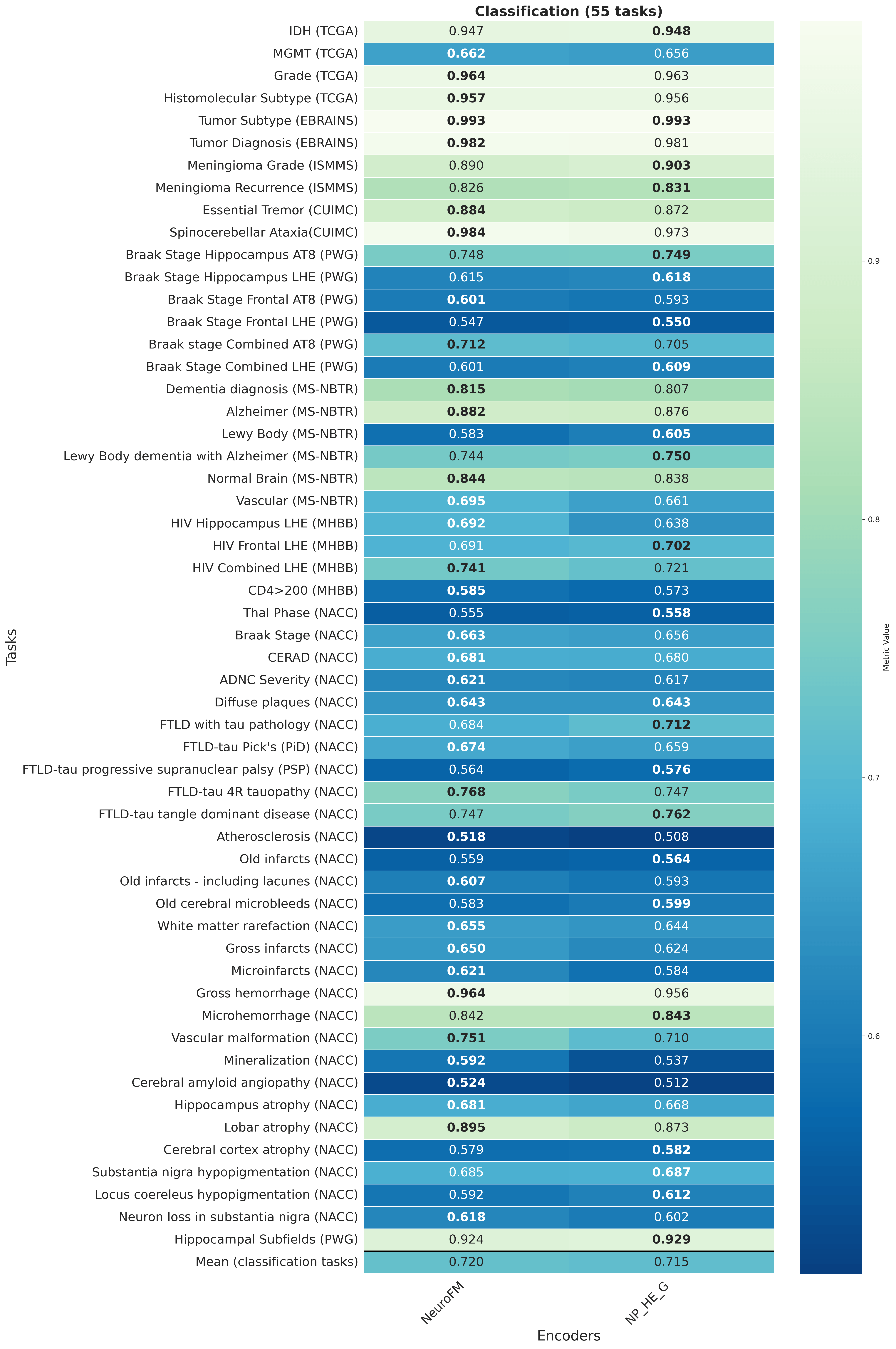
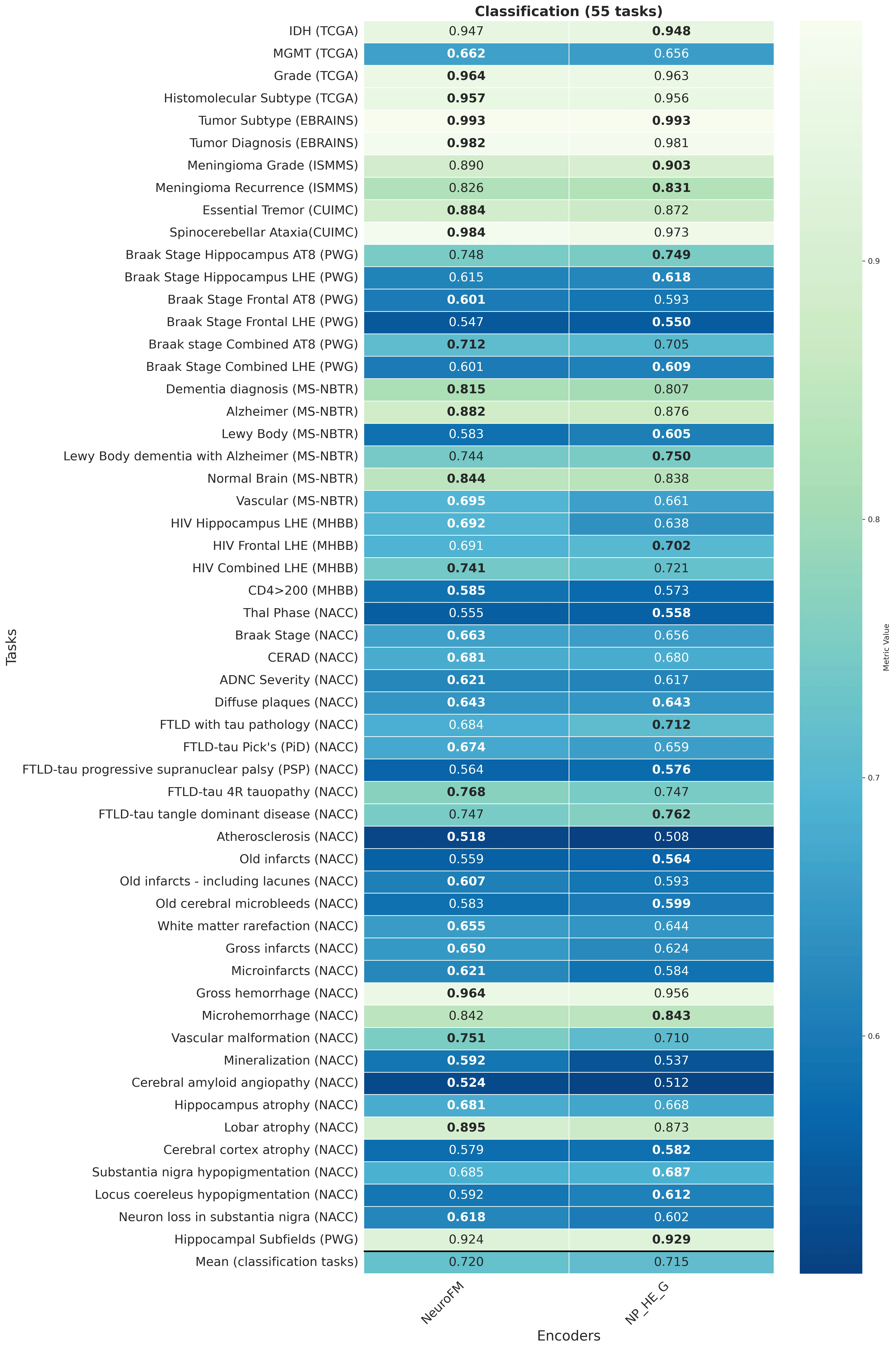
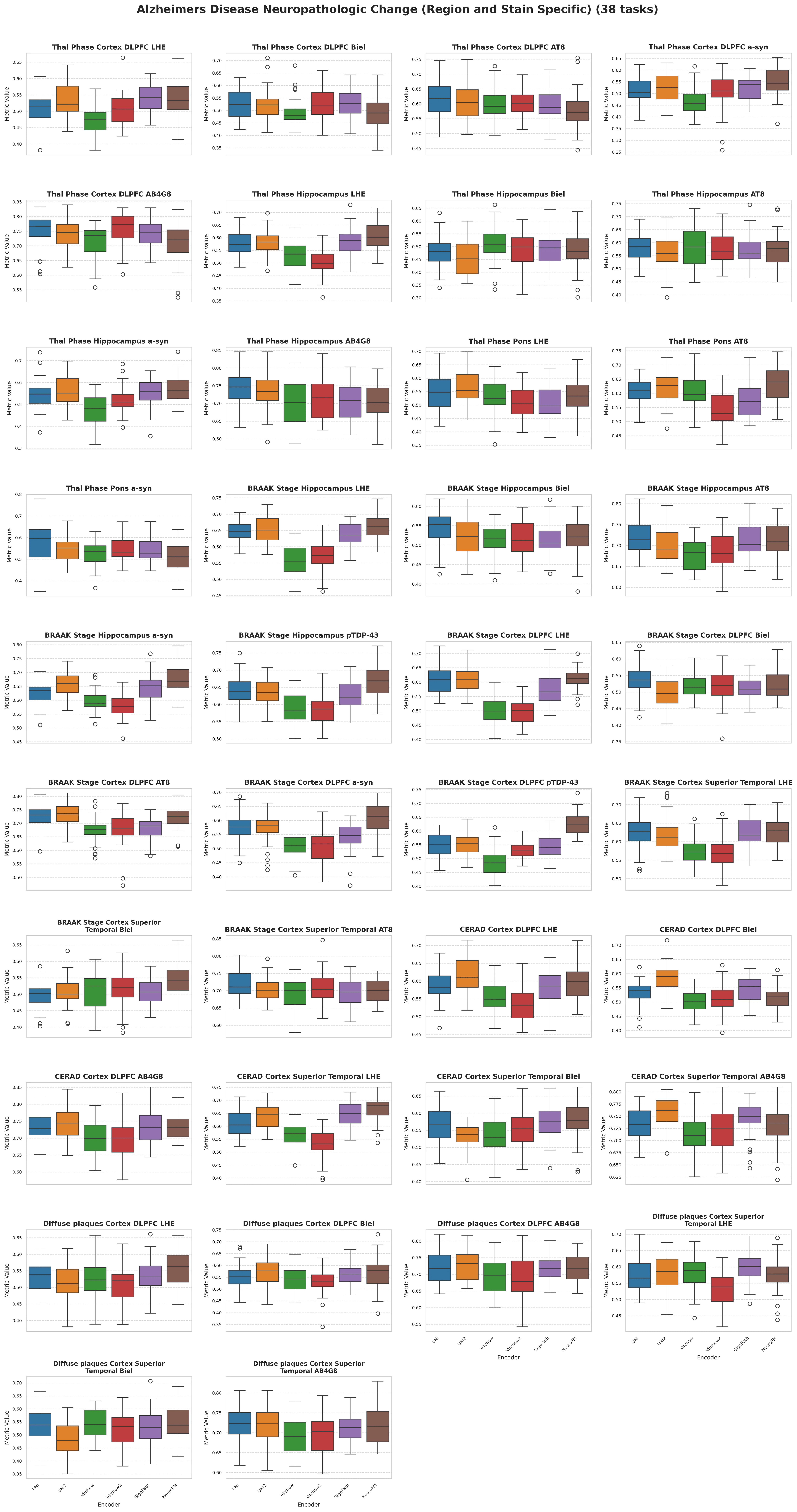
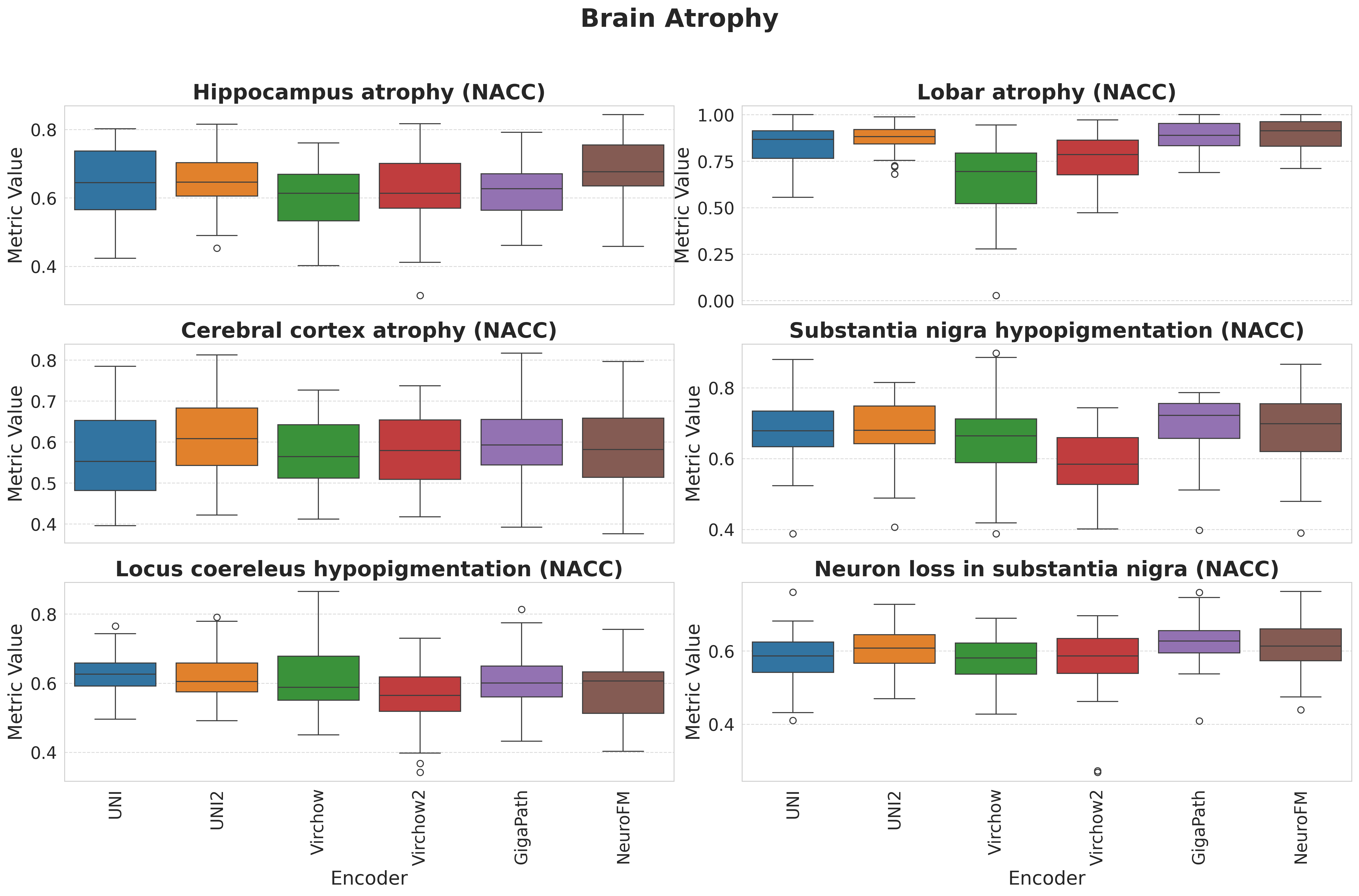
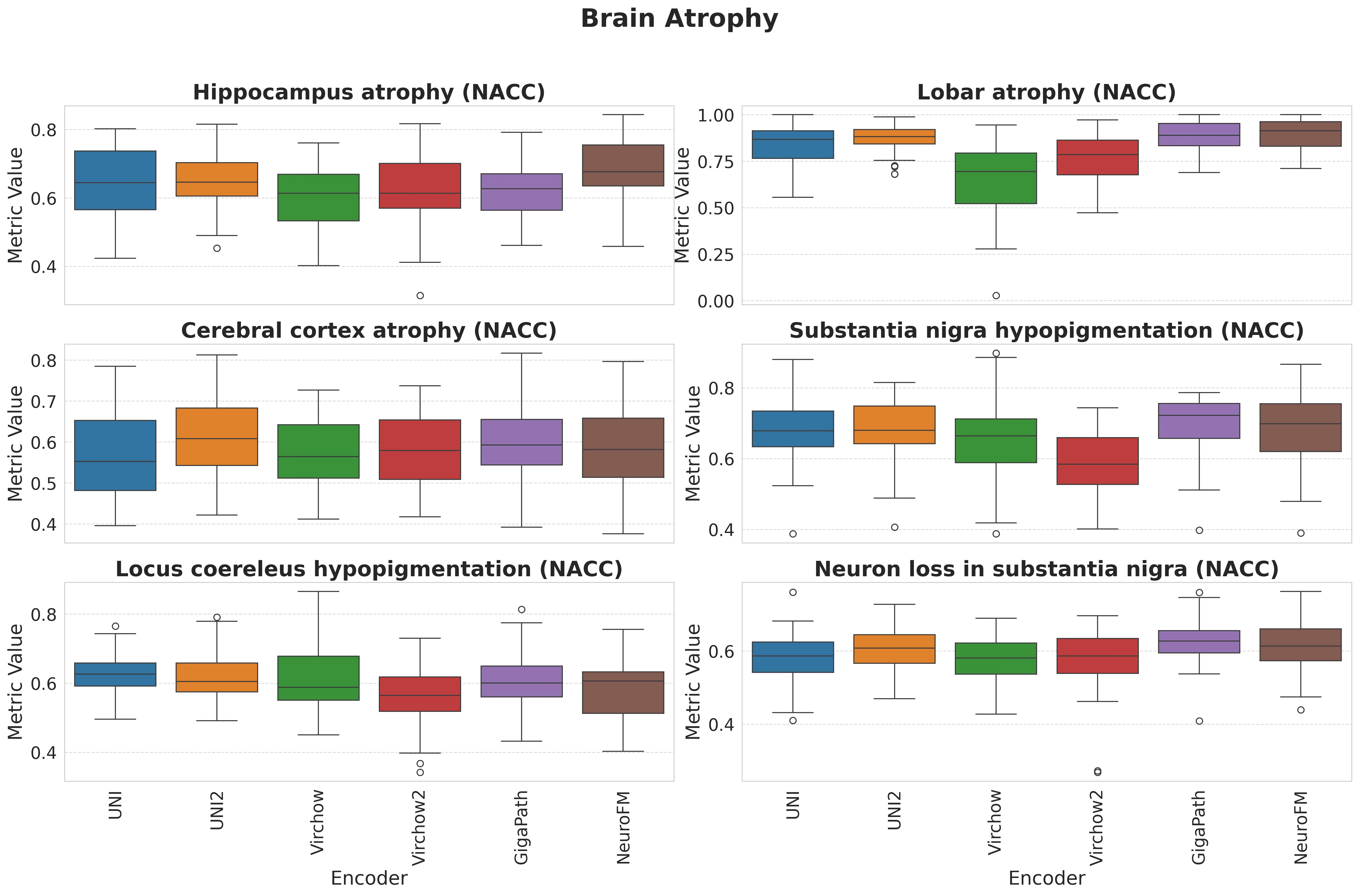
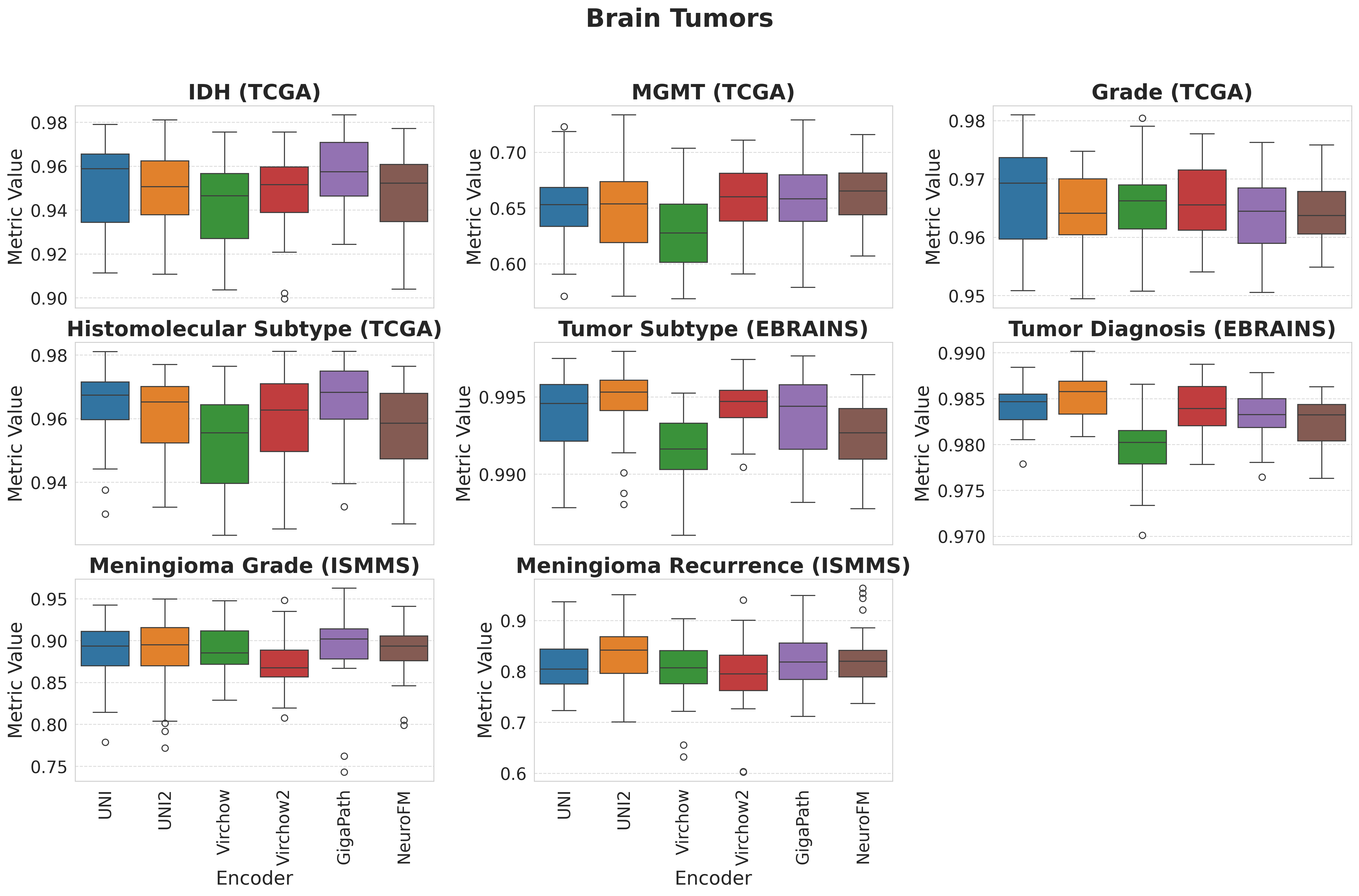
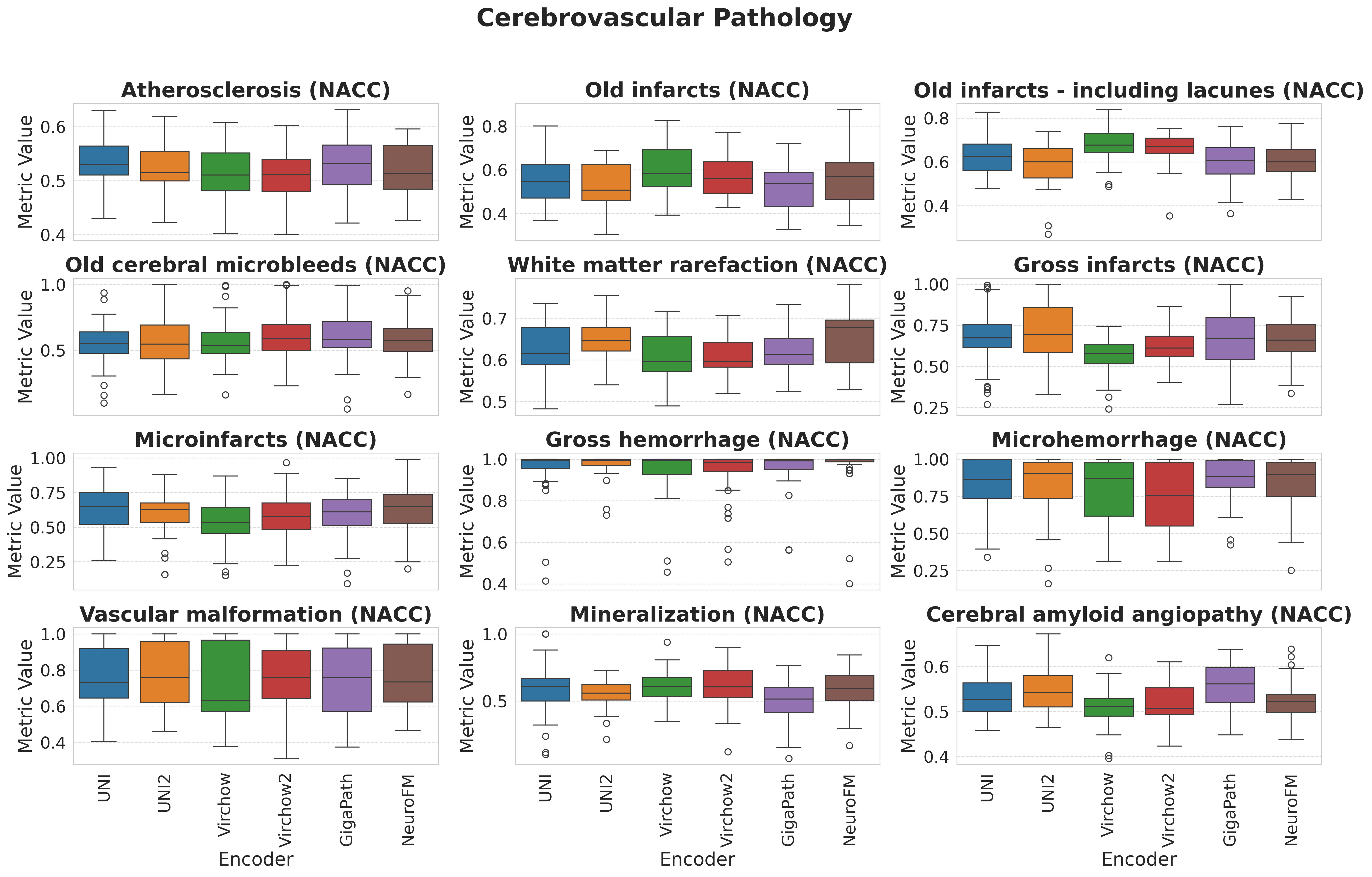
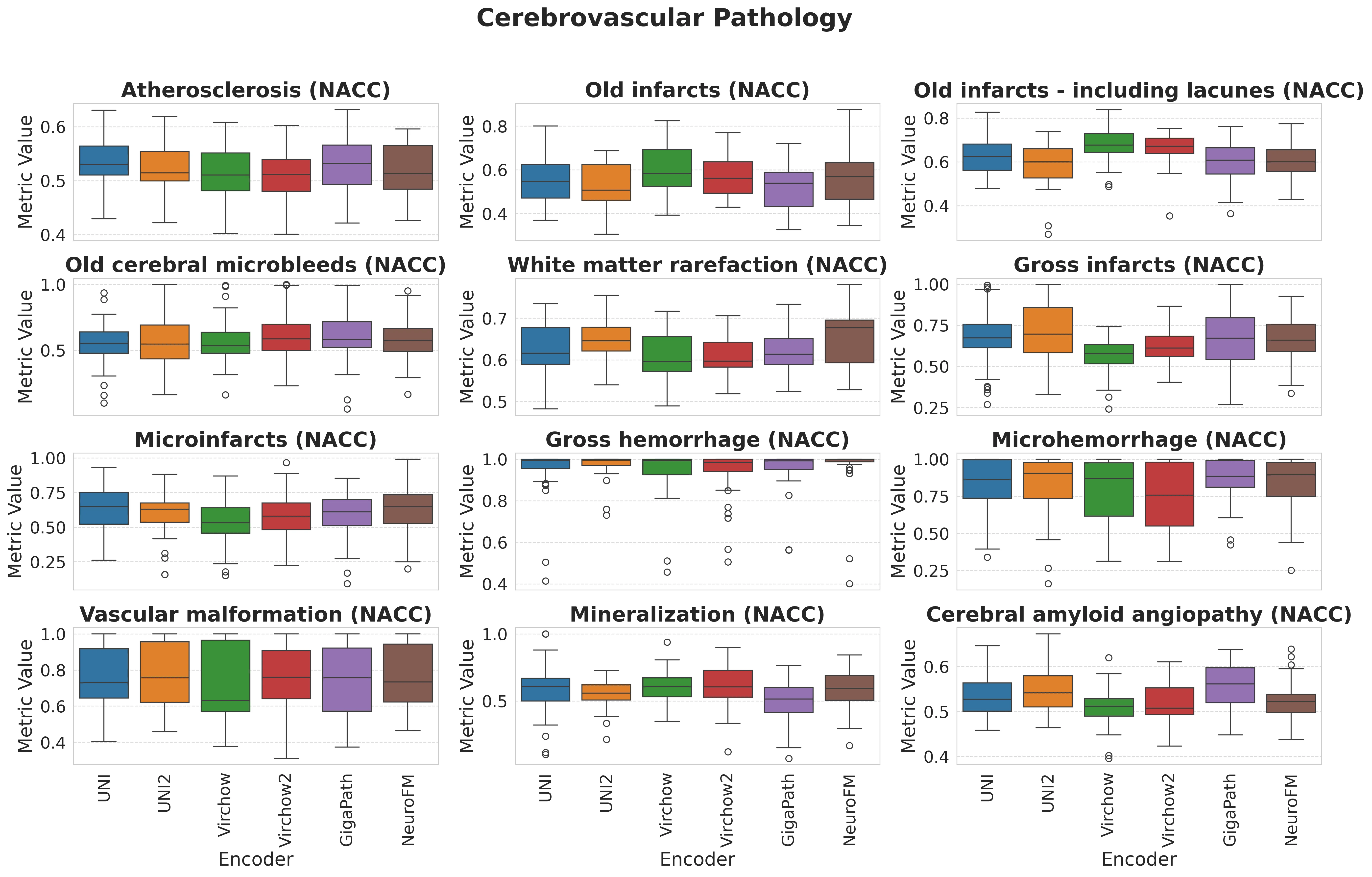
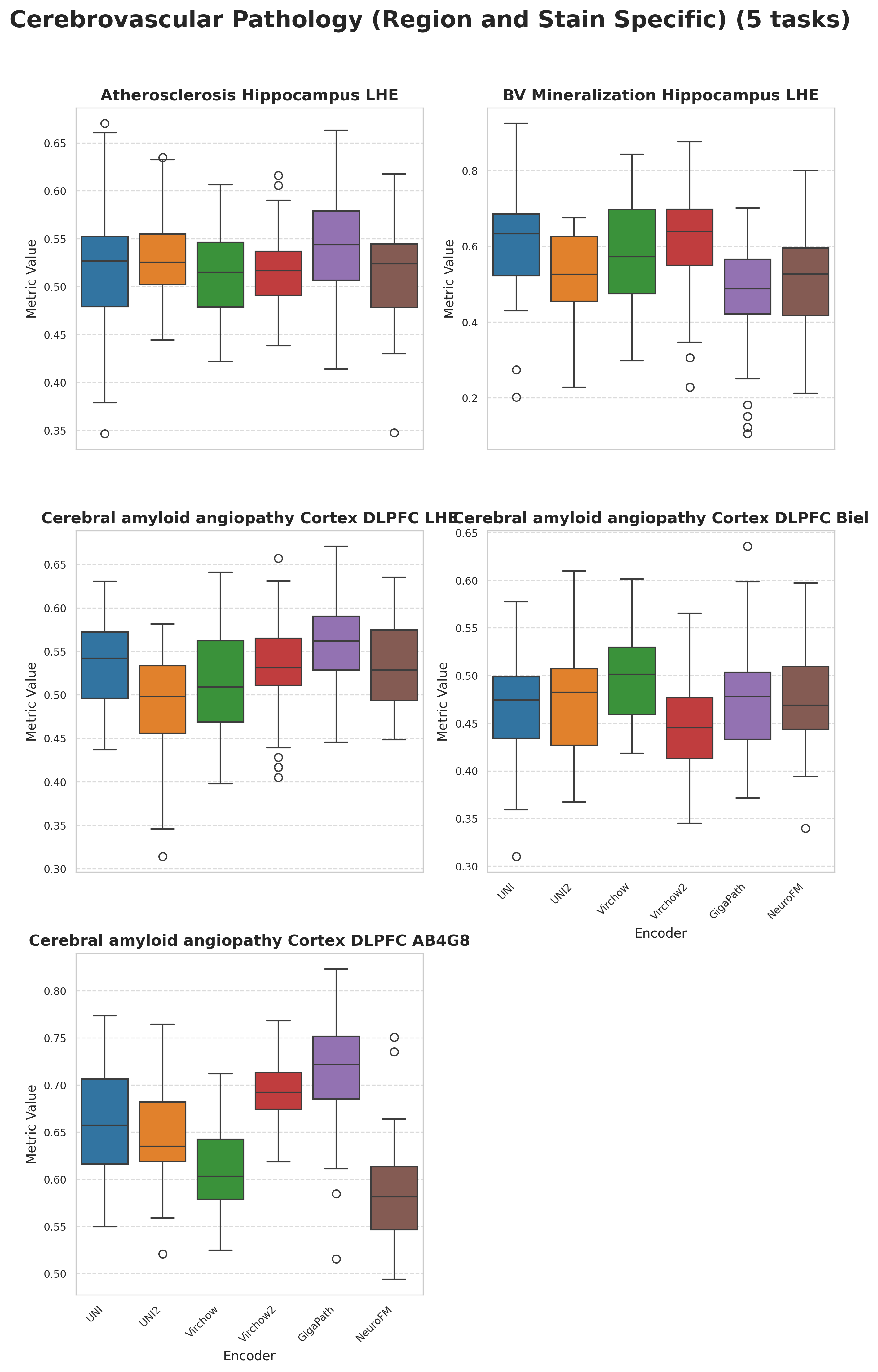
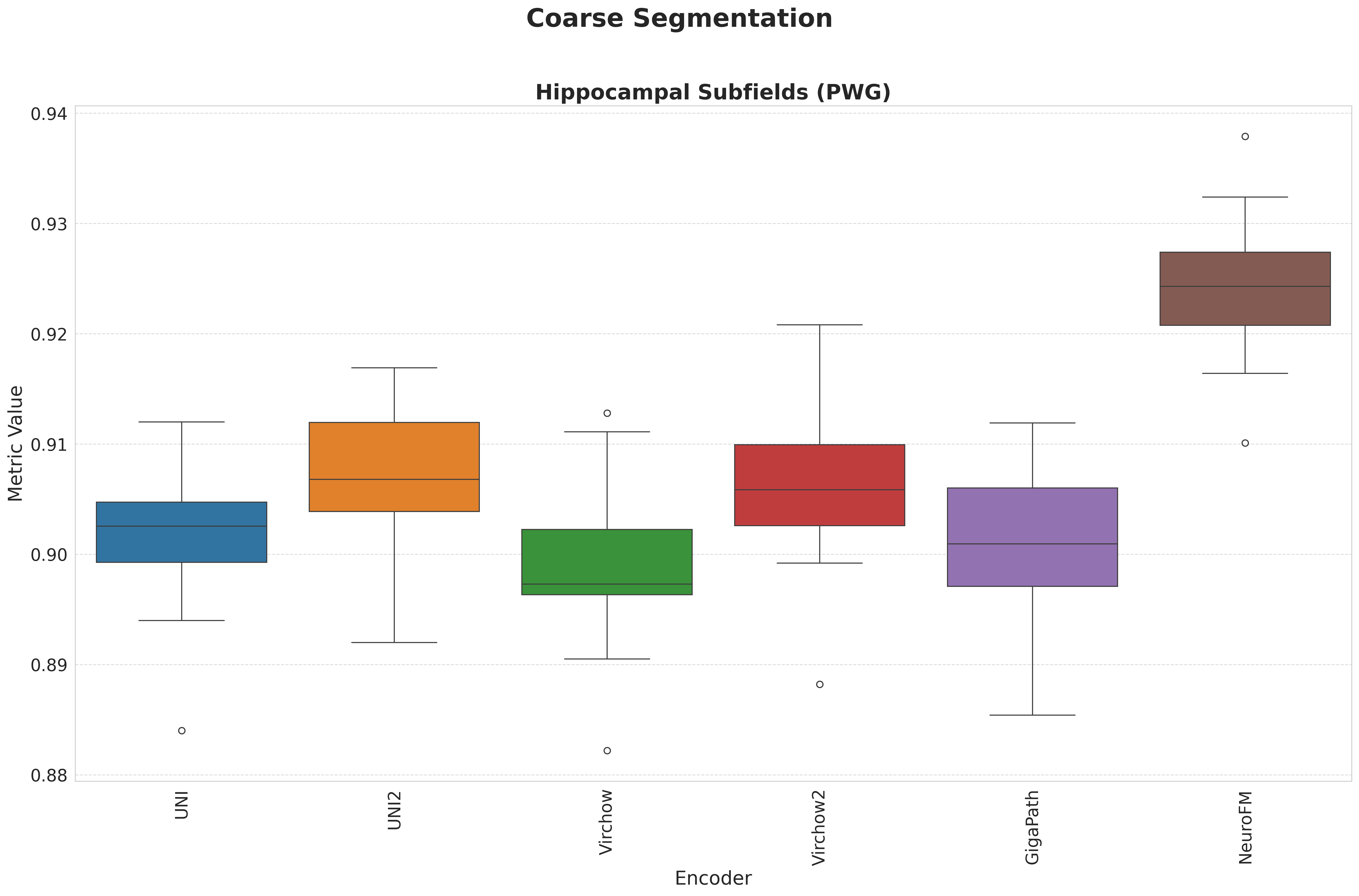
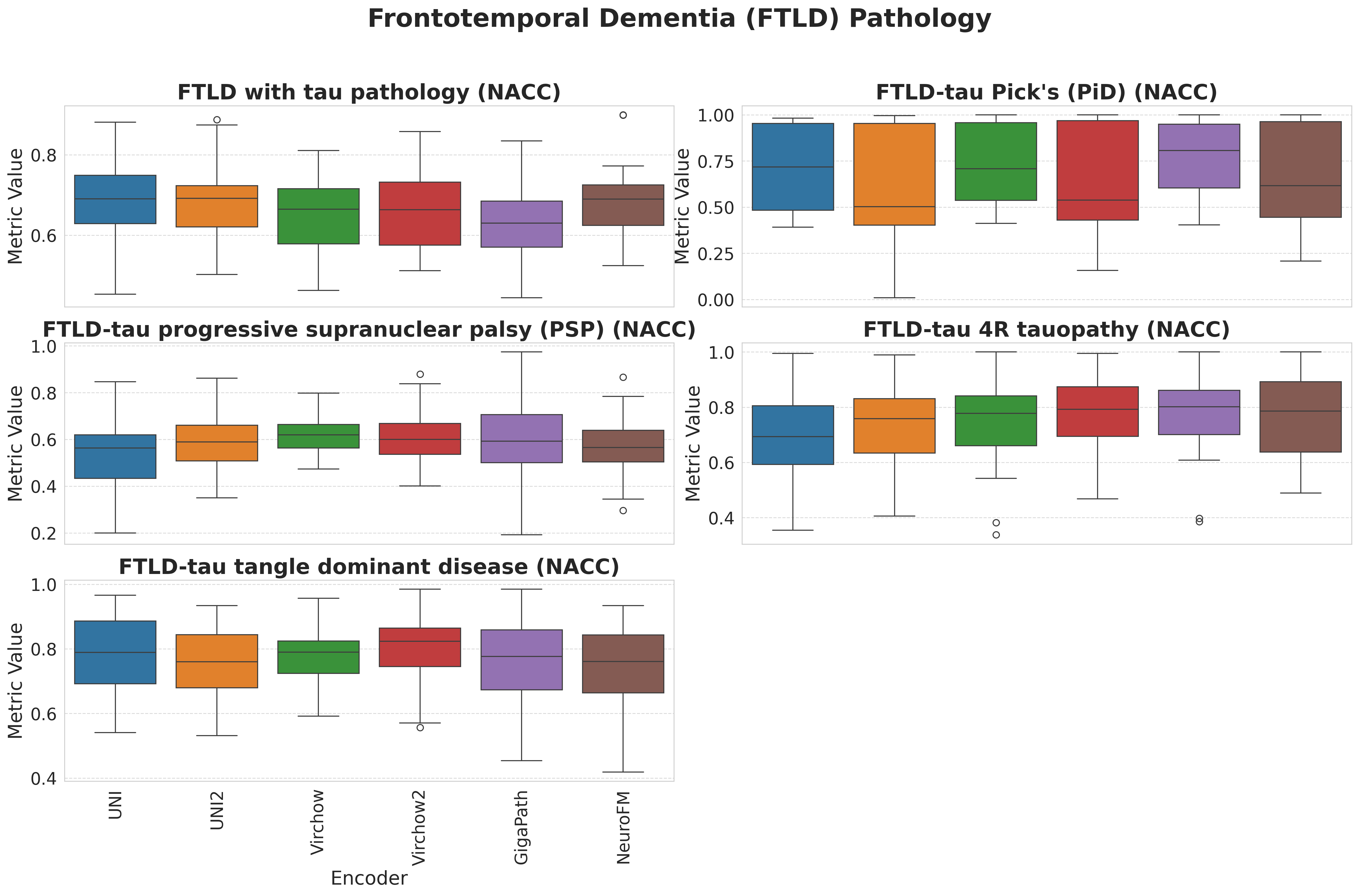
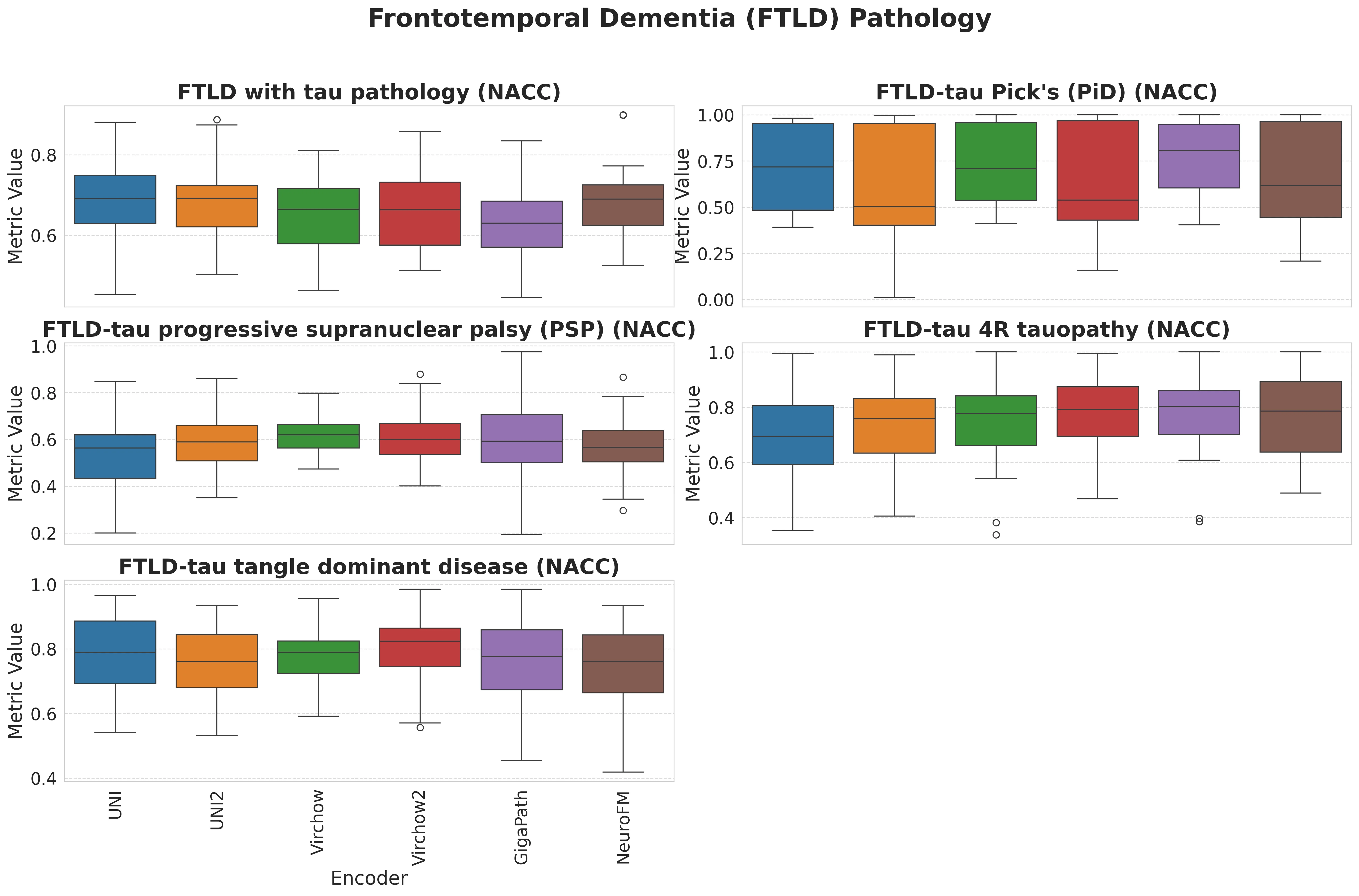
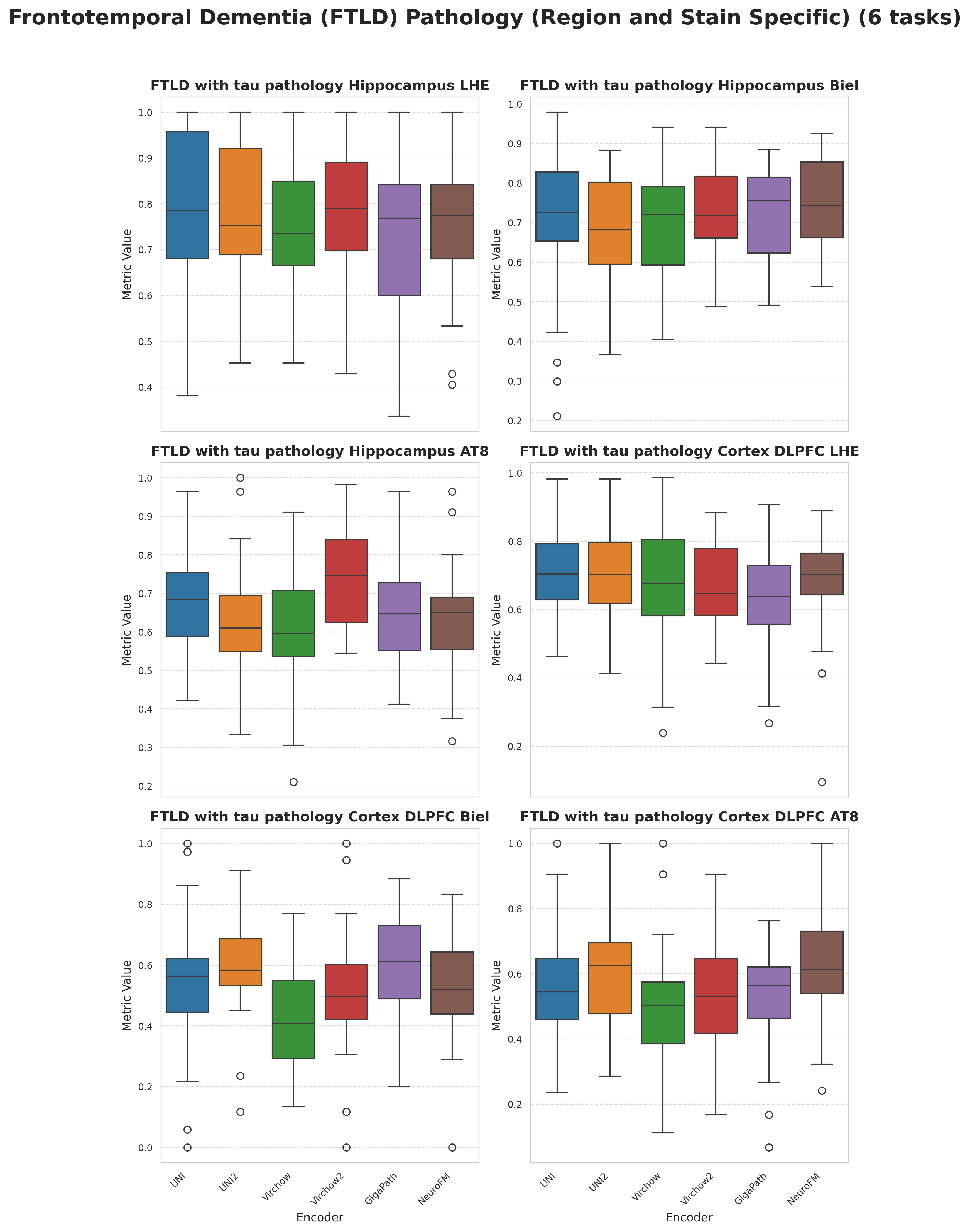
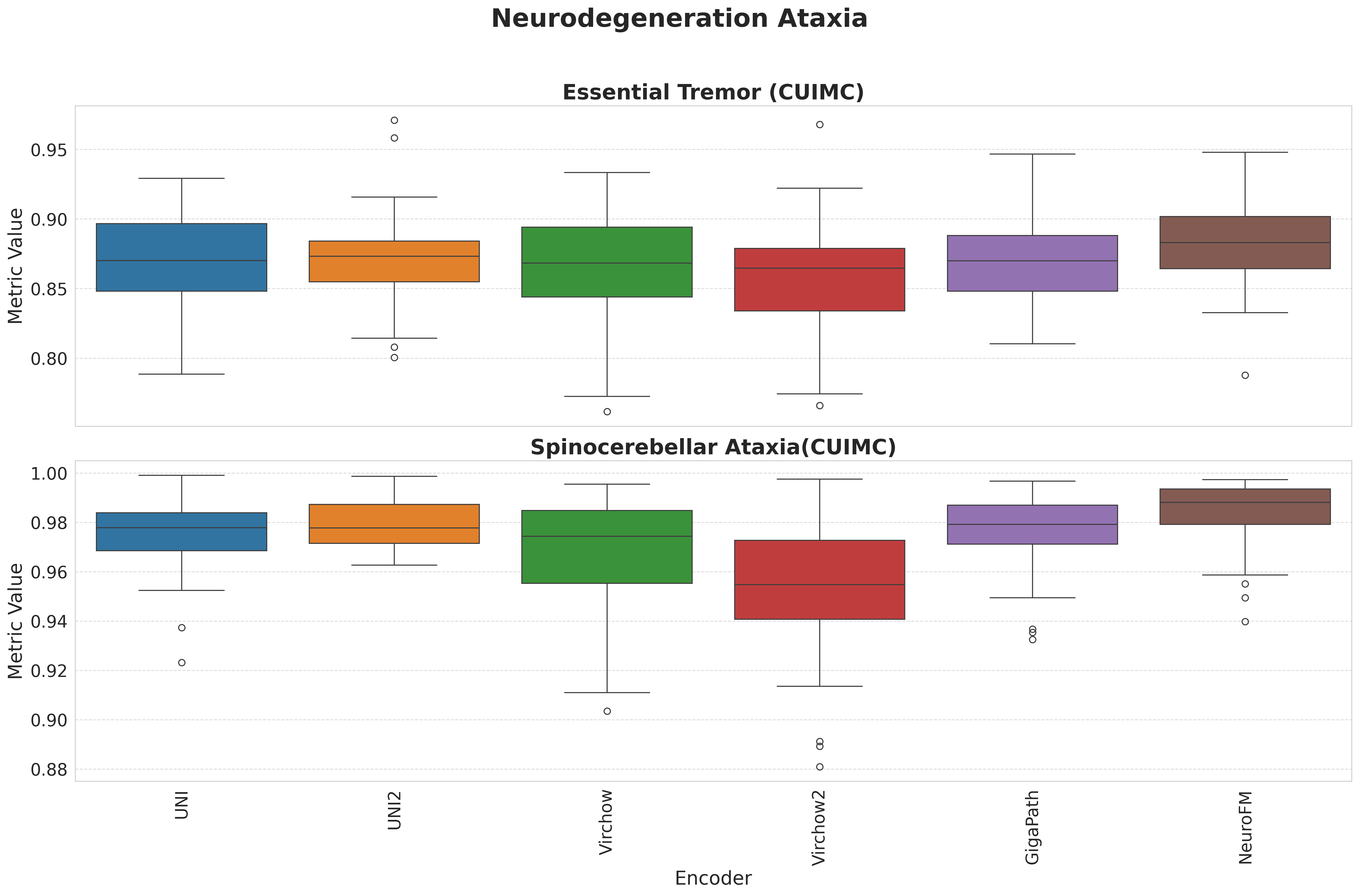
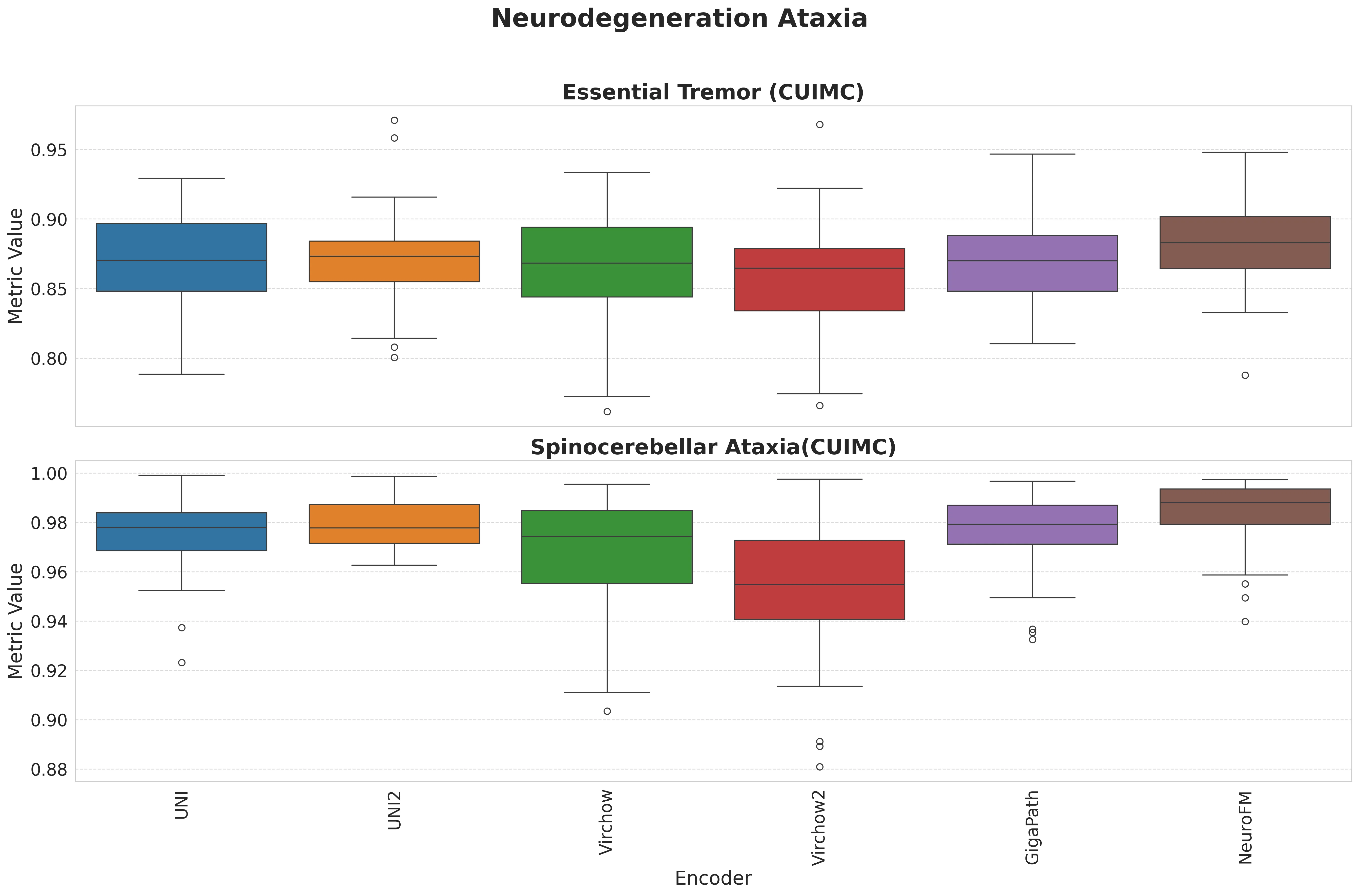
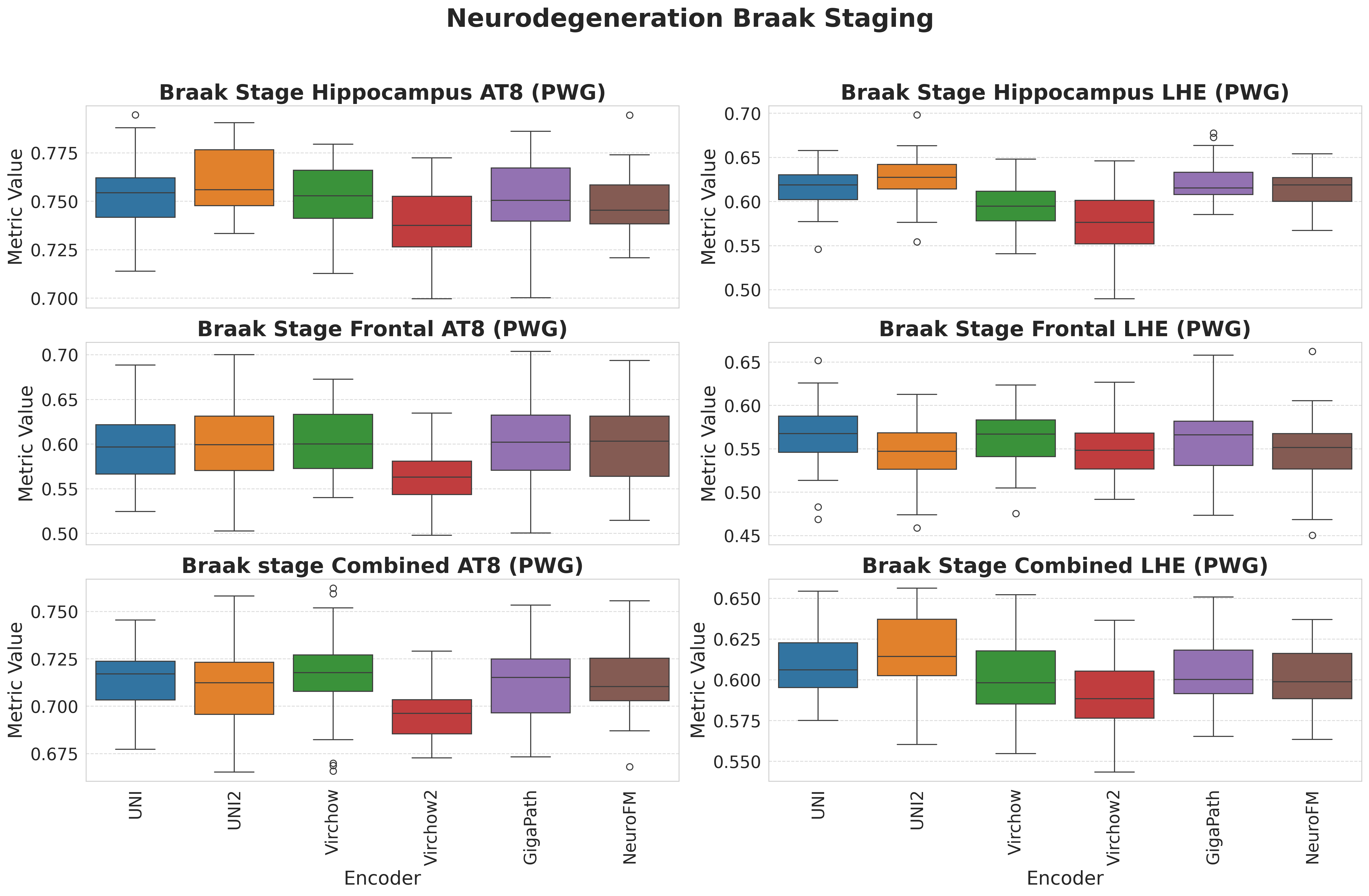
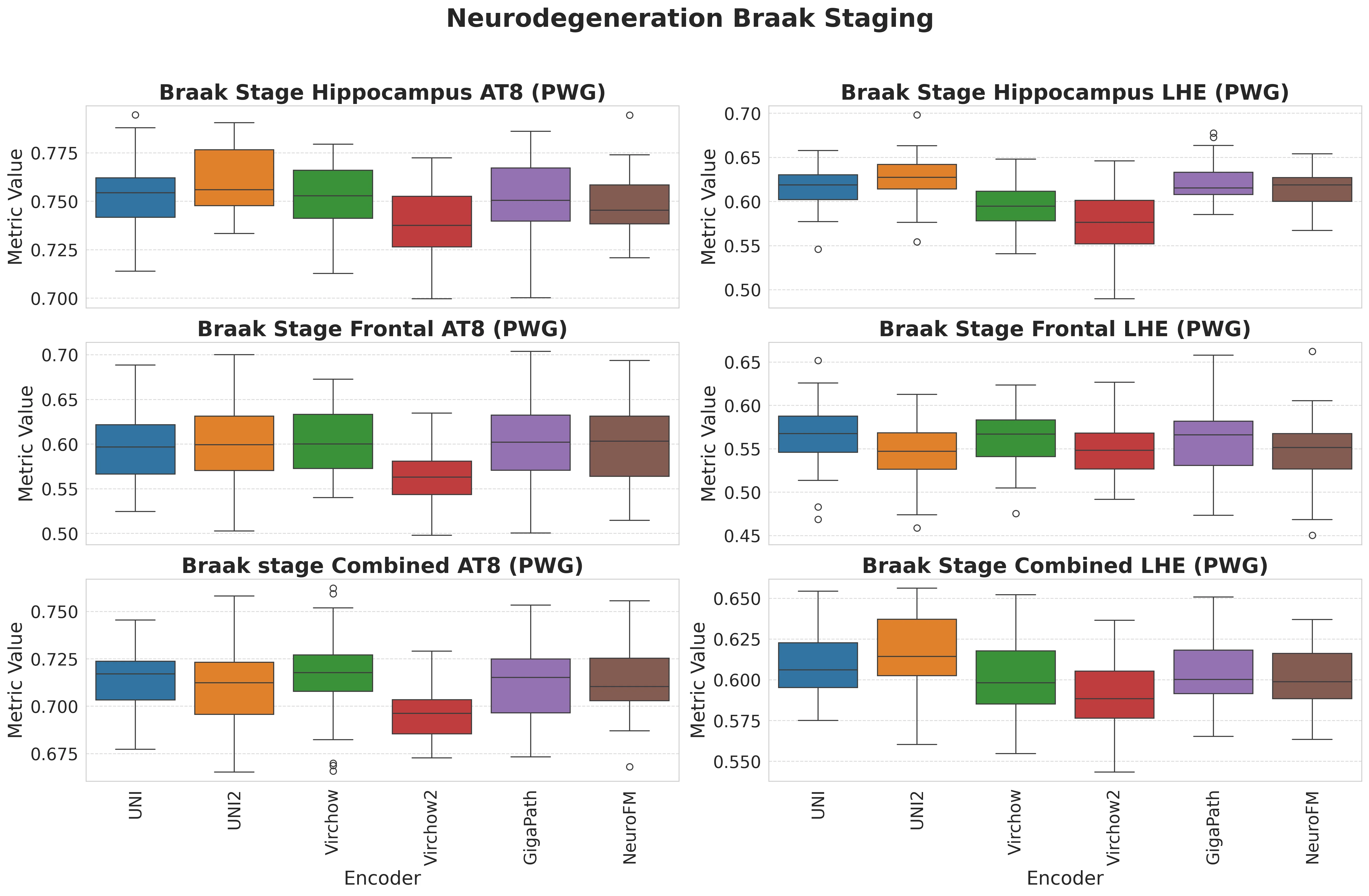
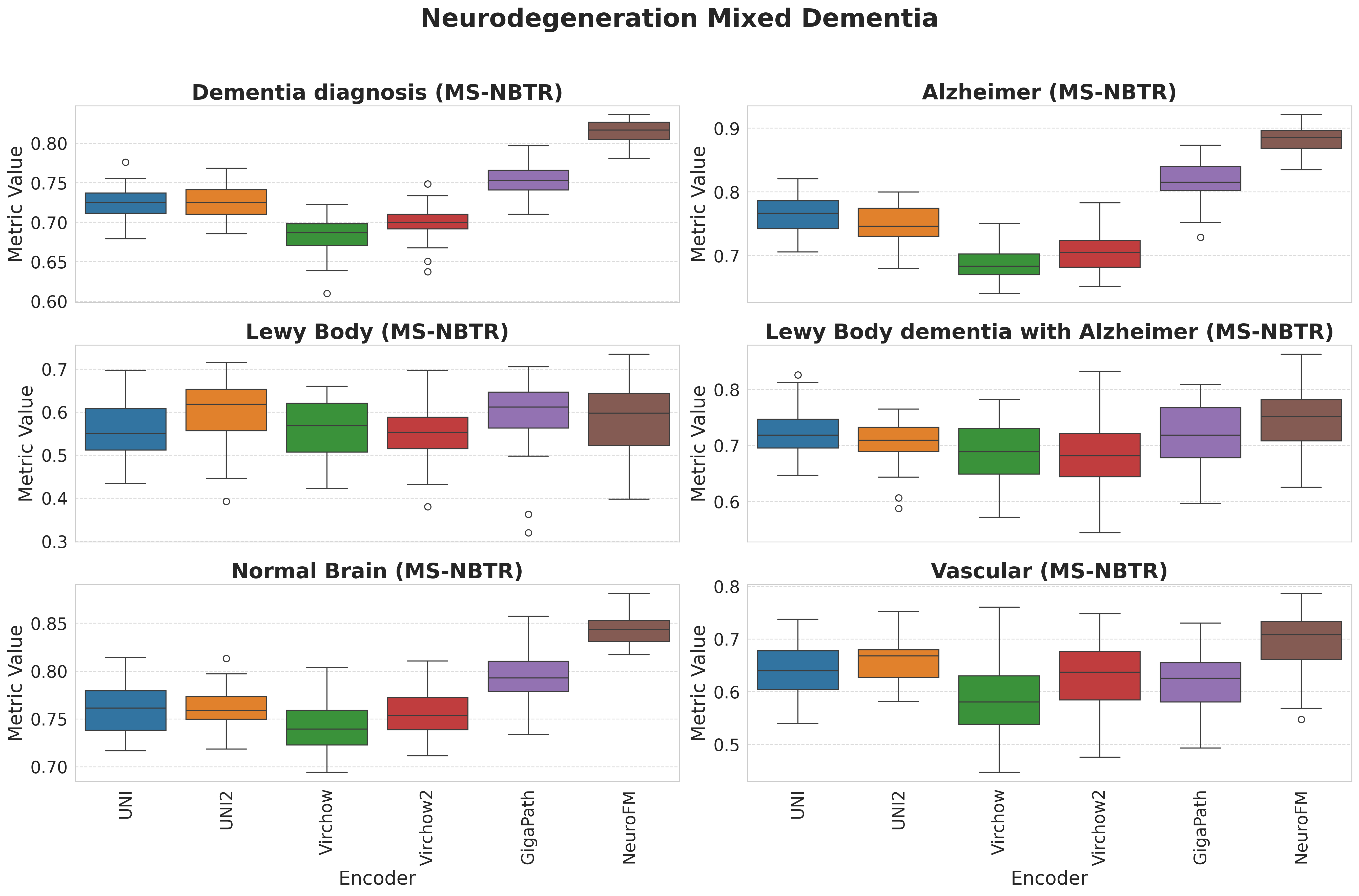
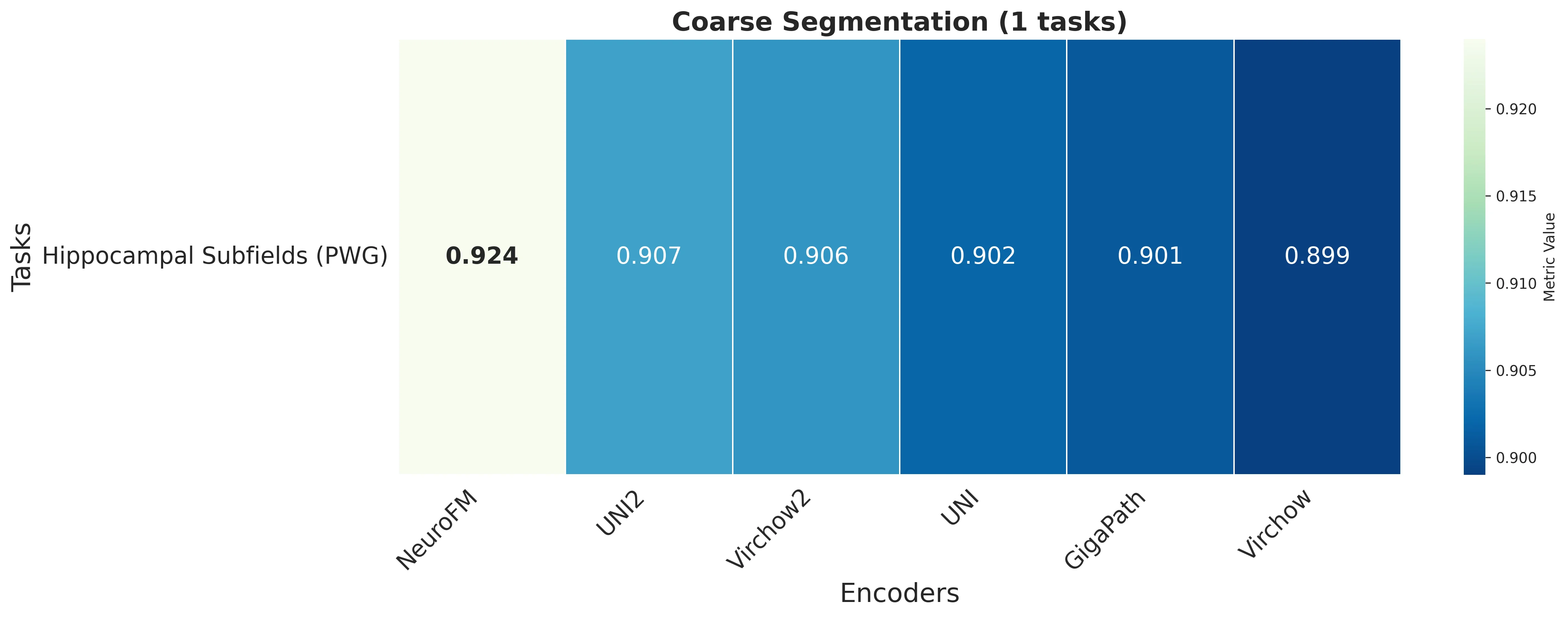
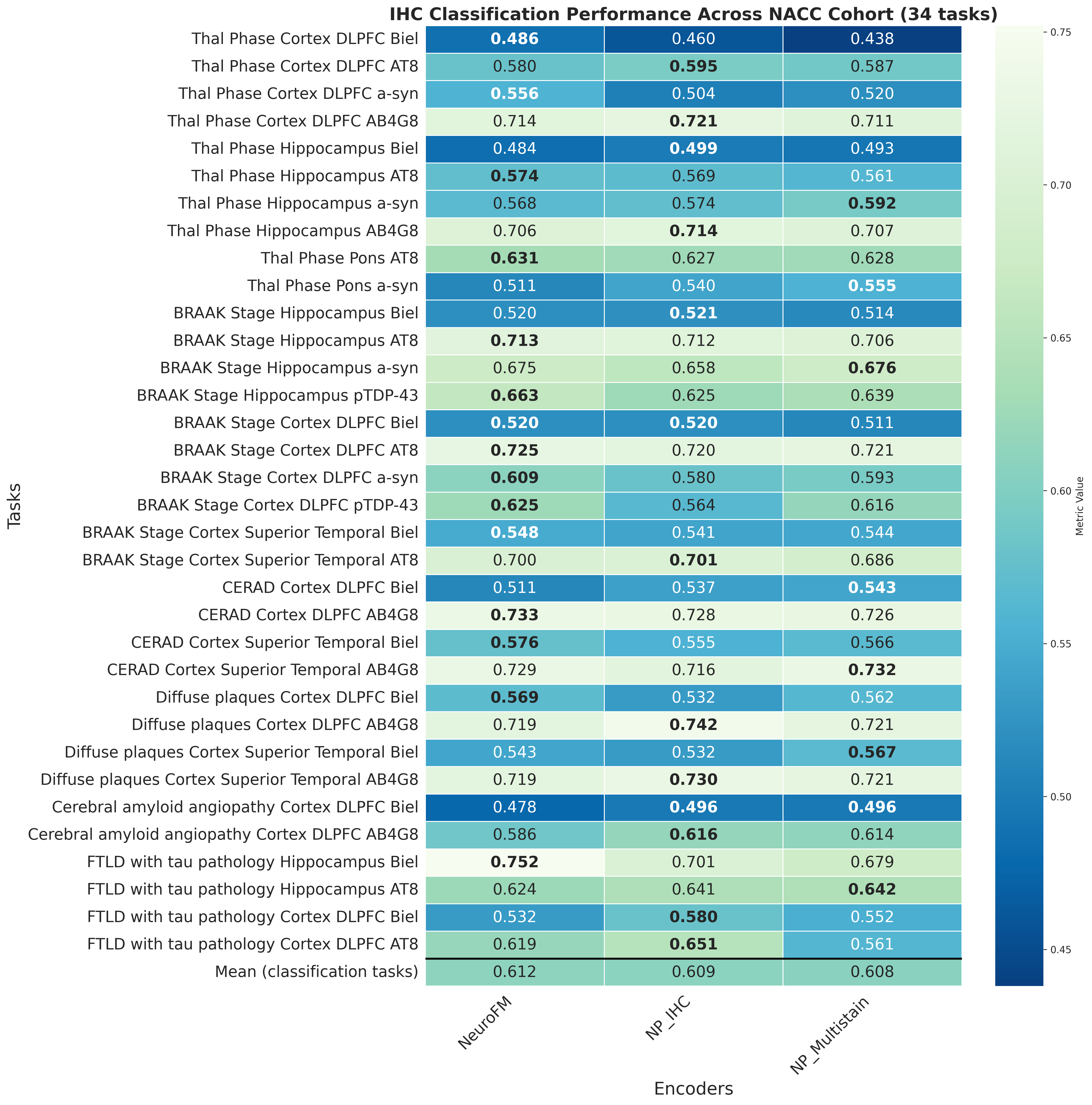
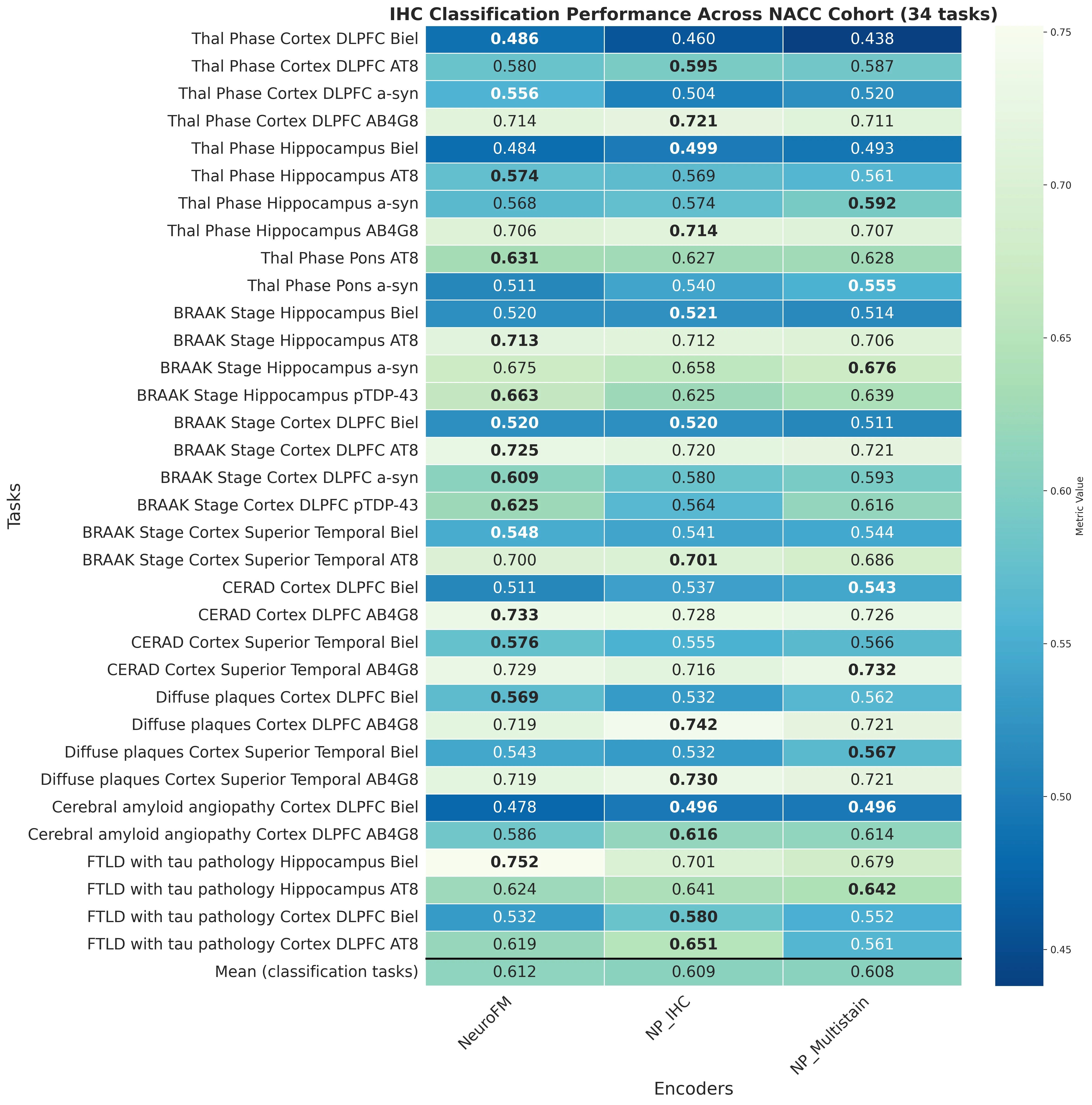
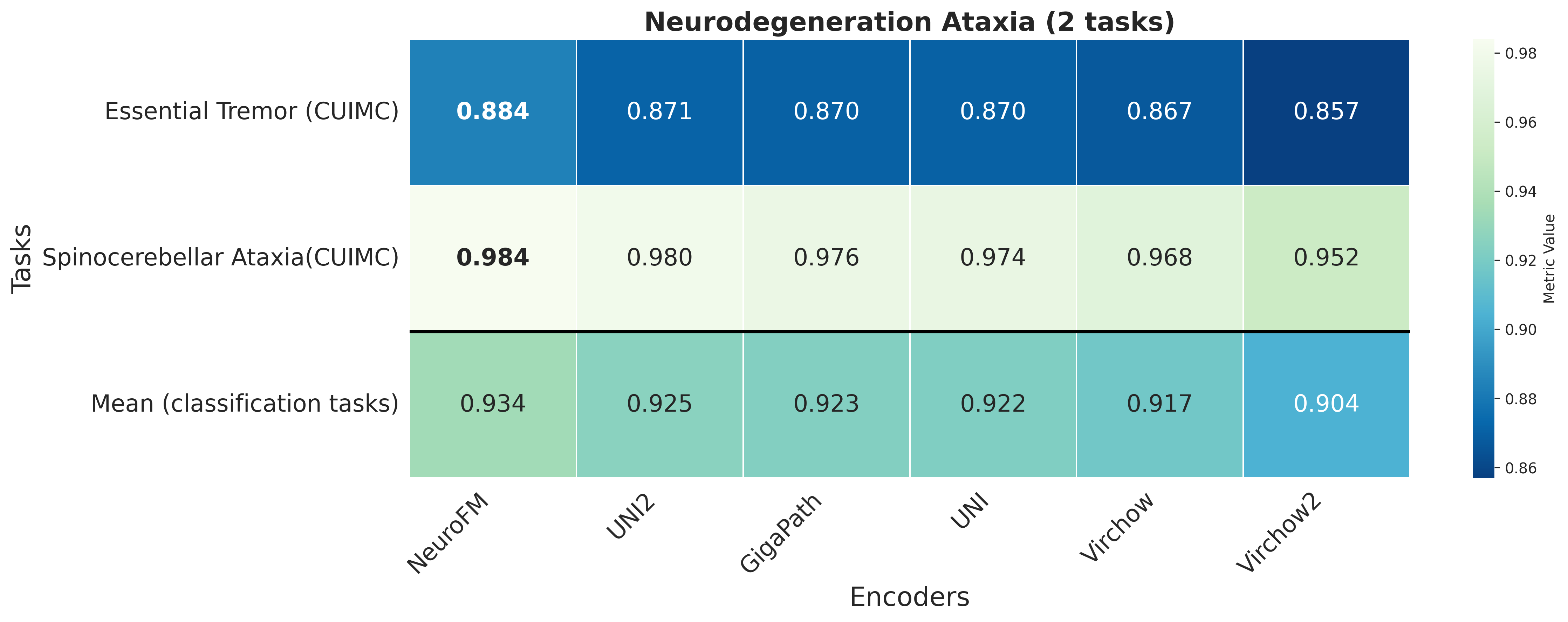
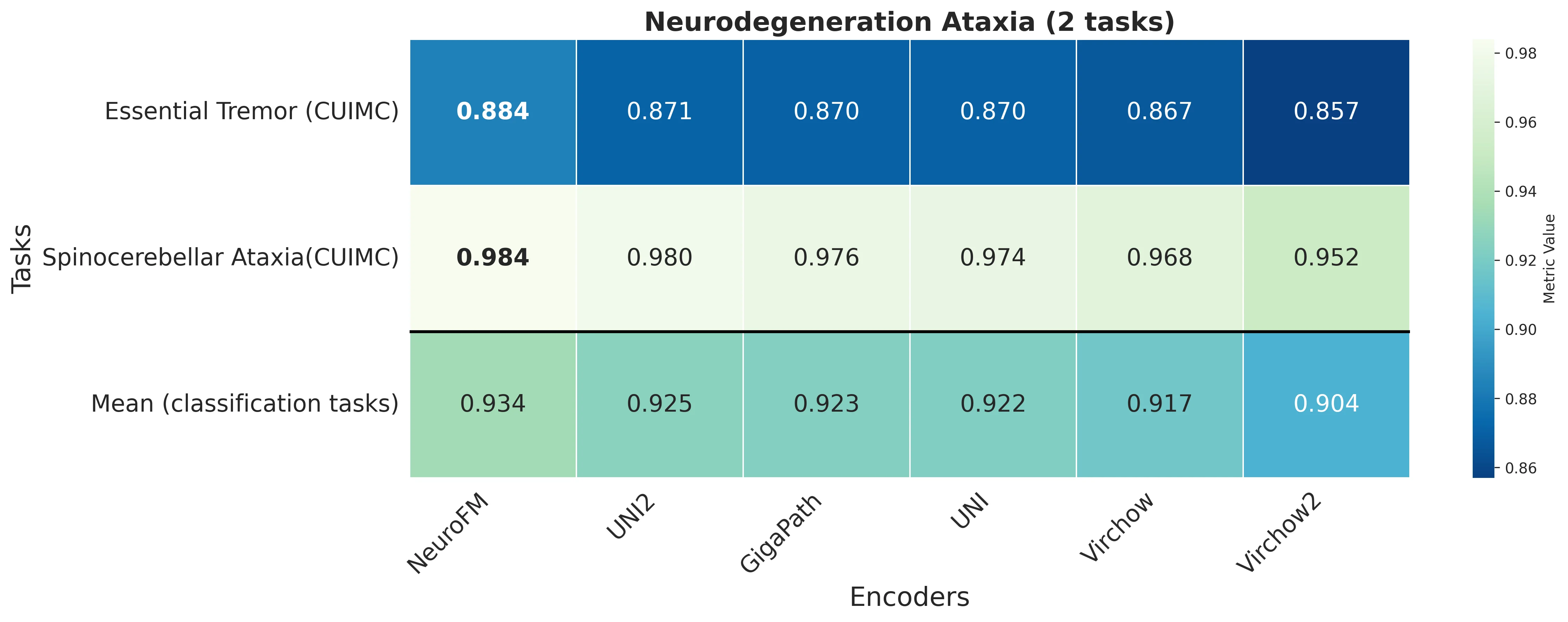
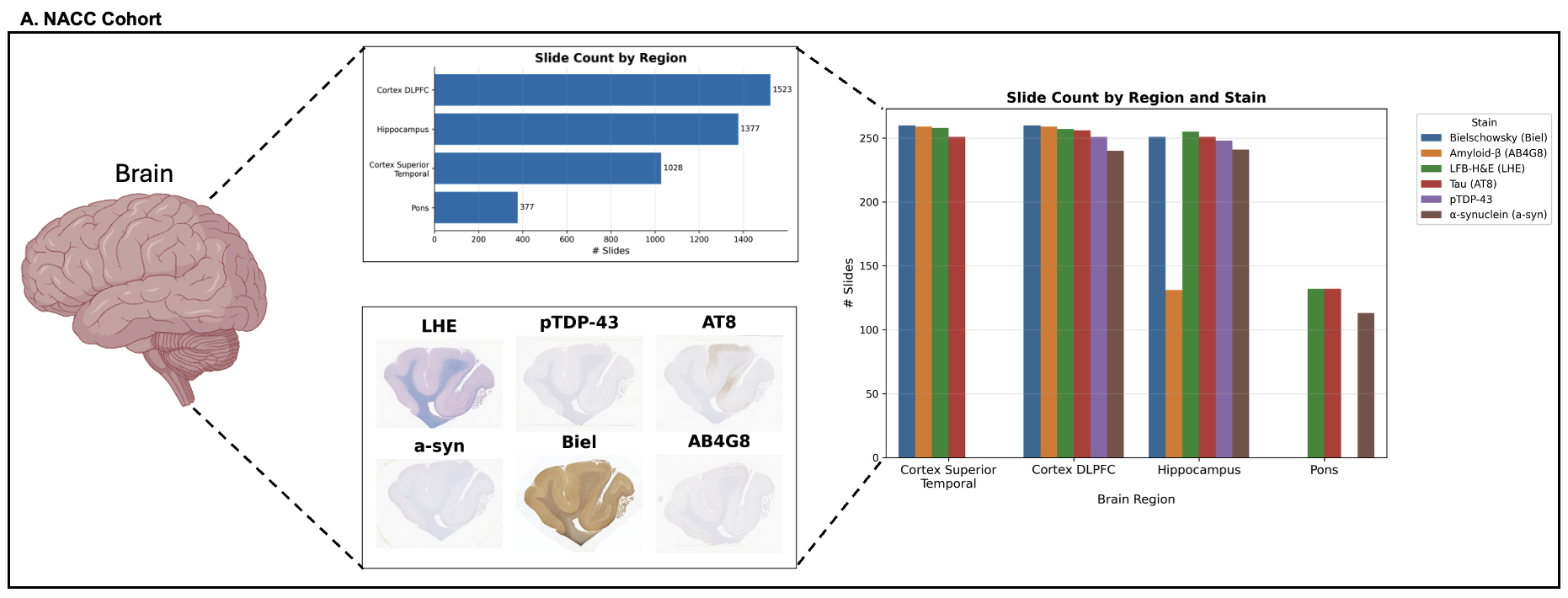
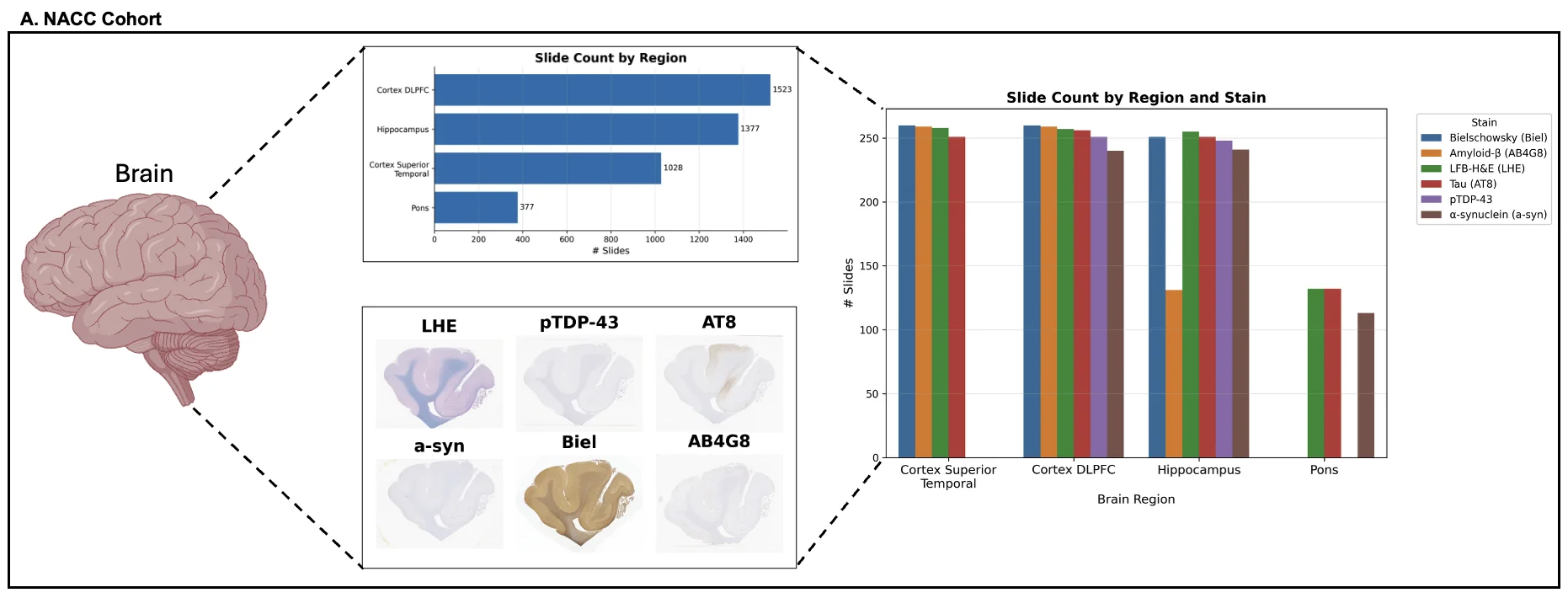
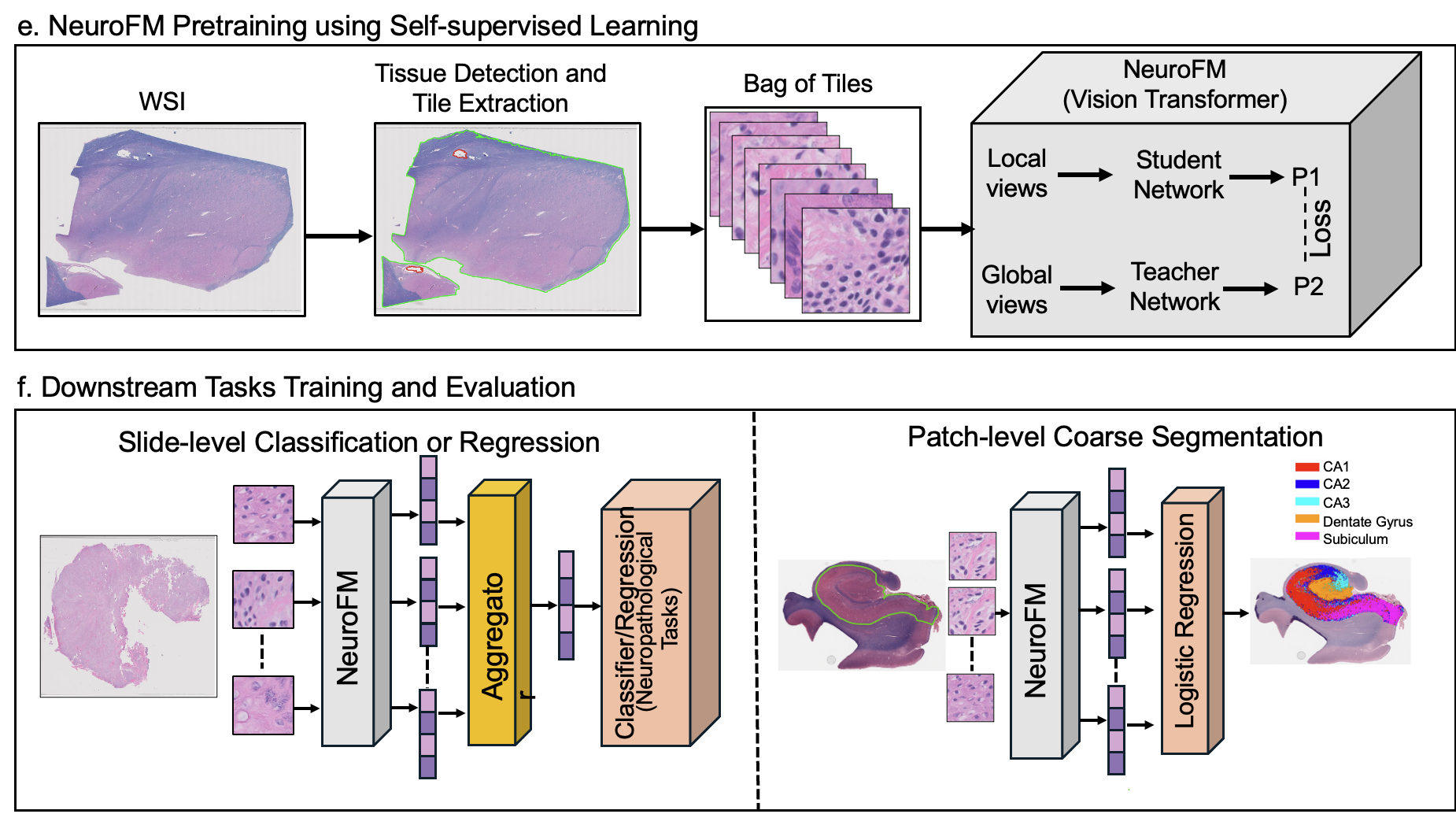
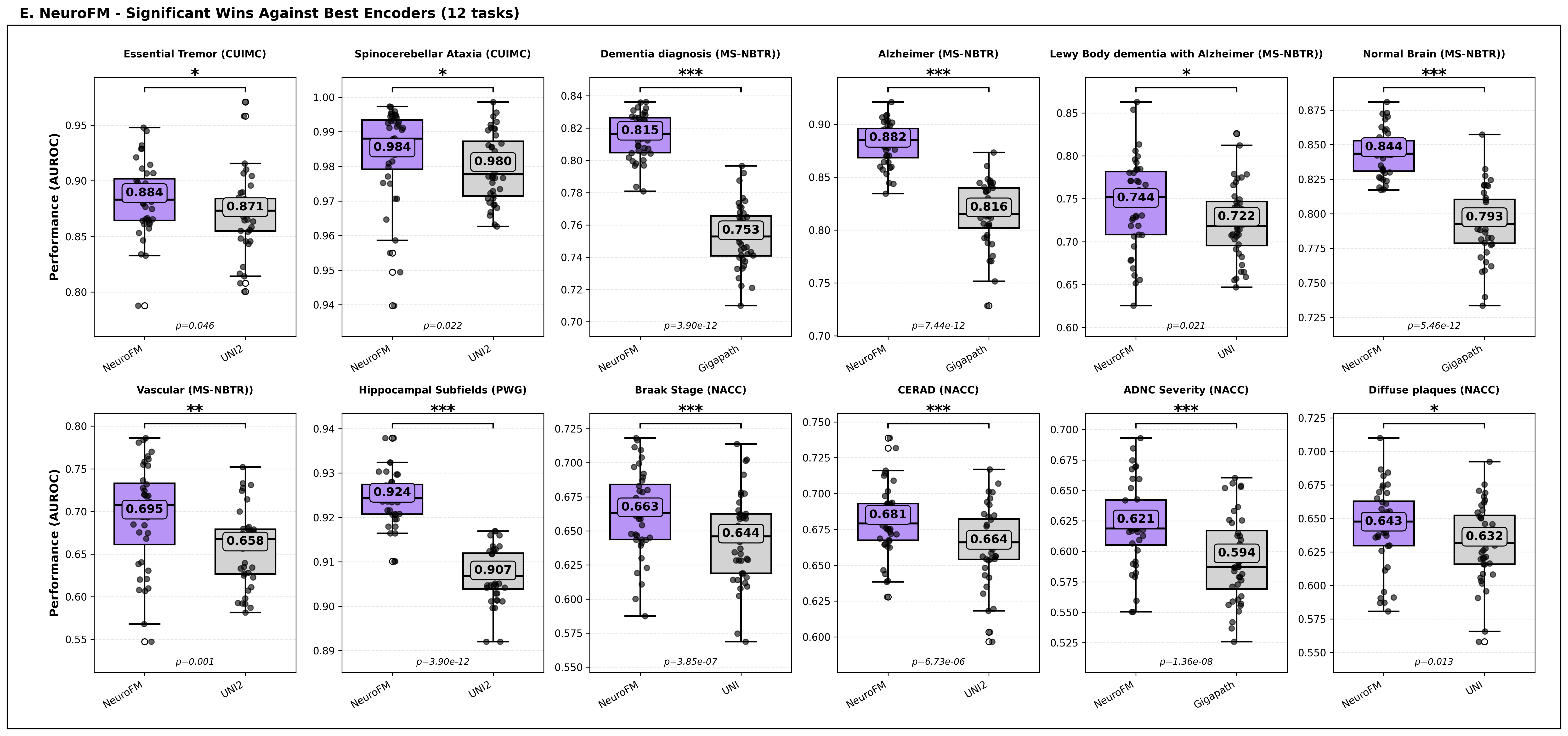
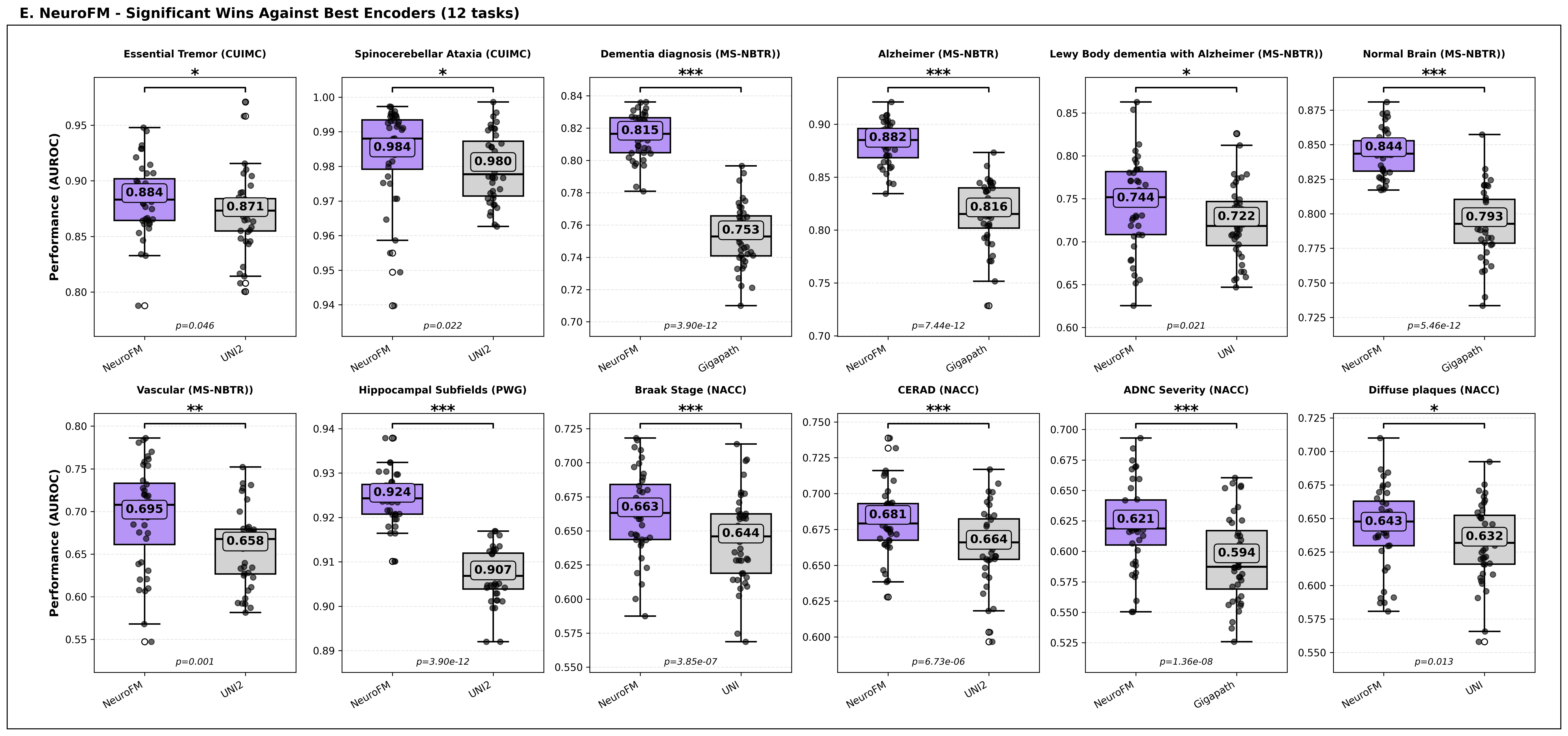
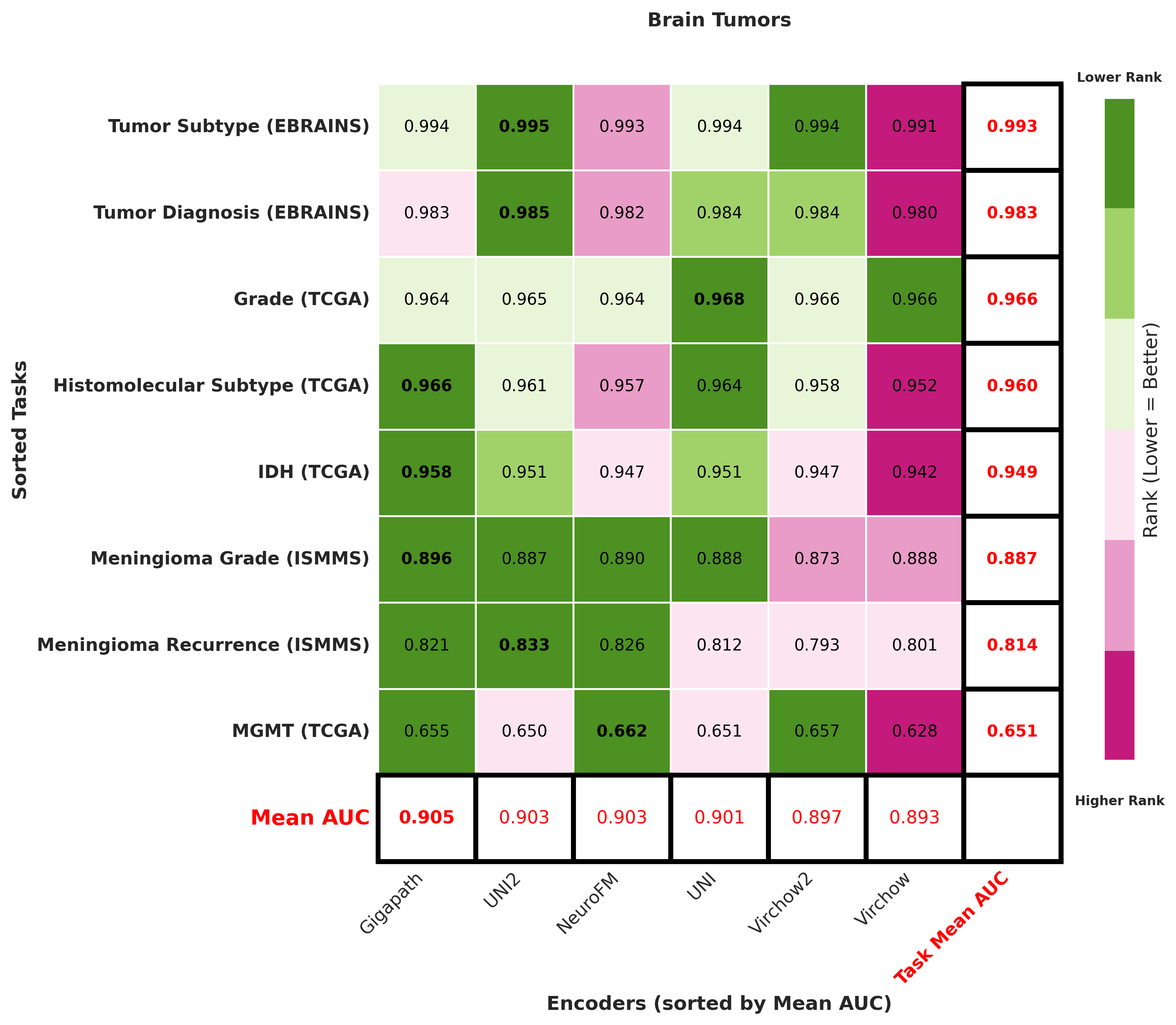
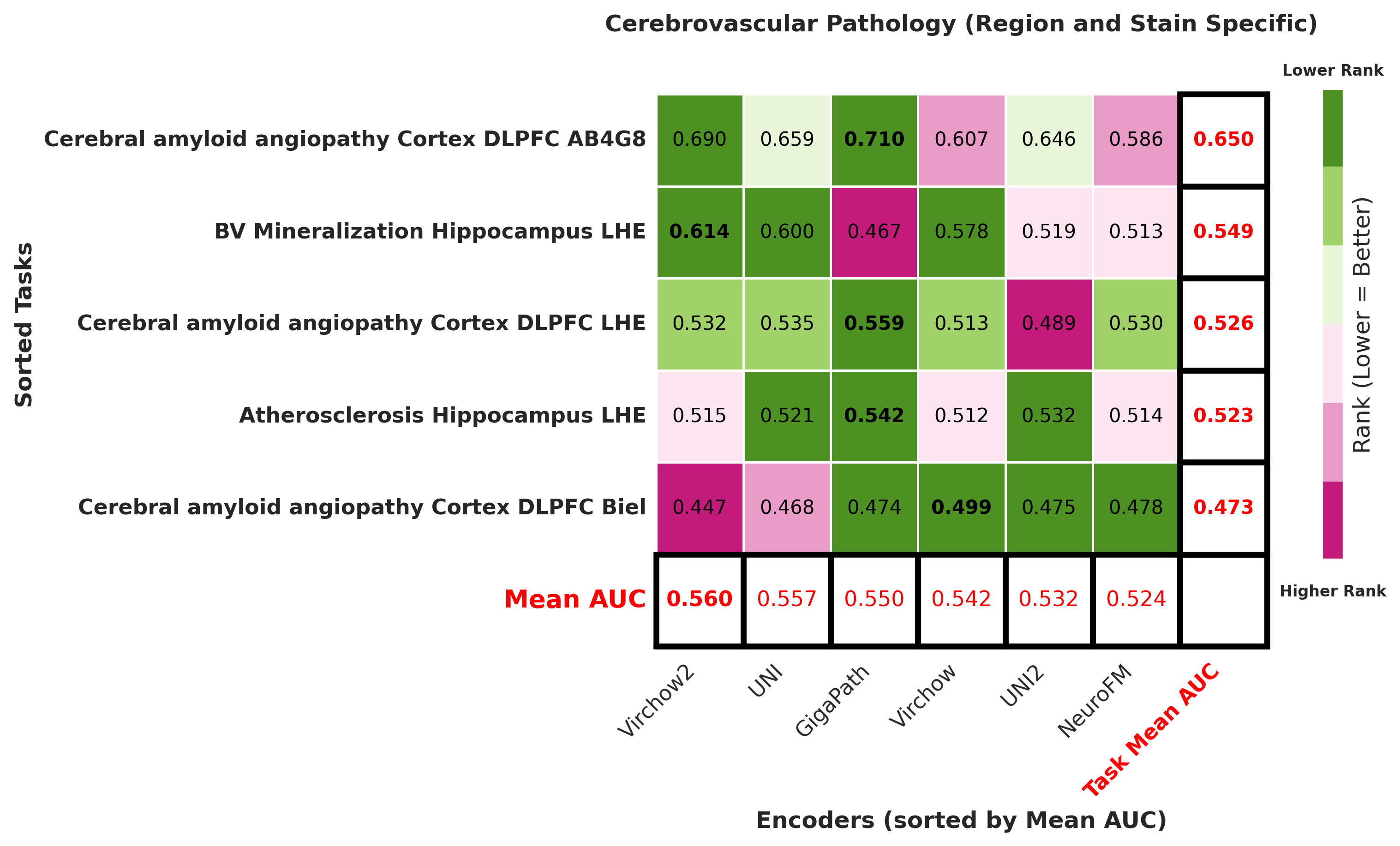
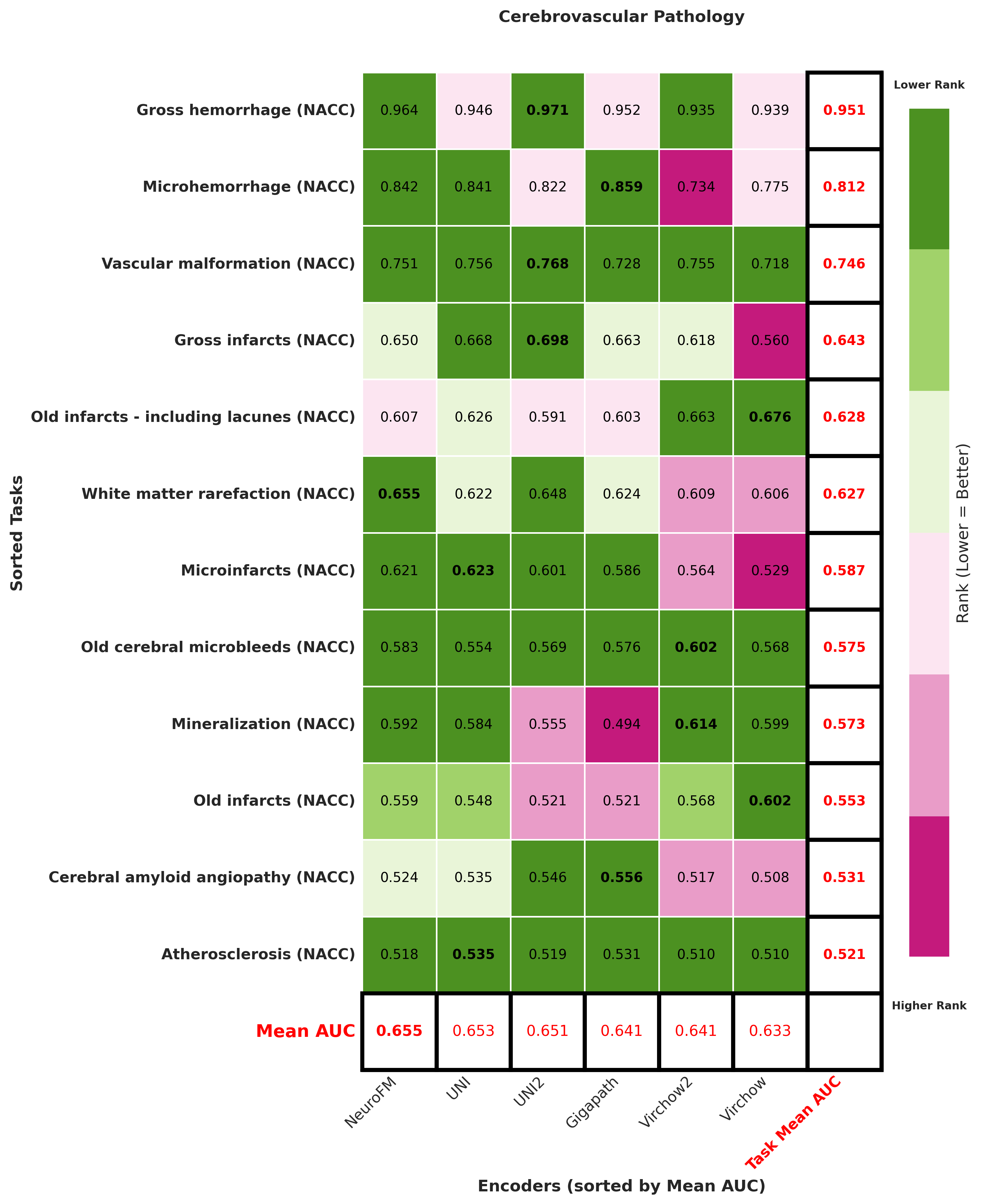
