In the process of patient diagnosis, non-invasive measurements are widely used due to their low risks and quick results. Electrocardiogram (ECG), as a non-invasive method to collect heart activities, is used to diagnose cardiac conditions. Analyzing the ECG typically requires domain expertise, which is a roadblock to applying artificial intelligence (AI) for healthcare. Through advances in self-supervised learning and foundation models, AI systems can now acquire and leverage domain knowledge without relying solely on human expertise. However, there is a lack of comprehensive analyses over the foundation models' performance on ECG. This study aims to answer the research question: "Are Foundation Models Useful for ECG Analysis?" To address it, we evaluate language/general time-series/ECG foundation models in comparison with time-series deep learning models. The experimental results show that general time-series/ECG foundation models achieve a top performance rate of 80%, indicating their effectiveness in ECG analysis. In-depth analyses and insights are provided along with comprehensive experimental results. This study highlights the limitations and potential of foundation models in advancing physiological waveform analysis. The data and code for this benchmark are publicly available at https://github.com/yuhaoxu99/ECGMultitasks-Benchmark.
Deep Dive into An Electrocardiogram Multi-task Benchmark with Comprehensive Evaluations and Insightful Findings.
In the process of patient diagnosis, non-invasive measurements are widely used due to their low risks and quick results. Electrocardiogram (ECG), as a non-invasive method to collect heart activities, is used to diagnose cardiac conditions. Analyzing the ECG typically requires domain expertise, which is a roadblock to applying artificial intelligence (AI) for healthcare. Through advances in self-supervised learning and foundation models, AI systems can now acquire and leverage domain knowledge without relying solely on human expertise. However, there is a lack of comprehensive analyses over the foundation models’ performance on ECG. This study aims to answer the research question: “Are Foundation Models Useful for ECG Analysis?” To address it, we evaluate language/general time-series/ECG foundation models in comparison with time-series deep learning models. The experimental results show that general time-series/ECG foundation models achieve a top performance rate of 80%, indicating thei
Electrocardiogram (ECG) records the heart's electrical activities via skin-placed electrodes [1], producing waveforms that decipher cardiac functions. Its non-invasive nature and ease of collection make ECG ideal for continuous monitoring and early detection of cardiovascular abnormalities. ECG is used for diagnosing arrhythmias [2], myocardial infarctions [3] and analyzing heart rate variability [4], highlighting its diverse utilities. However, ECG analysis is challenging due to individual variations, complex waveforms, and susceptibility to noises [5]. Traditional ECG analysis relies on specialized clinicians, December 2024 which is resource-intensive and does not scale well with large data volumes, increasing the risk of diagnostic errors. Advances in artificial intelligence (AI) have led to AI-assisted ECG diagnostics surpassing human performance [6].
AI models enhance ECG analysis by extracting rich features. Beyond detecting heart diseases, ECG can infer age [7], gender [7], blood pressure [8], and potassium levels [9]. Al-Zaiti et al. [6] used a random forest model that outperformed clinicians and FDA-approved systems in detecting acute myocardial ischemia. While traditional machine learning relies on feature engineering, potentially losing clinically relevant information, neural networks can use raw ECG signals, preserving critical information. Baloglu et al. [10] achieved over 99% accuracy in myocardial infarction detection using a convolutional neural network. However, neural networks require extensive labeled data, which may not always be available. Foundation models address this by leveraging largescale pre-training and task-specific fine-tuning. McKeen et al. [11] proposed ECG-FM, a transformer-based model pre-trained on 2.5 million samples, demonstrating the strong potential of unsupervised foundation models. Despite the emergence of ECG foundation models, fair and comprehensive evaluations on their effectiveness are lacking.
In this study, we construct a benchmark to fairly evaluate existing foundation models for ECG analysis, including large language model (LLM), time-series foundation model (TSFM), and an ECG foundation model (ECGFM), in contrast to traditional timeseries deep learning model (TSDL). We compare their performance across five tasks, assessing ECG data modeling from different perspectives: simple feature extraction (RR interval estimation), complex feature extraction (age estimation), balanced labels (gender classification), imbalanced labels (potassium abnormality prediction), and multi-class classification (arrhythmia detection). Our evaluation scenarios encompass zero-shot, fewshot, and fine-tuning approaches. Through these comparisons, we analyze the strengths and weaknesses of different models and explore the effectiveness of foundation models. We envision our findings can inspire advancements in using foundation models for physiological waveform analysis. Our code is open-sourced to support future research.
