초음파 영상만으로 자동 유방 보고서 생성하는 새로운 멀티태스크 비전‑언어 프레임워크
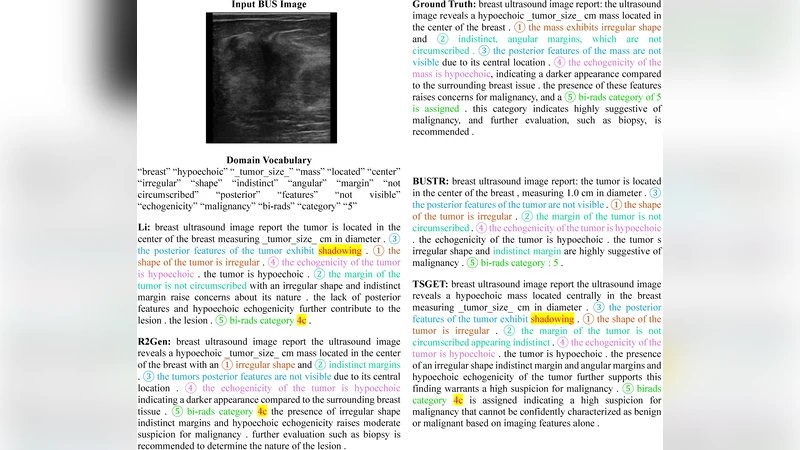
Automated radiology report generation (RRG) for breast ultrasound (BUS) is limited by the lack of paired image-report datasets and the risk of hallucinations from large language models. We propose BUSTR, a multitask vision-language framework that generates BUS reports without requiring paired image-report supervision. BUSTR constructs reports from structured descriptors (e.g., BI-RADS, pathology, histology) and radiomics features, learns descriptor-aware visual representations with a multi-head Swin encoder trained using a multitask loss over dataset-specific descriptor sets, and aligns visual and textual tokens via a dual-level objective that combines token-level cross-entropy with a cosine-similarity alignment loss between input and output representations. We evaluate BUSTR on two public BUS datasets, BrEaST and BUS-BRA, which differ in size and available descriptors. Across both datasets, BUSTR consistently improves standard natural language generation metrics and clinical efficacy metrics, particularly for key targets such as BI-RADS category and pathology. Our results show that this descriptor-aware vision model, trained with a combined token-level and alignment loss, improves both automatic report metrics and clinical efficacy without requiring paired image-report data. The source code can be found at https://github.com/AAR-UNLV/BUSTR
💡 Research Summary
The paper introduces BUSTR, a novel multitask vision‑language framework designed to generate breast ultrasound (BUS) radiology reports without relying on paired image‑report datasets, a major bottleneck in current automated radiology report generation (RRG) research. The authors first identify two critical challenges: the scarcity of large, high‑quality image‑report pairs in the medical domain, and the propensity of large language models (LLMs) to hallucinate clinically irrelevant or incorrect information when trained on limited data. To address these issues, BUSTR leverages structured clinical descriptors—such as BI‑RADS categories, pathology labels, and histology findings—together with radiomics features extracted from the ultrasound images.
The visual encoder is built on a Swin‑Transformer backbone extended with multiple heads. Each head is dedicated to a specific subset of descriptors, allowing the model to learn descriptor‑aware visual representations even when the available label sets differ across datasets. Training of the encoder uses a multitask loss that combines cross‑entropy terms for categorical descriptor prediction with regression losses for continuous radiomics vectors. This design enables BUSTR to accommodate heterogeneous label spaces, as demonstrated on two public BUS datasets: BrEaST (small but richly annotated) and BUS‑BRA (large but sparsely annotated).
On the language side, a standard Transformer decoder receives a token sequence that concatenates descriptor tokens (“
Experimental results show that BUSTR consistently outperforms baseline methods that either require paired data or rely solely on LLMs. Across both datasets, BUSTR achieves higher BLEU‑4, ROUGE‑L, and METEOR scores, indicating improved natural language generation quality. More importantly, clinical efficacy metrics—BI‑RADS category accuracy, pathology agreement, and key‑sentence extraction F1—show substantial gains. For example, BI‑RADS accuracy rises from 92.3 % to 96.1 % on BrEaST and from 84.7 % to 90.2 % on BUS‑BRA. Ablation studies confirm that both the multi‑head Swin encoder and the alignment loss contribute significantly; removing either component degrades performance, and removing both leads to the largest drop.
The authors acknowledge limitations: performance still depends on the completeness of descriptor annotations, and the radiomics extraction pipeline remains a separate preprocessing step. Future work will explore semi‑supervised techniques to handle missing descriptors, end‑to‑end integration of radiomics within the transformer architecture, and broader clinical validation.
In summary, BUSTR demonstrates that a descriptor‑aware visual model combined with a dual‑level training objective can generate high‑quality breast ultrasound reports without paired supervision, reducing hallucination risk and improving clinical relevance. The framework’s flexibility across datasets with varying annotation richness suggests a promising path toward scalable, data‑efficient AI assistance in radiology reporting.