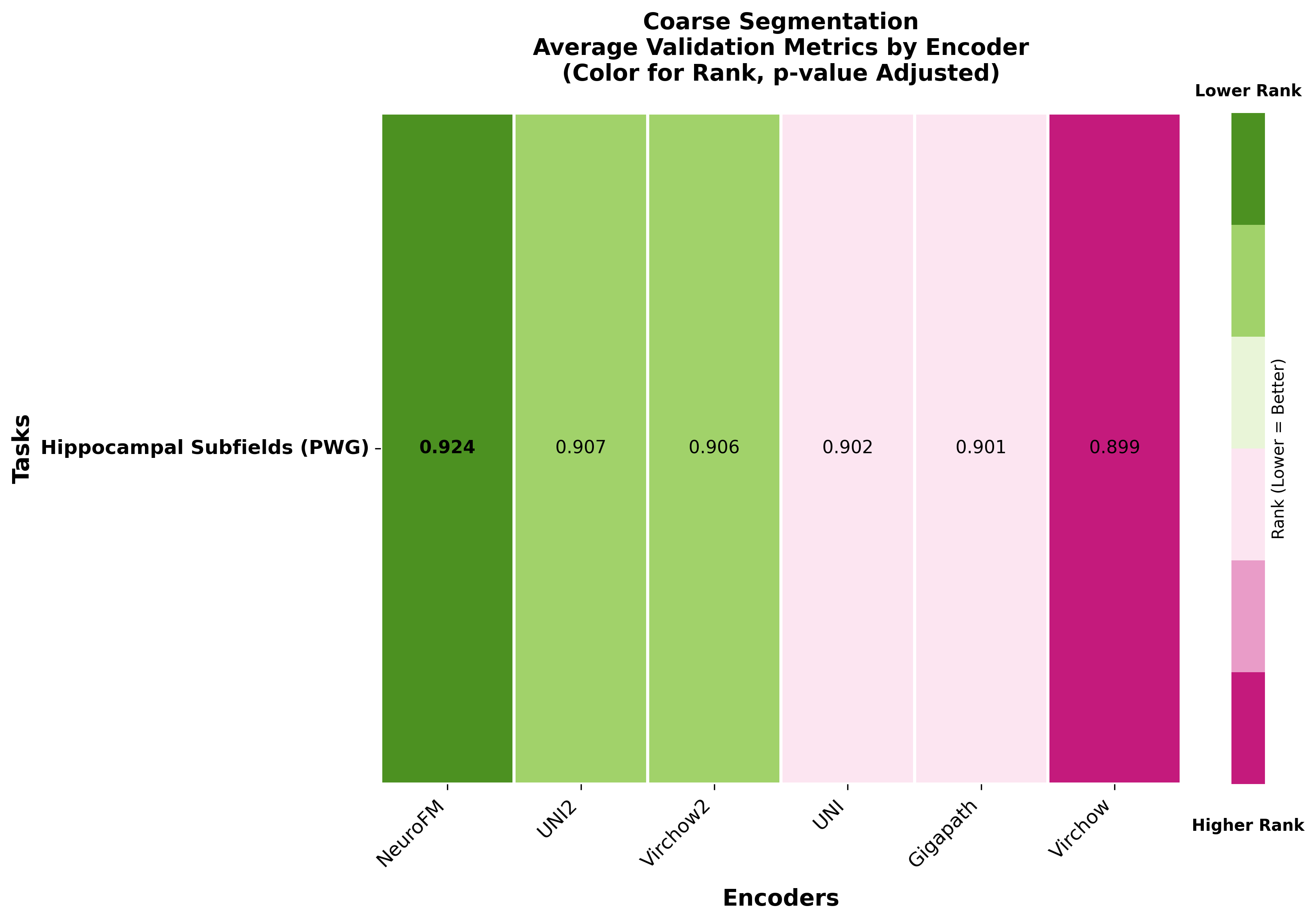
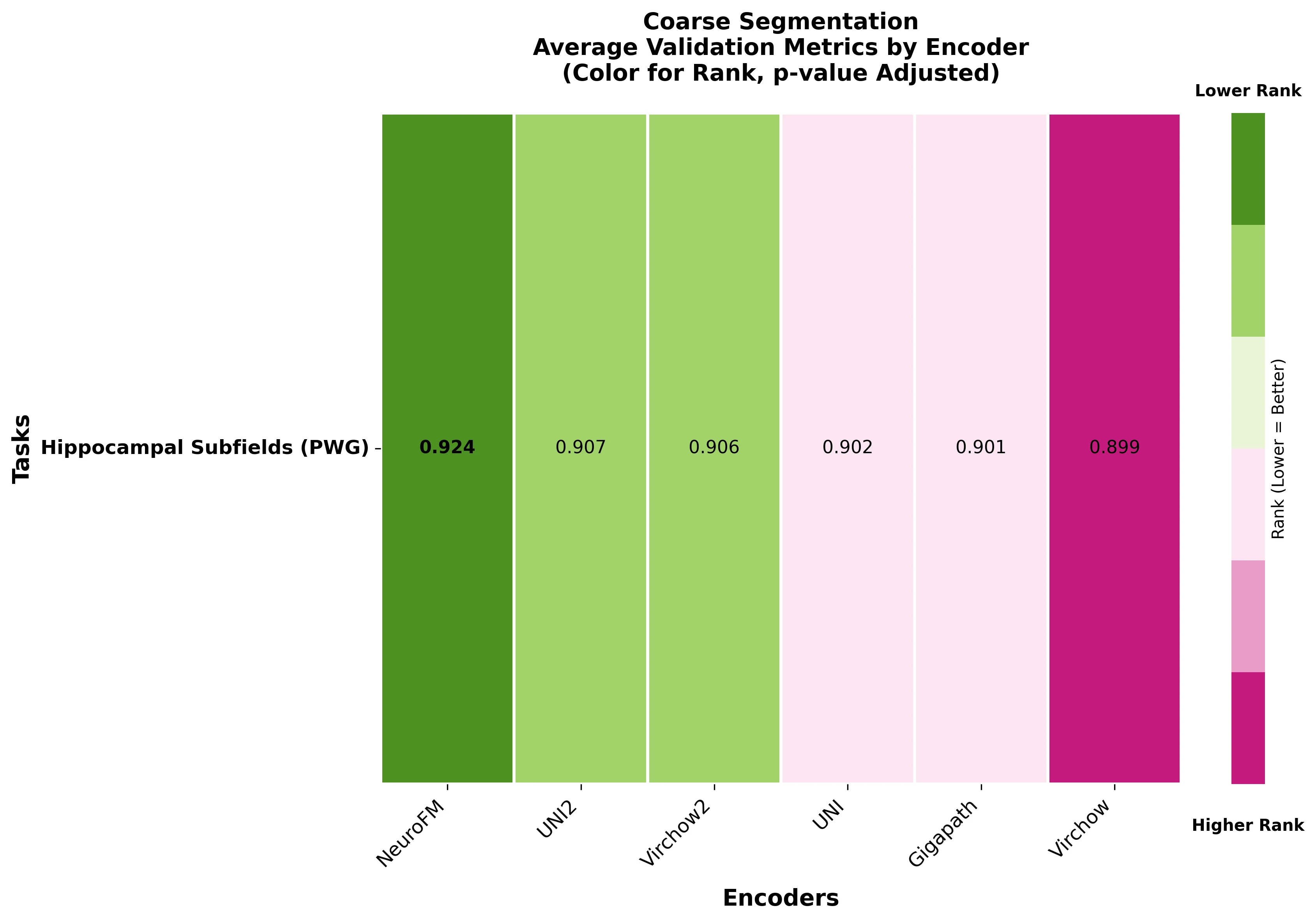
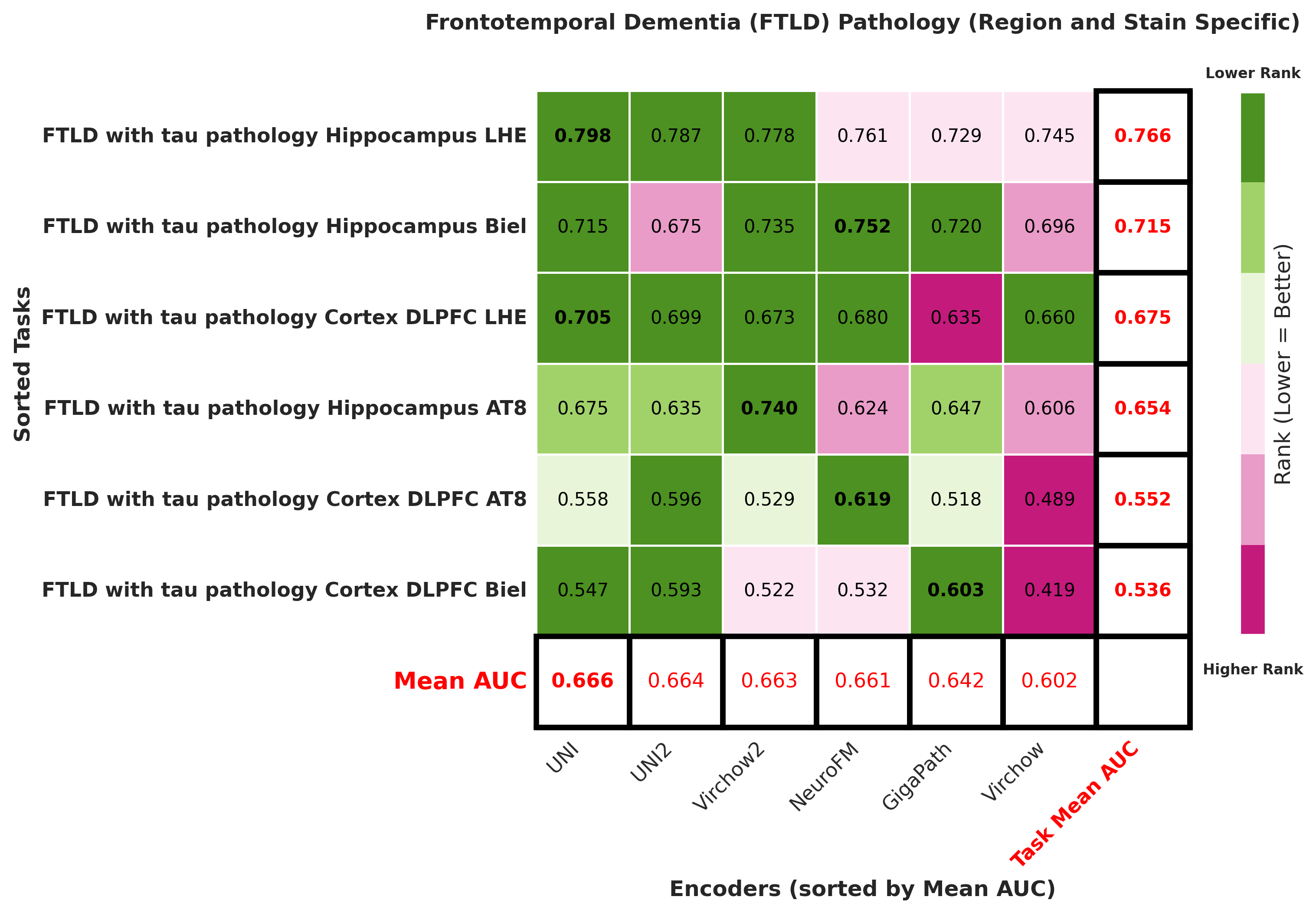
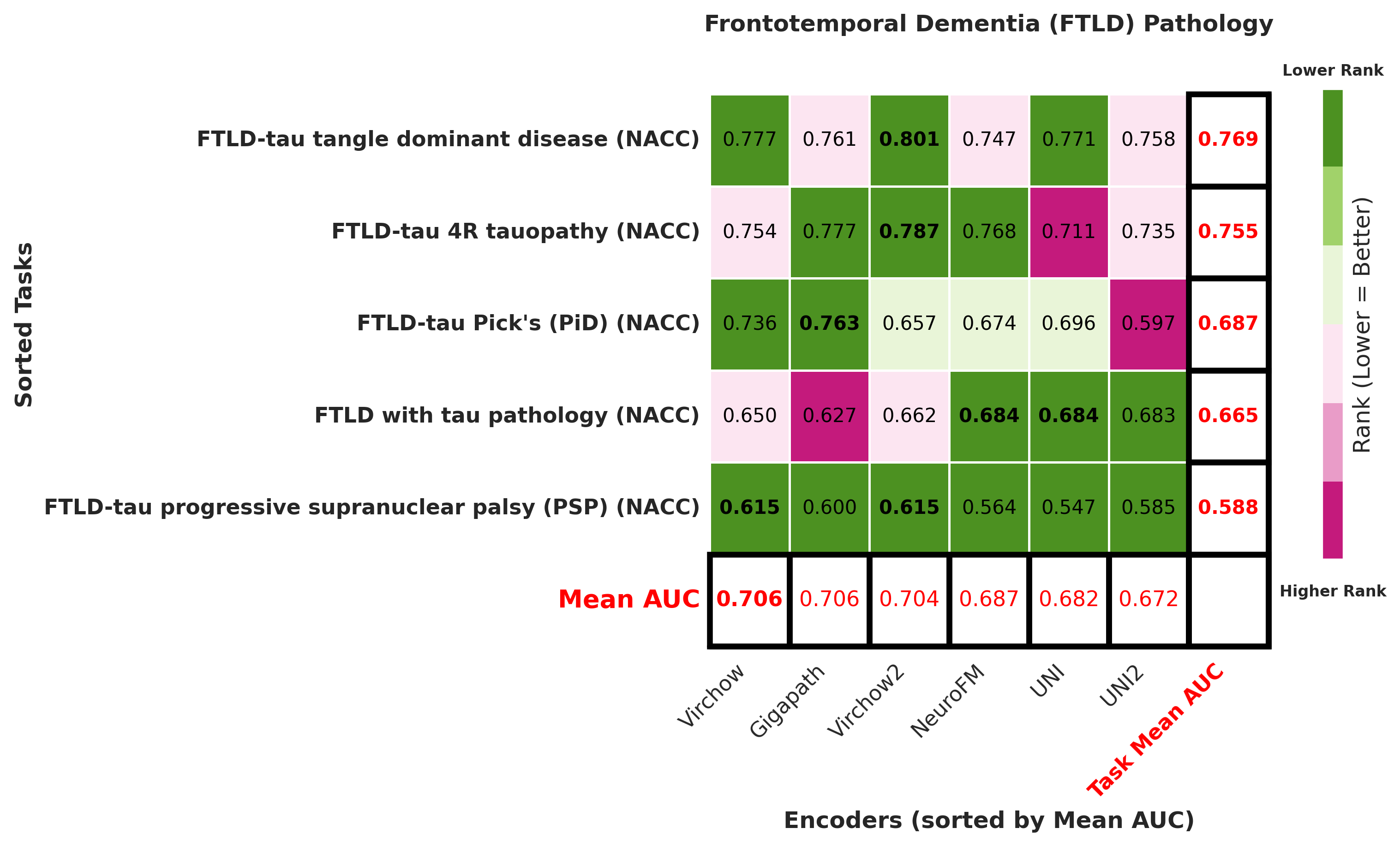
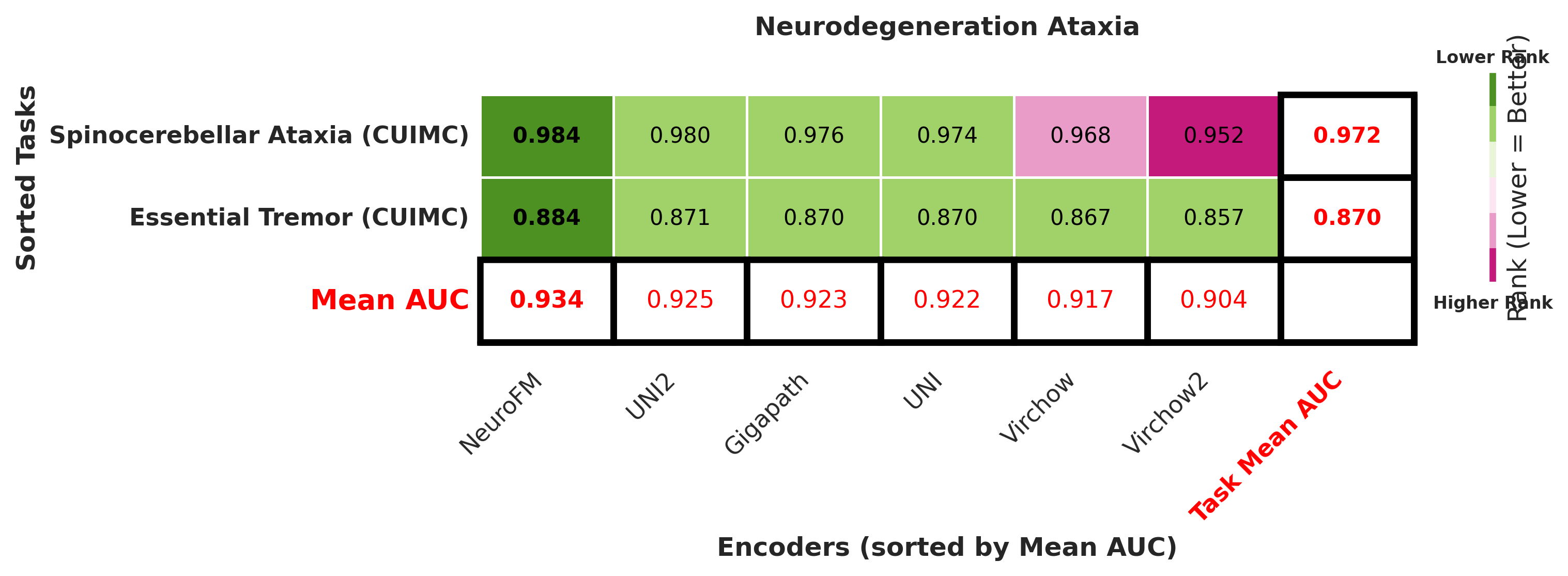
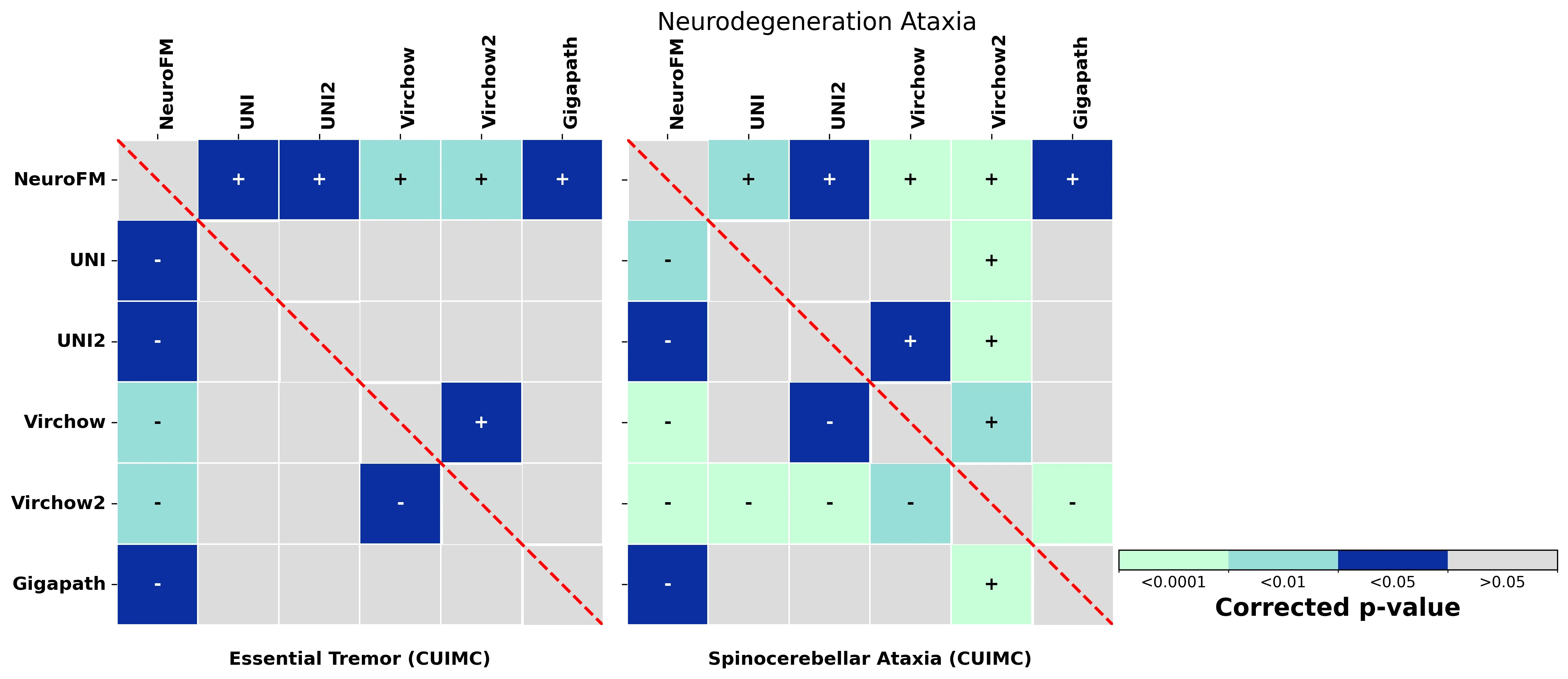
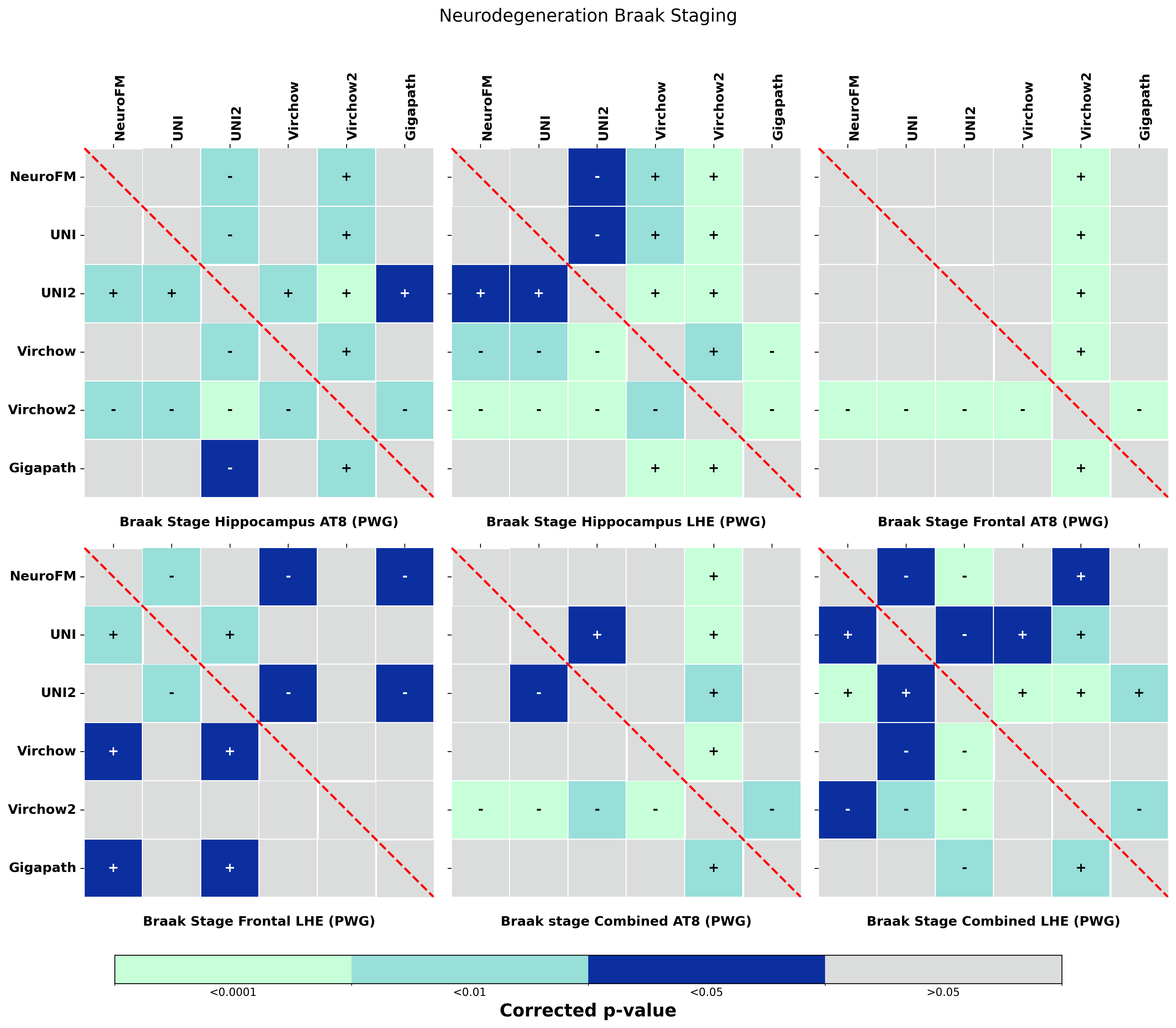
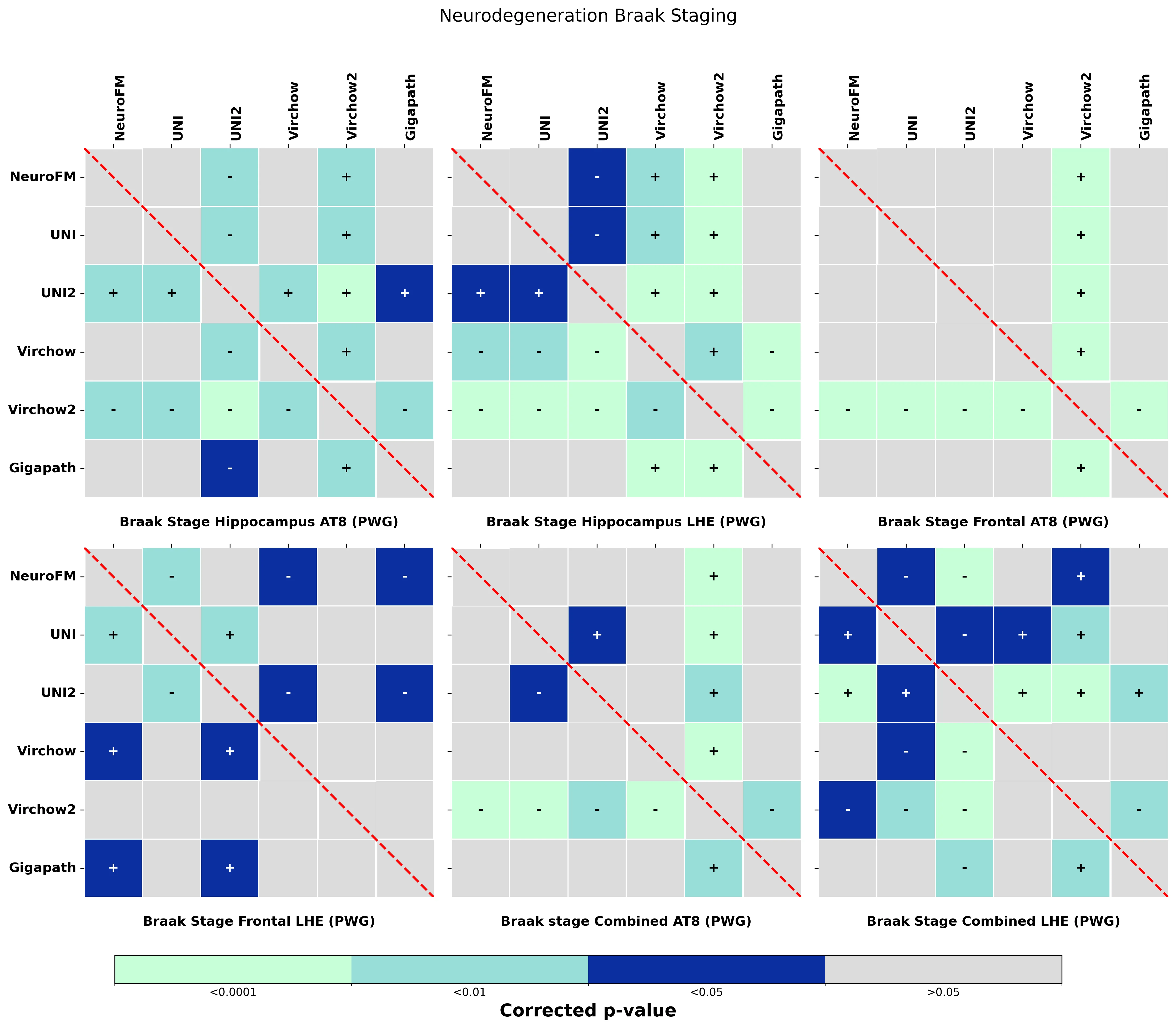
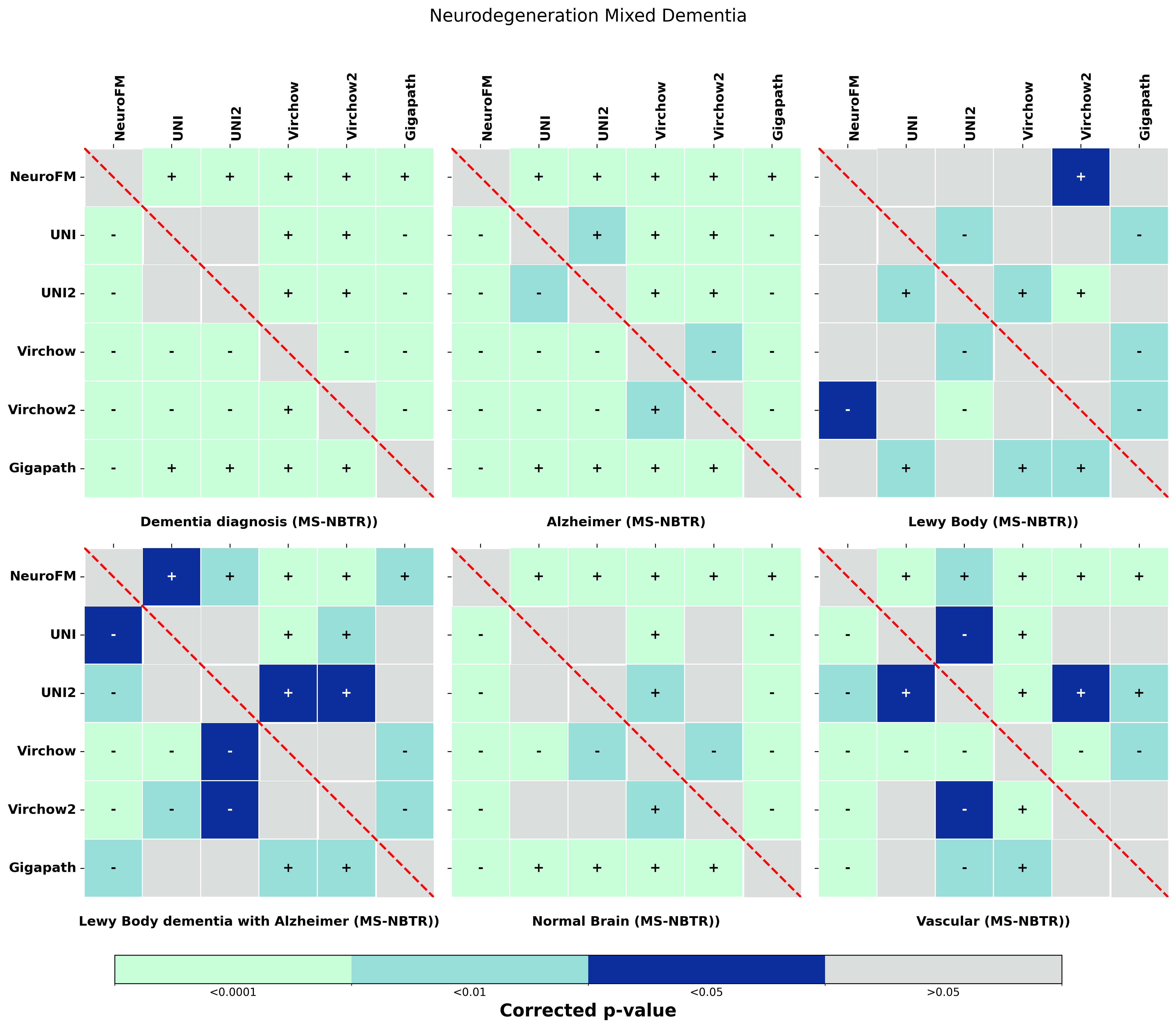
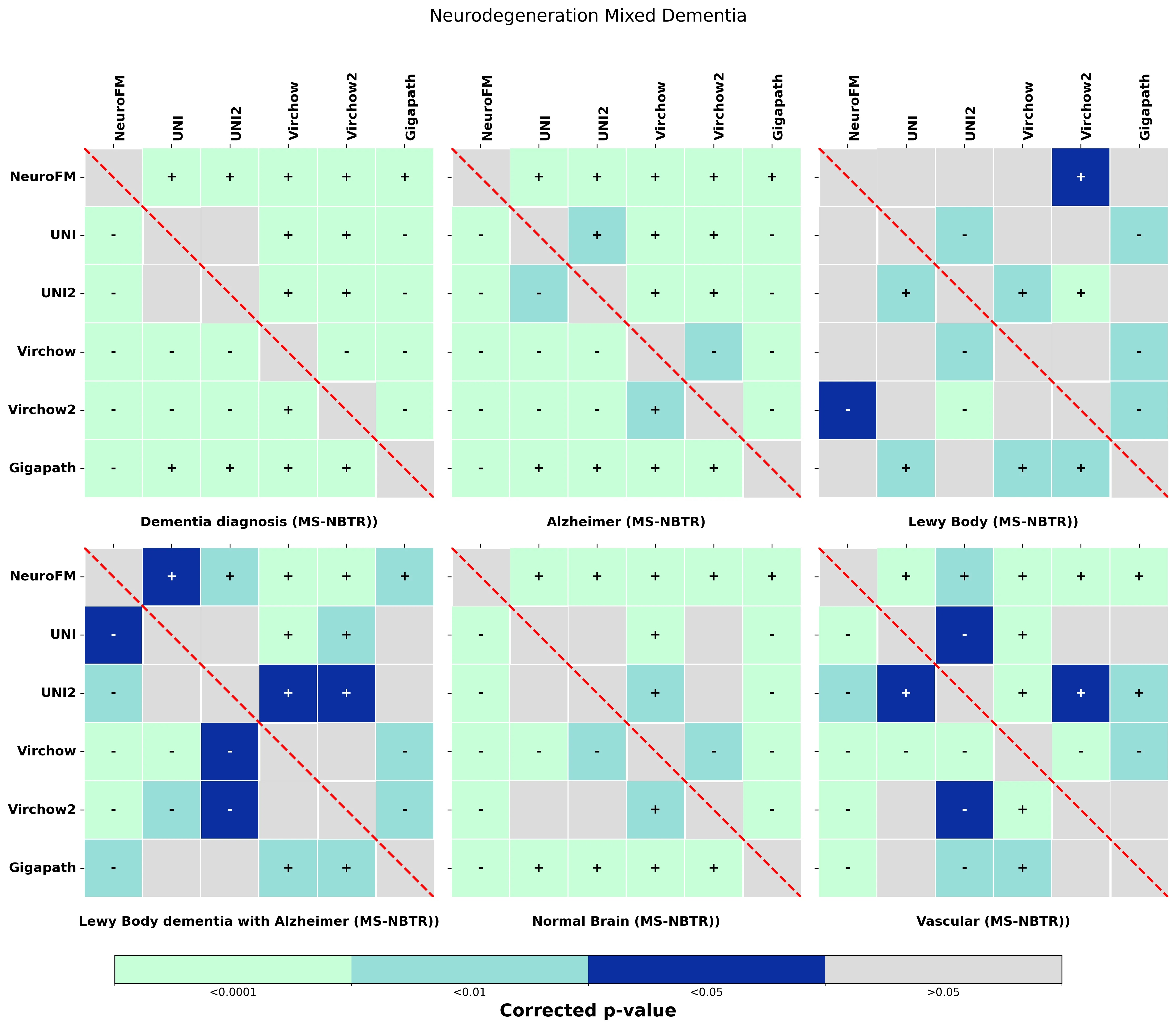
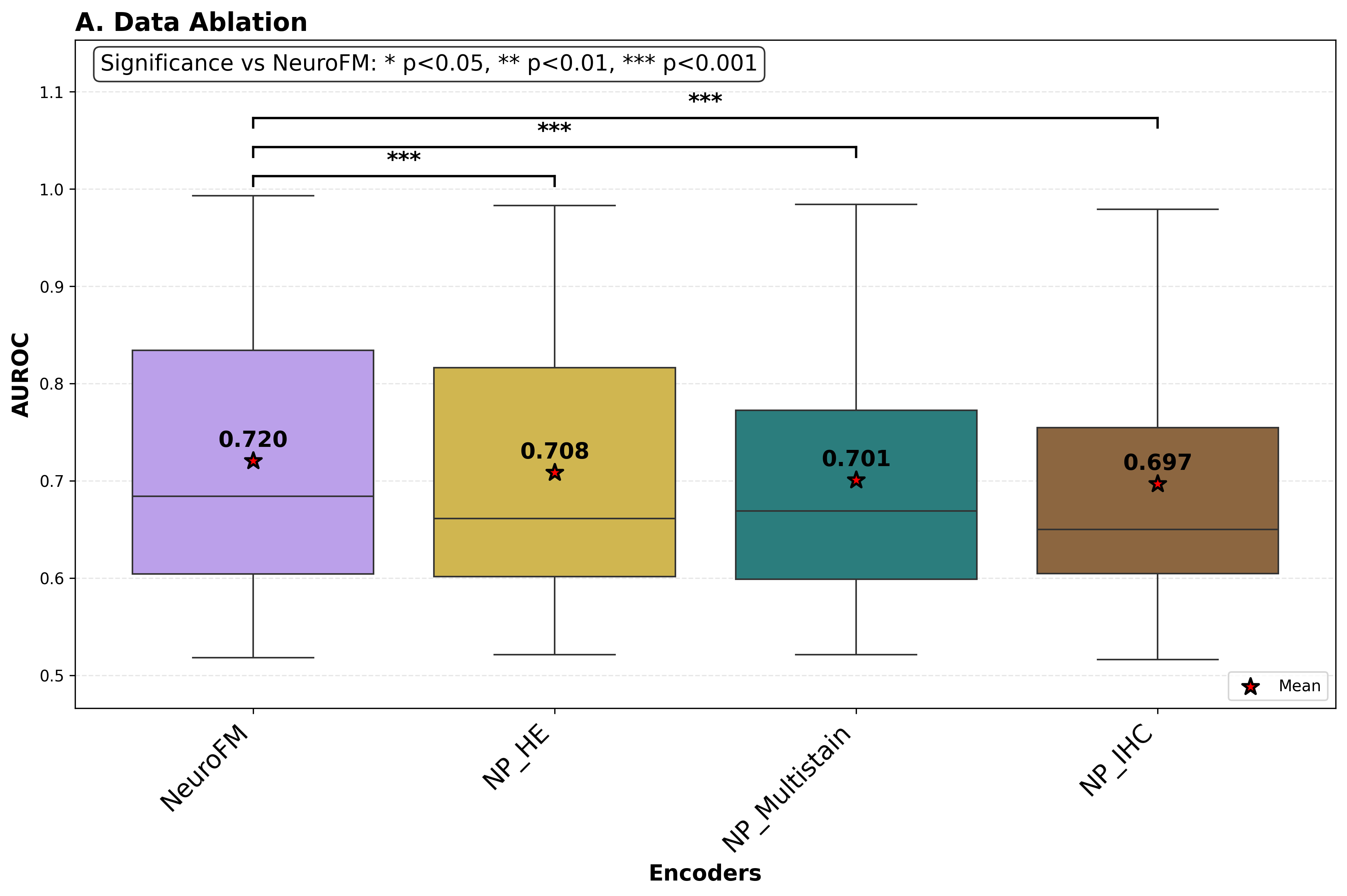
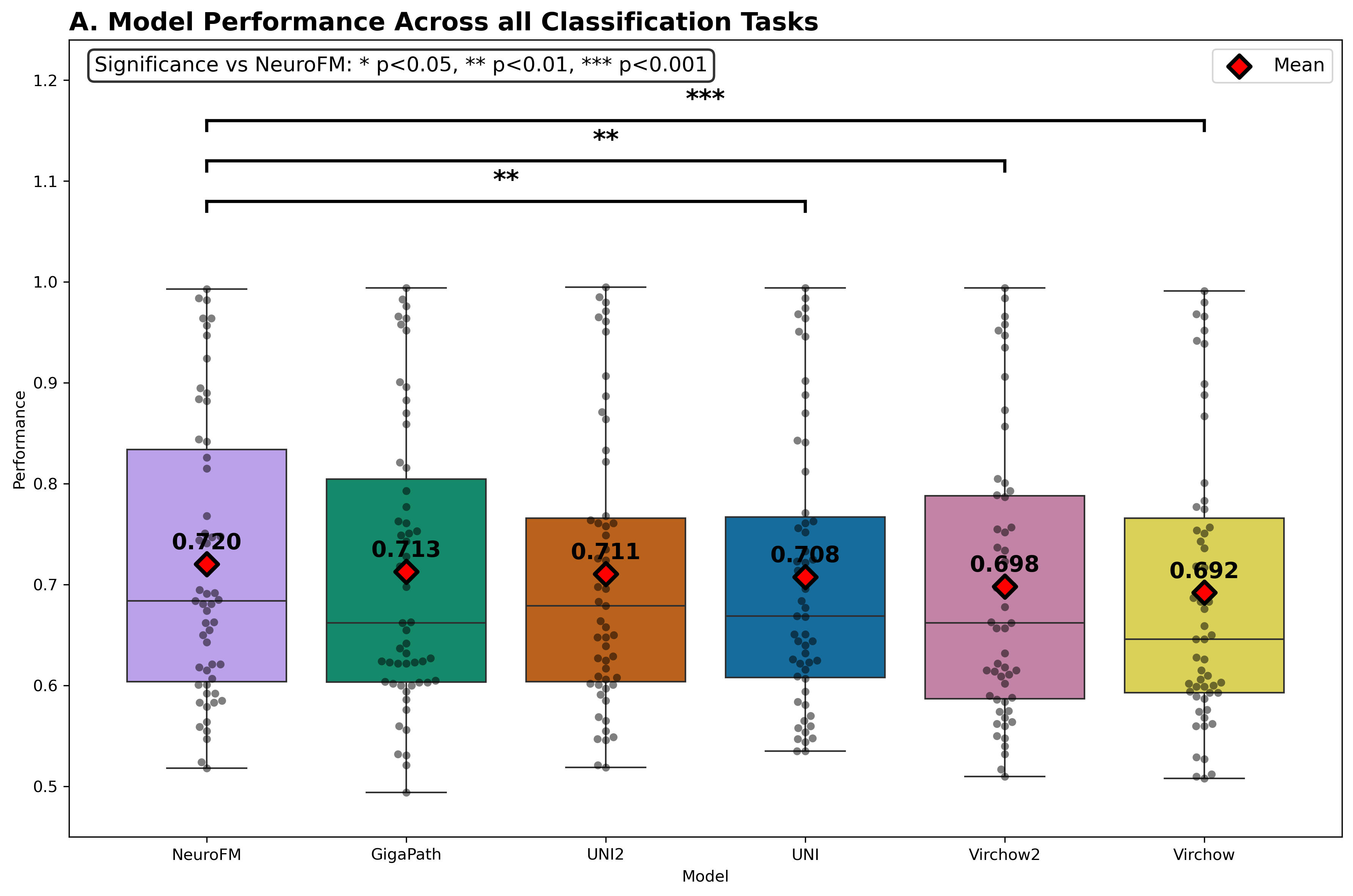
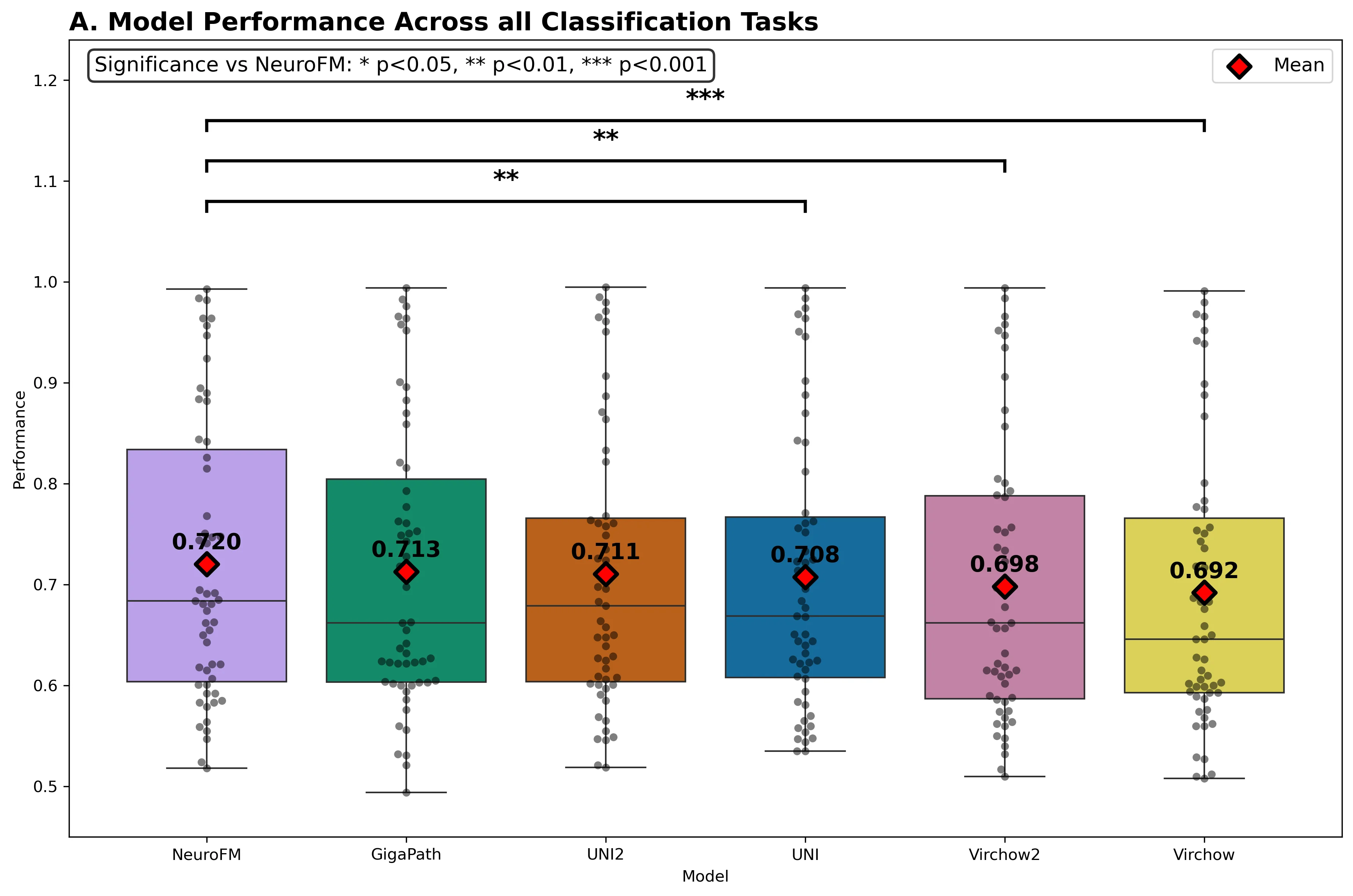
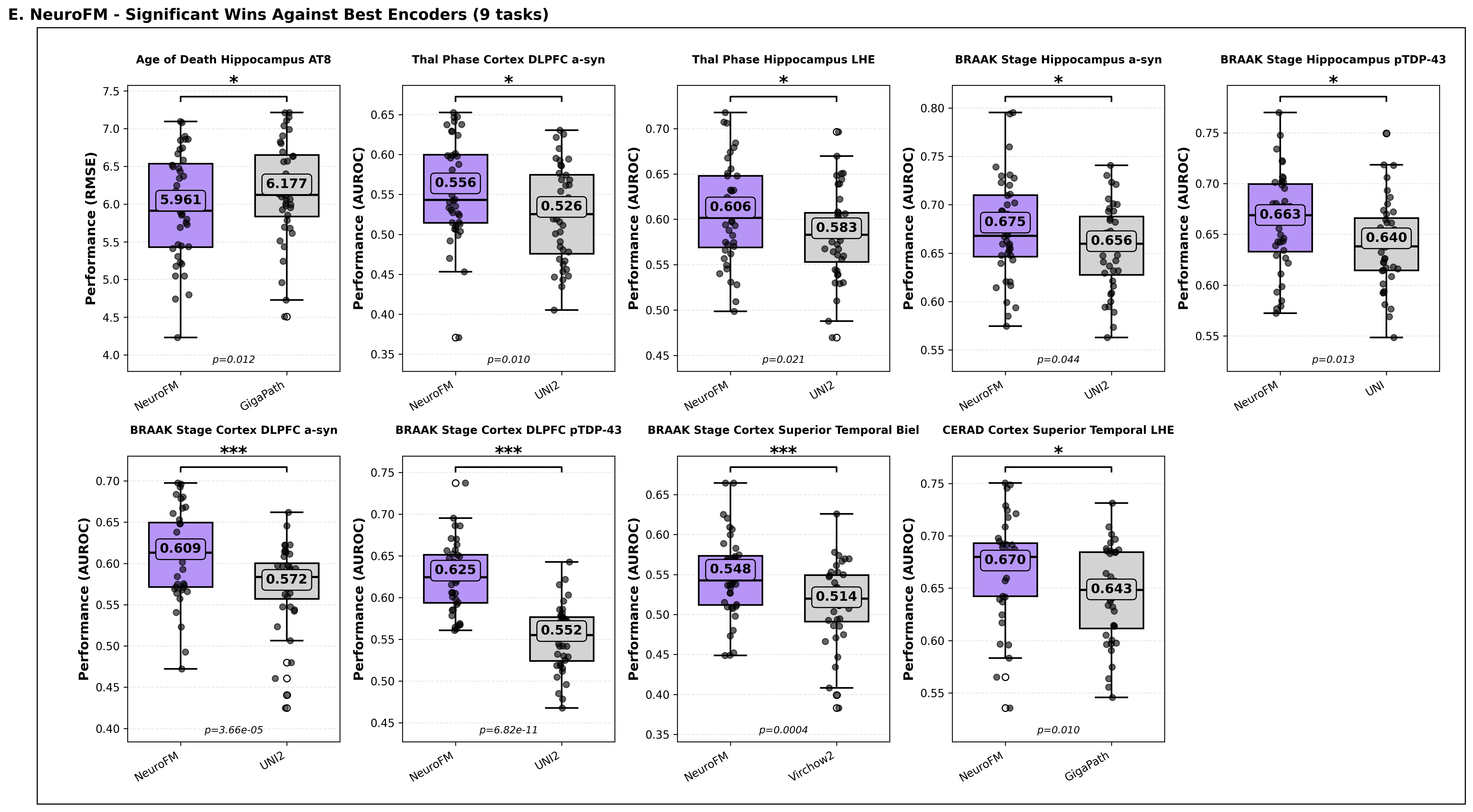
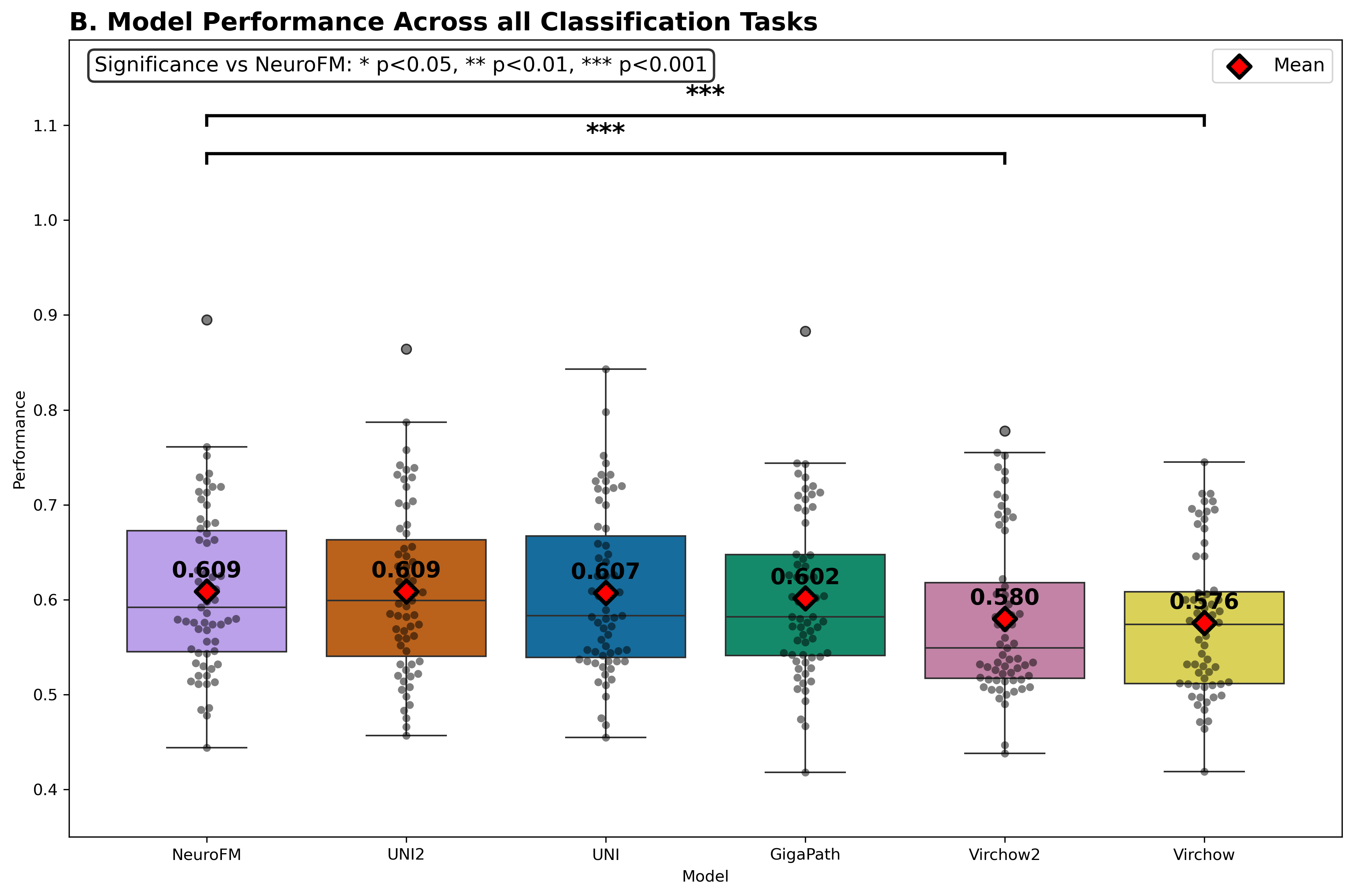
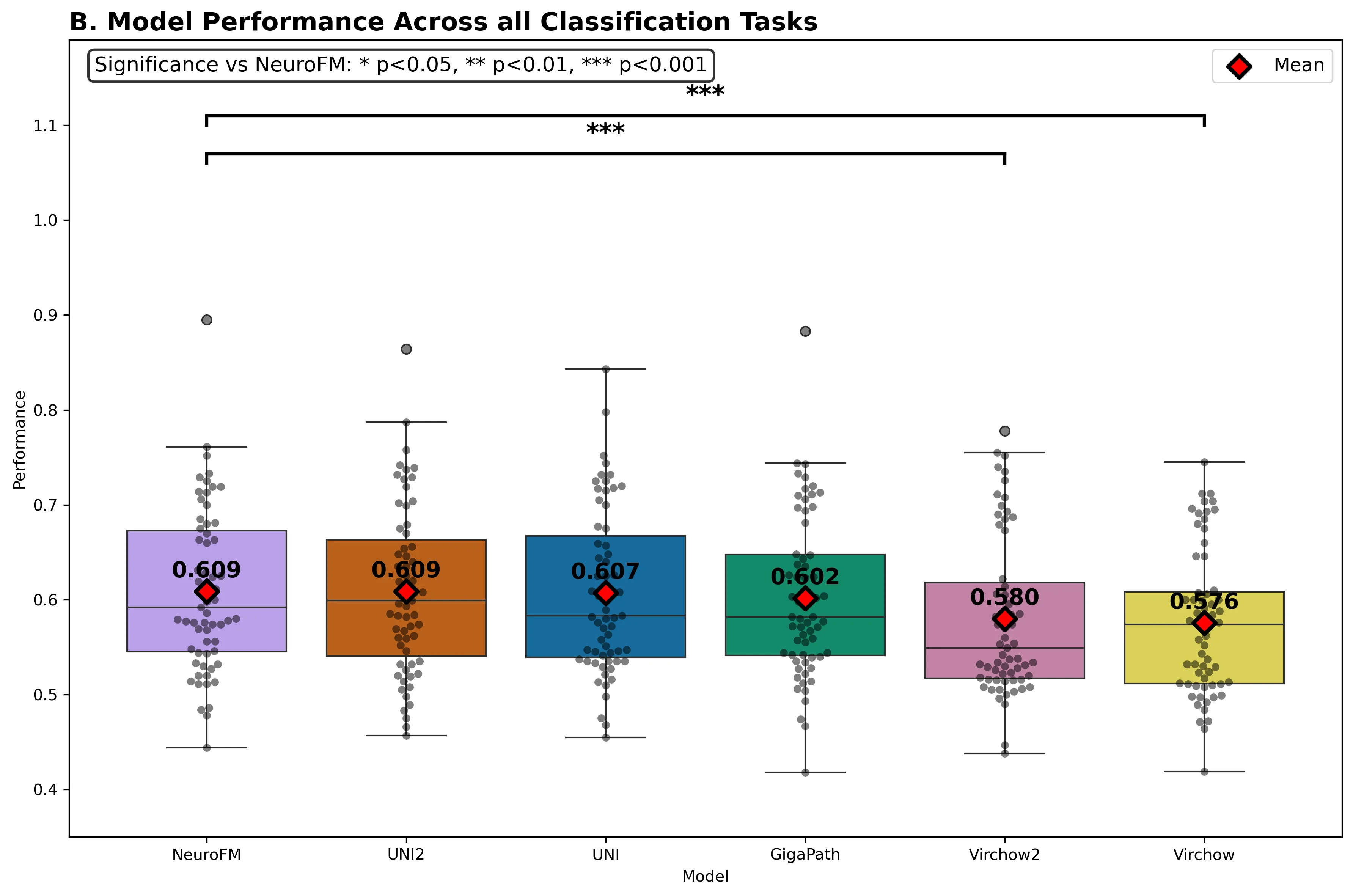
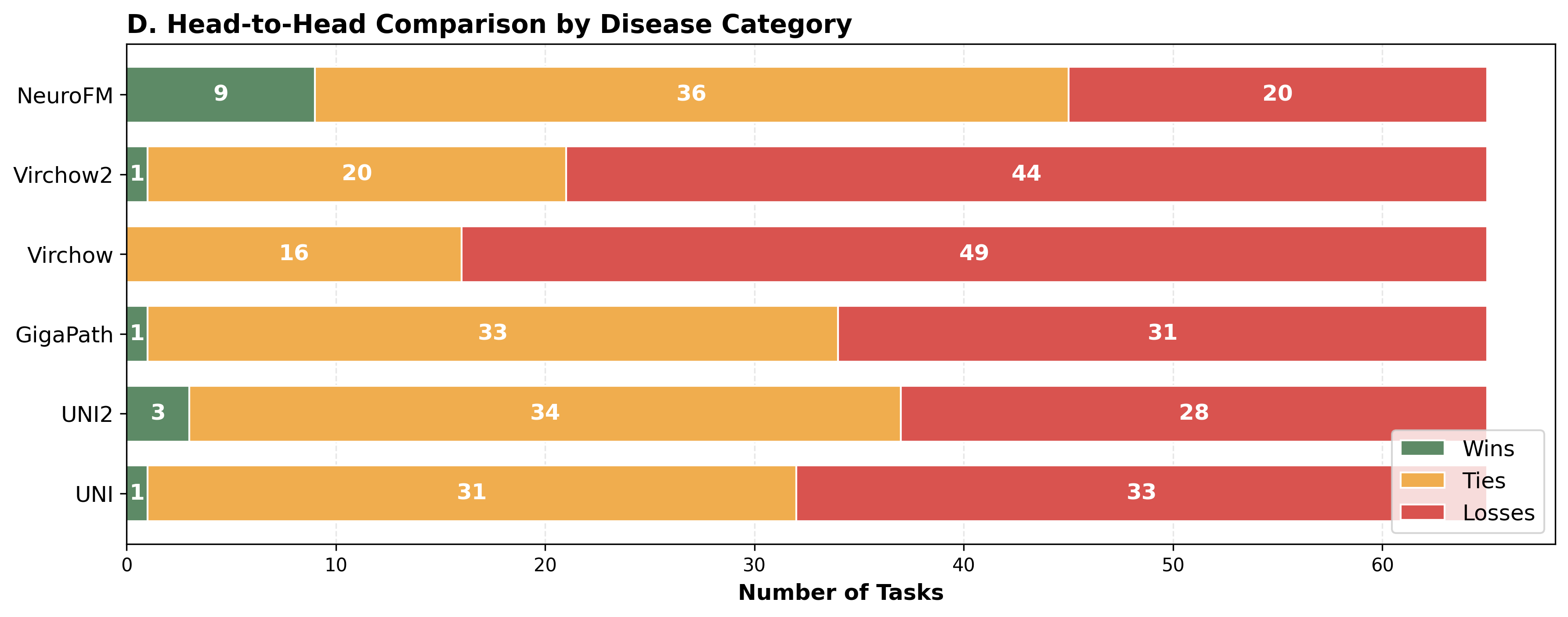
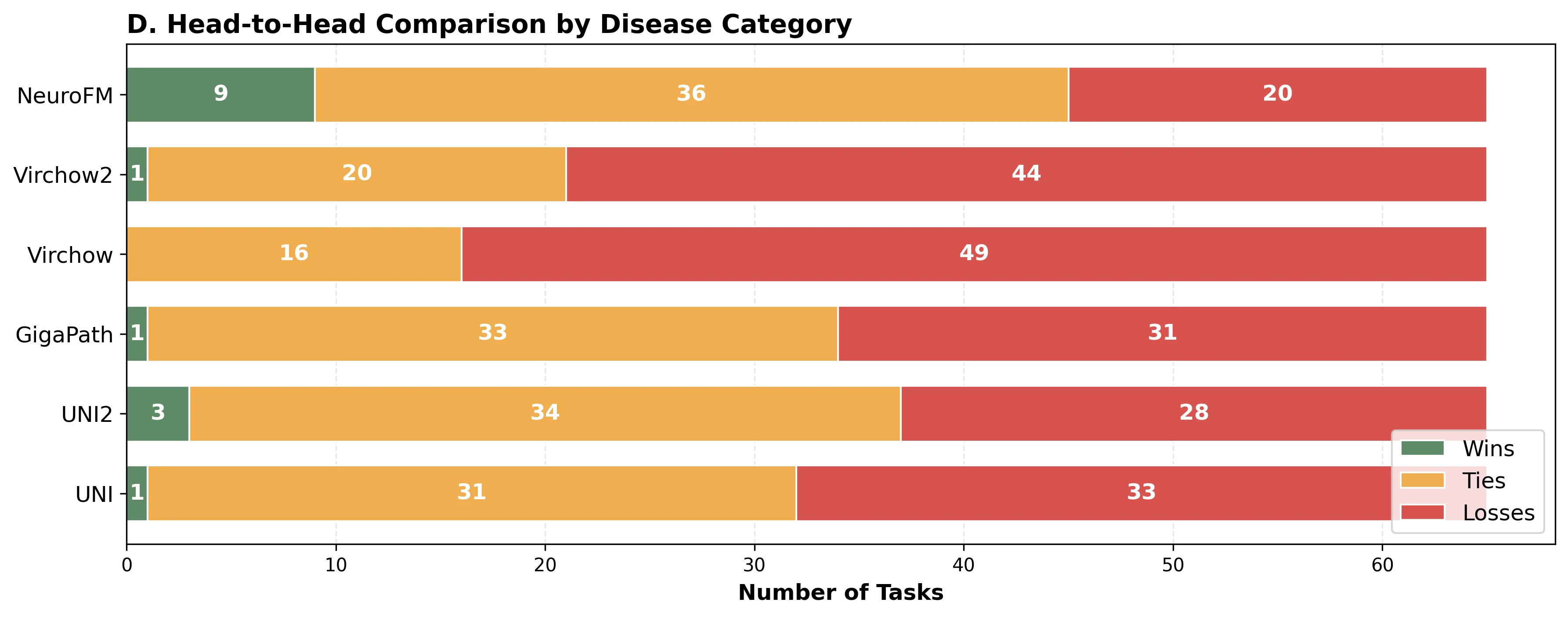
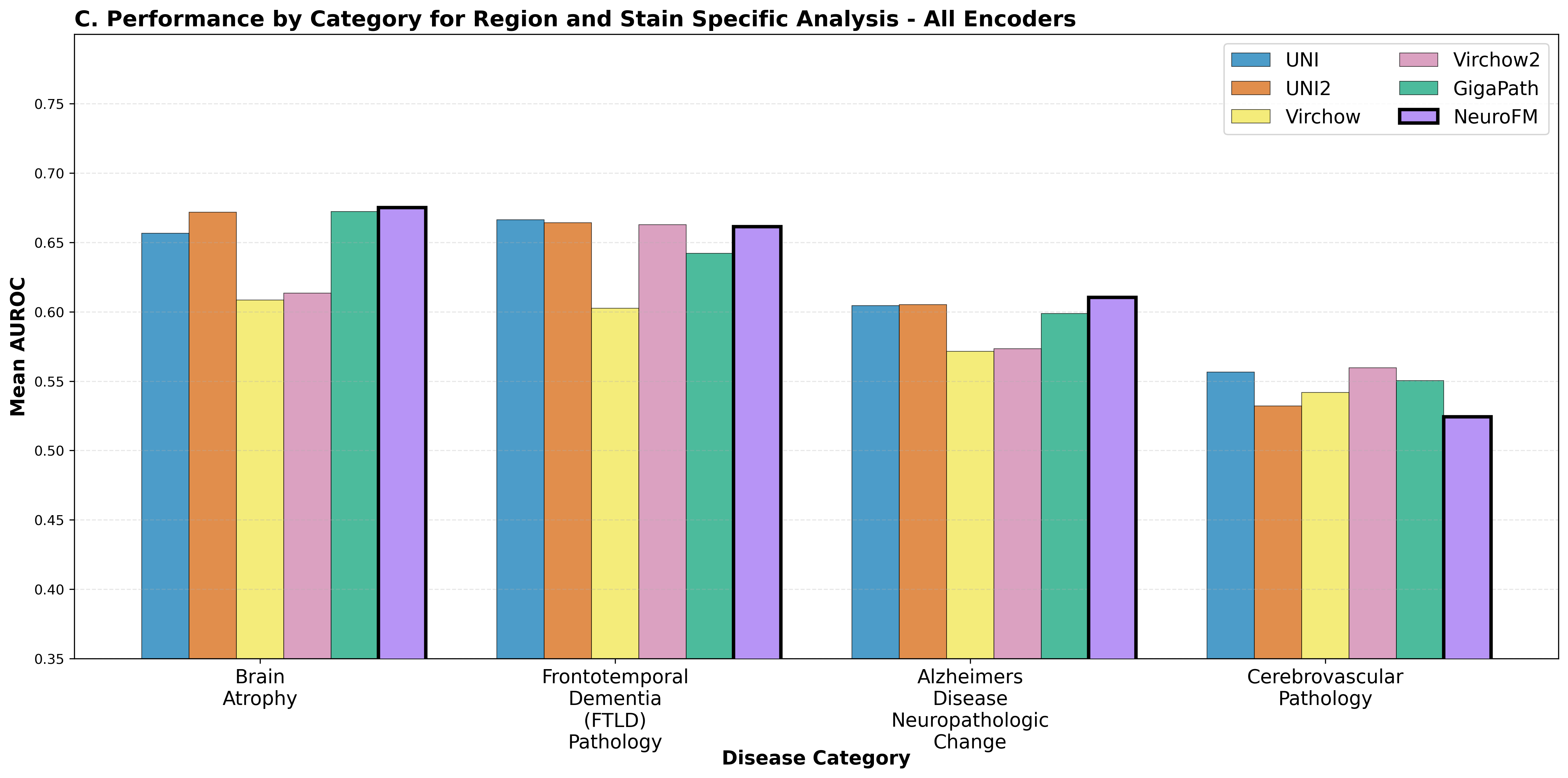
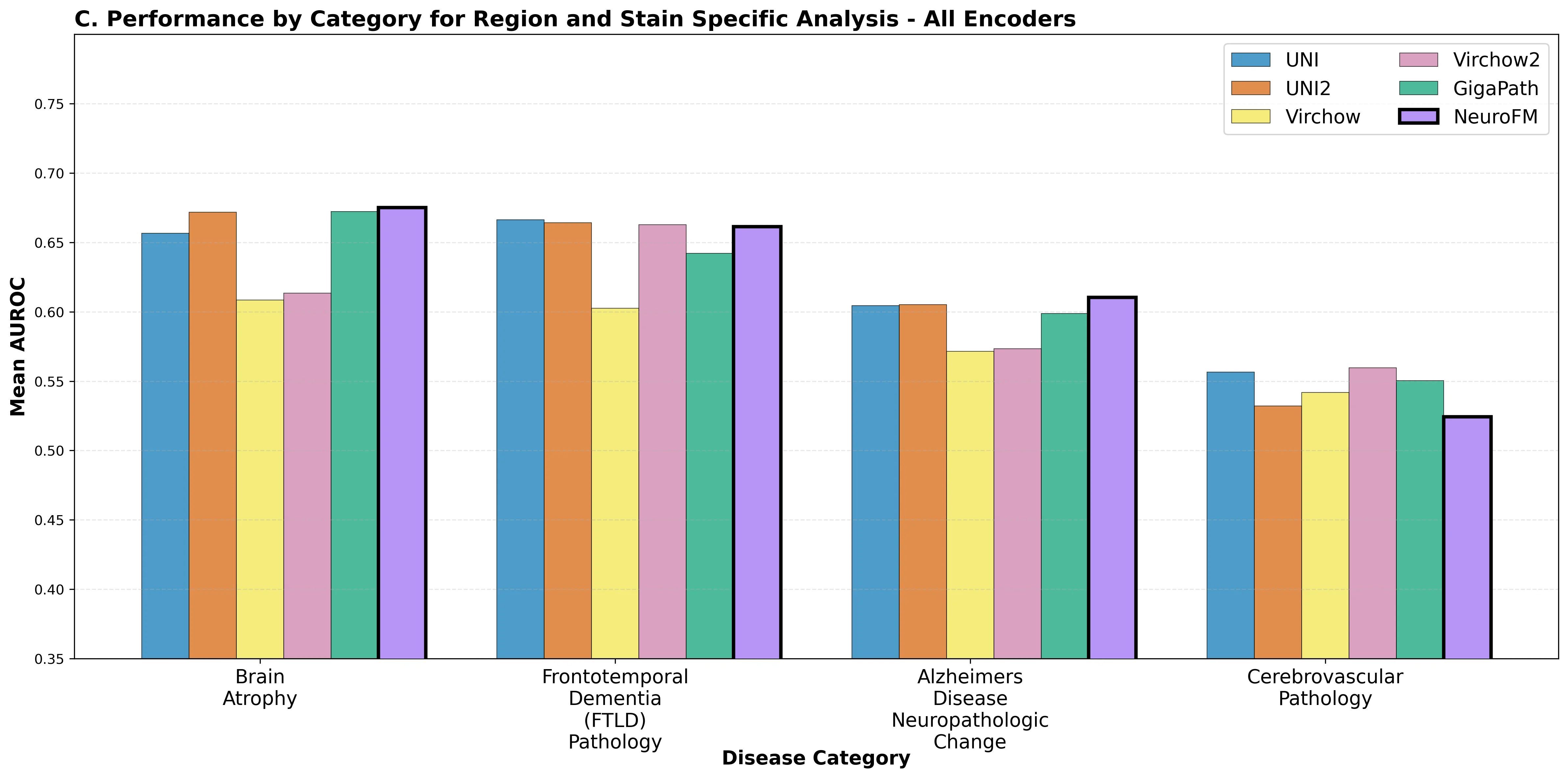
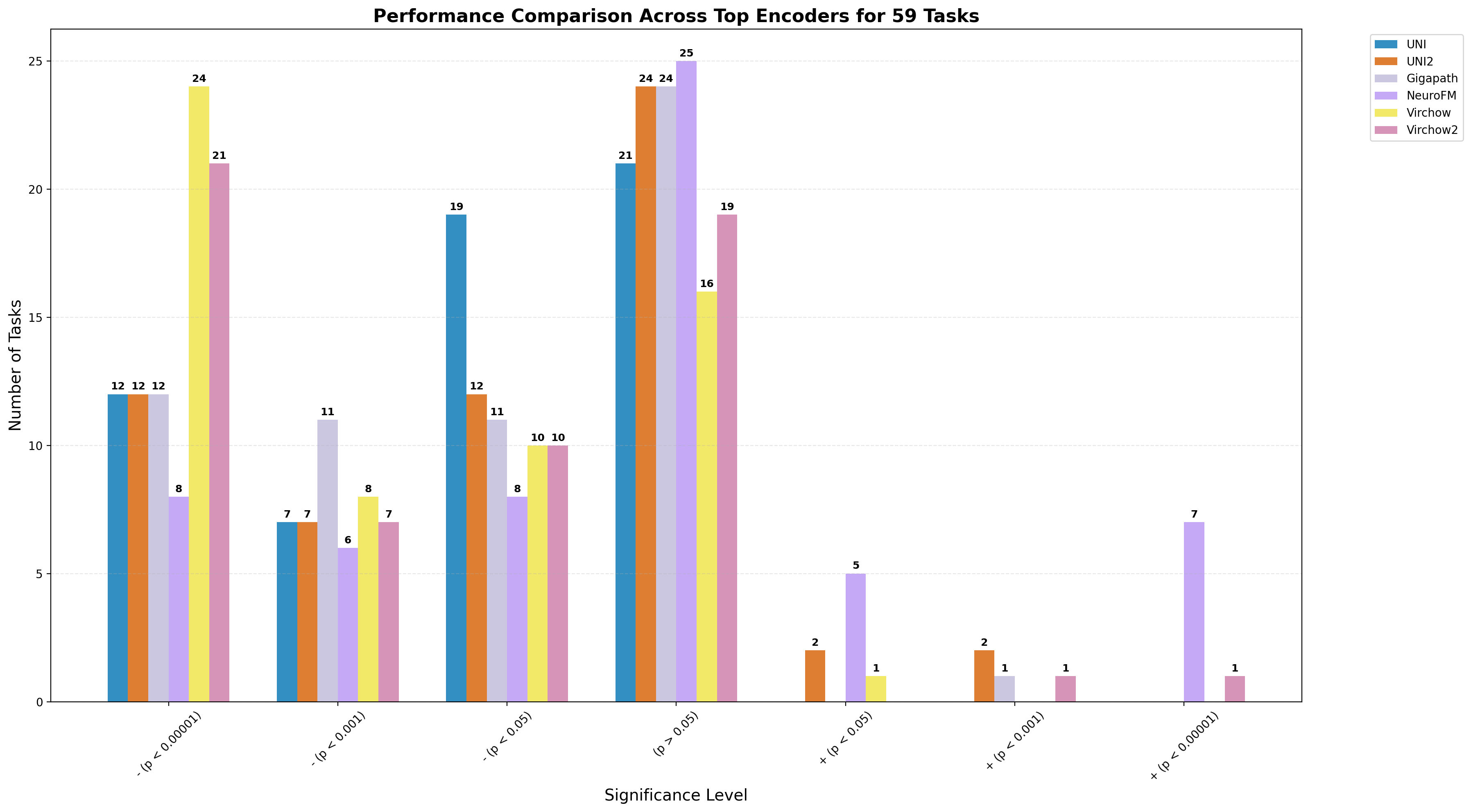
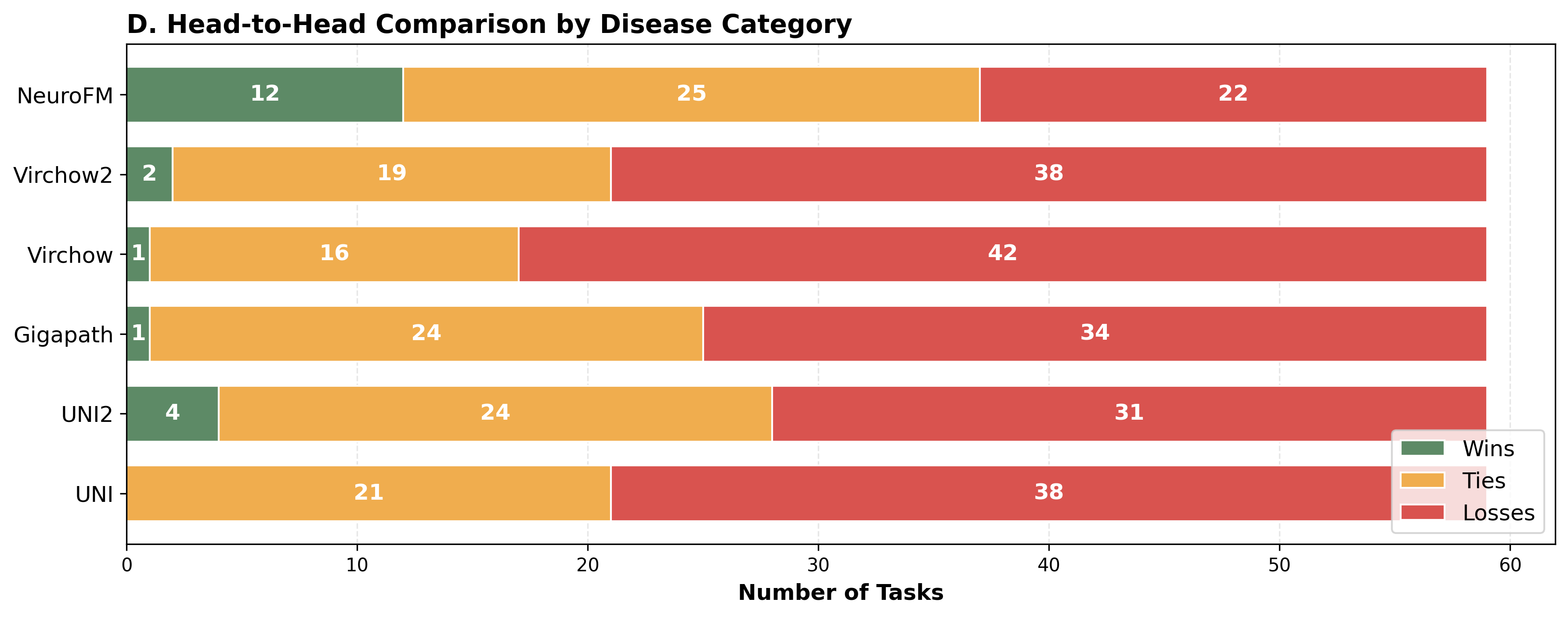
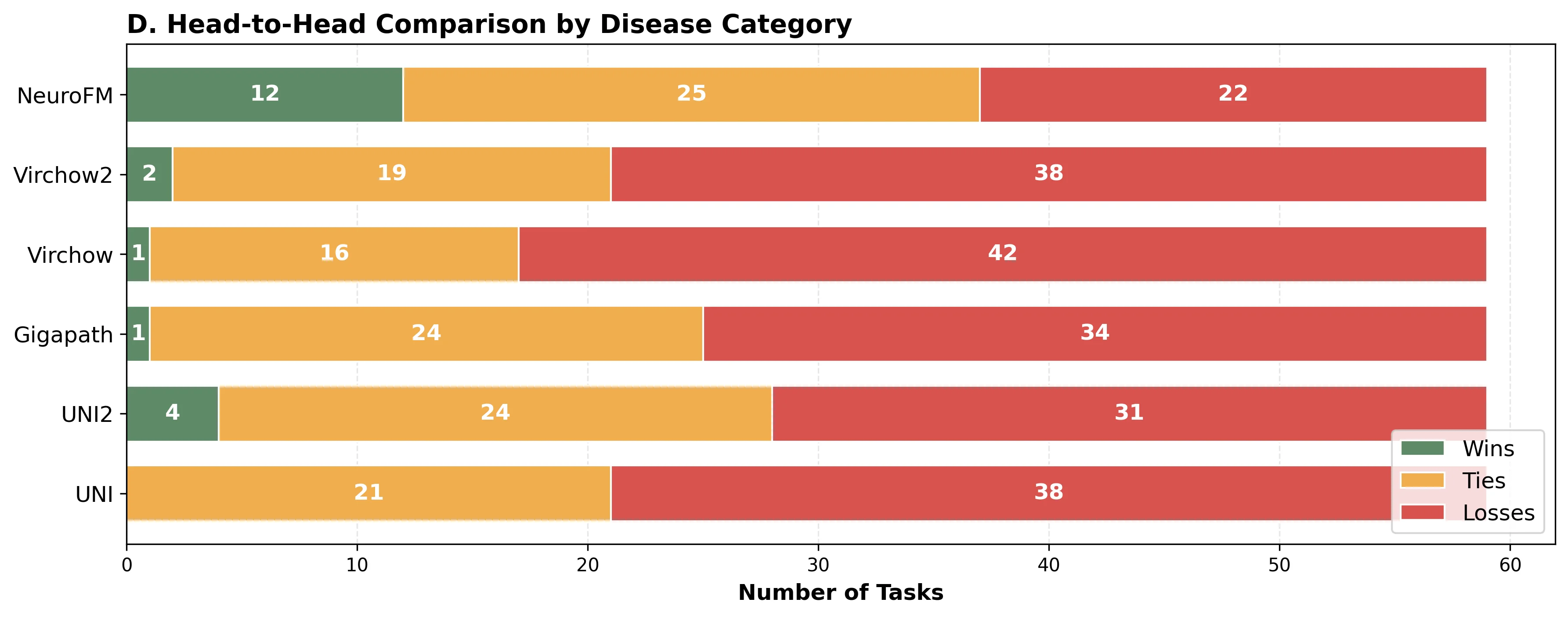
Foundation models have transformed computational pathology by providing generalizable representations from large-scale histology datasets. However, existing models are predominantly trained on surgical pathology data, which is enriched for non-nervous tissue and overrepresents neoplastic, inflammatory, metabolic, and other non-neurological