Our experiment is conducted on the MIMIC-IV-ECG [12] dataset, which is currently the largest publicly accessible ECG dataset, comprising 800,035 diagnostic electrocardiograms from 161,352 unique patients. Each ECG strip is 12-lead and 10 seconds in length with 500 Hz sampling rate, denoted by x ∈ R C×L where C = 12 and L = 10 × 500 = 5000. Downstream tasks. We evaluate the performance of the benchmark on the following tasks: (1) RR Interval Estimation. The RR interval, which represents the time between two R-wave peaks in an ECG, is directly calculated from the ECG signal. (2) Age Estimation. Patient age estimation involves analyzing ECG signal characteristics to estimate age, challenging the model to effectively interpret complex signal patterns correlated with physiological aging. (3) Gender Classification. Gender classification is a binary classification task with a roughly balanced ratio of 50% to 50%. (4) Potassium Abnormality Prediction. We use ECG strips to predict the Potassium (blood) lab test result which is taken between ECG recording time and one hour after the ECG time. This task is challengening, with imbalanced ratio of 97% (normal) to 3% (abnormal). (5) Arrhythmia Detection. We select the 14 most frequently occurring diagnoses, with the remaining ones grouped under “Others”, resulting in a total of 15 labels. Among these downstream tasks, RR interval estimation and age estimation are regression tasks, where the prediction target y ∈ R. Gender prediction and potassium abnormality prediction are binary classification tasks, where the prediction target y ∈ {0, 1}. Arrhythmia detection is multiclass classification task, where the prediction target y ∈ {1, 2, 3, . . . , M} (M = 15 denotes the phenotype of arrhythmia).
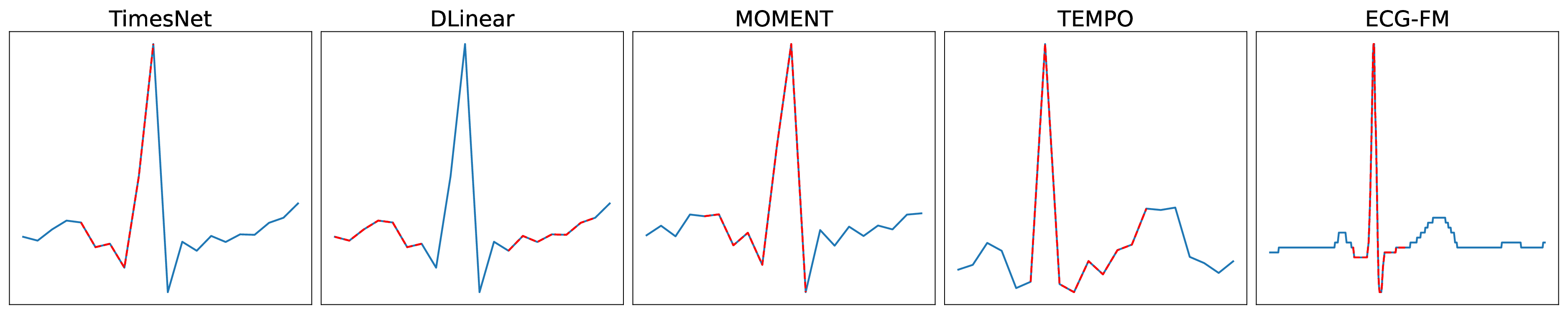
Evaluated Models. We select the following models for benchmarking: TimesNet [13], DLinear [14], GPT-2 [15], Llama 3.1 [16], MOMENT [17], TEMPO [18] and ECG-FM [11]. The details of these models are shown in Table 1. For the TSDL, TSFM, and E
…(Full text truncated)…
This content is AI-processed based on ArXiv data.