diseases. Neuropathology represents a markedly different domain of histopathology, characterized by unique cell types (neurons, glia, etc.), distinct cytoarchitecture, and disease-specific pathological features including neurofibrillary tangles, amyloid plaques, Lewy bodies, and pattern-specific neurodegeneration. This domain mismatch may limit the ability of general-purpose foundation models to capture the morphological patterns critical for interpreting neurodegenerative diseases such as Alzheimer's disease, Parkinson's disease, and cerebellar ataxias. To address this gap, we developed NeuroFM, a foundation model trained specifically on whole-slide images of brain tissue spanning diverse neurodegenerative pathologies. NeuroFM demonstrates superior performance compared to general-purpose models across multiple neuropathology-specific downstream tasks, including mixed dementia disease classification, hippocampal region segmentation, and neurodegenerative ataxia identification encompassing cerebellar essential tremor and spinocerebellar ataxia subtypes. This work establishes that domain-specialized foundation models trained on brain tissue can better capture neuropathology-specific features than models trained on general surgical pathology datasets. By tailoring foundation models to the unique morphological landscape of neurodegenerative diseases, NeuroFM enables more accurate and reliable AI-based analysis for brain disease diagnosis and research, setting a precedent for domain-specific model development in specialized areas of digital pathology.
Deep Dive into Domain-Specific Foundation Model Improves AI-Based Analysis of Neuropathology.
Foundation models have transformed computational pathology by providing generalizable representations from large-scale histology datasets. However, existing models are predominantly trained on surgical pathology data, which is enriched for non-nervous tissue and overrepresents neoplastic, inflammatory, metabolic, and other non-neurological diseases. Neuropathology represents a markedly different domain of histopathology, characterized by unique cell types (neurons, glia, etc.), distinct cytoarchitecture, and disease-specific pathological features including neurofibrillary tangles, amyloid plaques, Lewy bodies, and pattern-specific neurodegeneration. This domain mismatch may limit the ability of general-purpose foundation models to capture the morphological patterns critical for interpreting neurodegenerative diseases such as Alzheimer’s disease, Parkinson’s disease, and cerebellar ataxias. To address this gap, we developed NeuroFM, a foundation model trained specifically on whole-slide
Domain-Specific Foundation Model Improves AI-Based Analysis
of Neuropathology
Ruchika Verma1,2*, Shrishtee Kandoi3,4, Robina Afzal3,4, Shengjia Chen1,2,
Jannes Jegminat1,2, Michael W. Karlovich3,4, Melissa Umphlett4,
Timothy E. Richardson4, 21, Kevin Clare4, Quazi Hossain4, Jorge Samanamud4,
Phyllis L. Faust5, Elan D. Louis10, 11, Ann C. McKee12, 13, 14, 15, 16,
Thor D. Stein12, 13, 15, 16, Jonathan D. Cherry15, 16, 12, 14, 13, 18, Jesse Mez15, 16, 14,
Anya C. McGoldrick19, 20, Dalilah D. Quintana Mora19,20,
Melissa J. Nirenberg19,3, 20, 21, Ruth H. Walker19,3, 20, 21, Yolfrankcis Mendez19, 20,
Susan Morgello3, 4, 7, Dennis W. Dickson8, Melissa E. Murray8,
Carlos Cordon-Cardo6, Nadejda M. Tsankova4, 7, Jamie M. Walker4, 7, 21,
Diana K. Dangoor4, 9, 1, 7, 20, 21, Stephanie McQuillan4, 20, Emma L. Thorn4, 20,
Claudia De Sanctis4, 1, 7, 9, 20, 21, Shuying Li22, 17, Thomas J. Fuchs1,2,
Kurt Farrell4,1, John F. Crary1,4,7,9,20,21*, Gabriele Campanella1,2*
1Windreich Department of AI and Human Health, Icahn School of Medicine at Mount Sinai,
New York, NY, USA.
2Hasso Plattner Institute for Digital Health at Mount Sinai, Icahn School of Medicine at
Mount Sinai, New York, NY, USA.
3Department of Neurology, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
4Department of Pathology, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
5Department of Pathology and Cell Biology, Columbia University, New York, NY, USA.
6Department of Pathology, Memorial Sloan-Kettering Cancer Center, New York, NY, USA.
7Department of Neuroscience, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
8Department of Neuroscience, Mayo Clinic College of Medicine, Jacksonville, FL, USA.
9Ronald M. Loeb Center for Alzheimer’s Disease, Icahn School of Medicine at Mount Sinai,
New York, NY, USA.
10Department of Neurology, University of Texas Southwestern Medical Center, Dallas, TX,
USA.
11Peter O’Donnell Jr. Brain Institute, University of Texas Southwestern Medical Center,
Dallas, TX, USA.
12VA Boston Healthcare System, Boston, MA, USA.
13Department of Pathology and Laboratory Medicine, Boston University Chobanian &
Avedisian School of Medicine, Boston University, Boston, MA, USA.
14Department of Neurology, Boston University Chobanian & Avedisian School of Medicine,
Boston, MA, USA.
15Boston University Alzheimer’s Disease Research Center, Boston University Chobanian &
Avedisian School of Medicine, Boston, MA, USA.
16Boston University Chronic Traumatic Encephalopathy Center, Boston University
Chobanian & Avedisian School of Medicine, Boston University, Boston, MA, USA.
17Department of Computer Science and Software Engineering, Miami University, Oxford,
OH, USA.
18Department of Anatomy & Neurobiology, Boston University Chobanian & Avedisian
School of Medicine, Boston, MA, USA.
19Department of Neurology, James J. Peters Veterans Affairs Medical Center, Bronx, NY,
USA.
1
arXiv:2512.05993v1 [cs.CV] 30 Nov 2025
20Neuropathology Brain Bank & Research CoRE, Icahn School of Medicine at Mount Sinai,
New York, NY, USA.
21Friedman Brain Institute, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
22Department of Chemical, Paper and Biomedical Engineering, Miami University, Oxford,
OH, USA.
*Corresponding author(s). E-mail(s): ruchika@mssm.edu; john.crary@mountsinai.org;
gabriele.campanella@mssm.edu;
Abstract
Foundation models have transformed computational pathology by providing generalizable representa-
tions from large-scale histology datasets. However, existing models are predominantly trained on surgical
pathology data, which is enriched for non-nervous tissue and overrepresents neoplastic, inflammatory,
metabolic, and other non-neurological diseases. Neuropathology represents a markedly different domain
of histopathology, characterized by unique cell types (neurons, glia, etc.), distinct cytoarchitecture, and
disease-specific pathological features including neurofibrillary tangles, amyloid plaques, Lewy bodies,
and pattern-specific neurodegeneration. This domain mismatch may limit the ability of general-purpose
foundation models to capture the morphological patterns critical for interpreting neurodegenerative dis-
eases such as Alzheimer’s disease, Parkinson’s disease, and cerebellar ataxias. To address this gap,
we developed NeuroFM, a foundation model trained specifically on whole-slide images of brain tissue
spanning diverse neurodegenerative pathologies. NeuroFM demonstrates superior performance compared
to general-purpose models across multiple neuropathology-specific downstream tasks, including mixed
dementia disease classification, hippocampal region segmentation, and neurodegenerative ataxia identifi-
cation encompassing cerebellar essential tremor and spinocerebellar ataxia subtypes. This work establishes
that domain-specialized foundation models trained on brain tissue can better capture neuropathology-
specific features than models trained on general surgical pathology datasets. By tailoring foundation
m
…(Full text truncated)…
This content is AI-processed based on ArXiv data.